| GO ID | Tissue | Disease Stage | Description | Gene Ratio | Bg Ratio | pvalue | p.adjust | Count |
| GO:004603422 | Liver | HCC | ATP metabolic process | 198/7958 | 277/18723 | 8.30e-23 | 1.55e-20 | 198 |
| GO:004854522 | Liver | HCC | response to steroid hormone | 206/7958 | 339/18723 | 6.81e-12 | 2.92e-10 | 206 |
| GO:007149622 | Liver | HCC | cellular response to external stimulus | 191/7958 | 320/18723 | 3.40e-10 | 1.13e-08 | 191 |
| GO:007138322 | Liver | HCC | cellular response to steroid hormone stimulus | 128/7958 | 204/18723 | 3.92e-09 | 1.04e-07 | 128 |
| GO:003297022 | Liver | HCC | regulation of actin filament-based process | 222/7958 | 397/18723 | 3.81e-08 | 8.30e-07 | 222 |
| GO:001631121 | Liver | HCC | dephosphorylation | 230/7958 | 417/18723 | 1.00e-07 | 1.96e-06 | 230 |
| GO:005123521 | Liver | HCC | maintenance of location | 185/7958 | 327/18723 | 1.70e-07 | 3.12e-06 | 185 |
| GO:007121421 | Liver | HCC | cellular response to abiotic stimulus | 183/7958 | 331/18723 | 1.59e-06 | 2.26e-05 | 183 |
| GO:010400421 | Liver | HCC | cellular response to environmental stimulus | 183/7958 | 331/18723 | 1.59e-06 | 2.26e-05 | 183 |
| GO:005165112 | Liver | HCC | maintenance of location in cell | 119/7958 | 214/18723 | 7.11e-05 | 6.39e-04 | 119 |
| GO:19026007 | Liver | HCC | proton transmembrane transport | 87/7958 | 157/18723 | 7.27e-04 | 4.42e-03 | 87 |
| GO:00712601 | Liver | HCC | cellular response to mechanical stimulus | 48/7958 | 81/18723 | 1.72e-03 | 8.96e-03 | 48 |
| GO:00096124 | Liver | HCC | response to mechanical stimulus | 113/7958 | 216/18723 | 2.20e-03 | 1.10e-02 | 113 |
| GO:00093146 | Liver | HCC | response to radiation | 223/7958 | 456/18723 | 3.08e-03 | 1.43e-02 | 223 |
| GO:015010411 | Liver | HCC | transport across blood-brain barrier | 50/7958 | 87/18723 | 3.42e-03 | 1.55e-02 | 50 |
| GO:001023211 | Liver | HCC | vascular transport | 50/7958 | 88/18723 | 4.69e-03 | 2.02e-02 | 50 |
| GO:00551191 | Liver | HCC | relaxation of cardiac muscle | 13/7958 | 17/18723 | 4.77e-03 | 2.02e-02 | 13 |
| GO:00860642 | Liver | HCC | cell communication by electrical coupling involved in cardiac conduction | 17/7958 | 25/18723 | 8.94e-03 | 3.45e-02 | 17 |
| GO:0010882 | Liver | HCC | regulation of cardiac muscle contraction by calcium ion signaling | 18/7958 | 27/18723 | 9.75e-03 | 3.66e-02 | 18 |
| GO:0010881 | Liver | HCC | regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion | 15/7958 | 22/18723 | 1.35e-02 | 4.80e-02 | 15 |
| Hugo Symbol | Variant Class | Variant Classification | dbSNP RS | HGVSc | HGVSp | HGVSp Short | SWISSPROT | BIOTYPE | SIFT | PolyPhen | Tumor Sample Barcode | Tissue | Histology | Sex | Age | Stage | Therapy Types | Drugs | Outcome |
| ATP1A2 | SNV | Missense_Mutation | | c.144G>T | p.Glu48Asp | p.E48D | P50993 | protein_coding | tolerated(0.16) | benign(0.029) | TCGA-A2-A25A-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Unspecific | Cytoxan | SD |
| ATP1A2 | SNV | Missense_Mutation | rs755353911 | c.1529N>A | p.Arg510His | p.R510H | P50993 | protein_coding | deleterious(0.01) | probably_damaging(1) | TCGA-A8-A09I-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Hormone Therapy | anastrozole | SD |
| ATP1A2 | SNV | Missense_Mutation | | c.2996N>T | p.Asp999Val | p.D999V | P50993 | protein_coding | deleterious(0) | probably_damaging(0.999) | TCGA-A8-A09Z-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| ATP1A2 | SNV | Missense_Mutation | | c.1079N>G | p.Glu360Gly | p.E360G | P50993 | protein_coding | deleterious(0.01) | probably_damaging(1) | TCGA-AQ-A04J-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Chemotherapy | cytoxan | SD |
| ATP1A2 | SNV | Missense_Mutation | novel | c.1055A>G | p.Lys352Arg | p.K352R | P50993 | protein_coding | deleterious(0.03) | probably_damaging(0.999) | TCGA-AR-A1AM-01 | Breast | breast invasive carcinoma | Female | <65 | III/IV | Chemotherapy | adriamycin | SD |
| ATP1A2 | SNV | Missense_Mutation | | c.250A>G | p.Thr84Ala | p.T84A | P50993 | protein_coding | deleterious(0.01) | possibly_damaging(0.577) | TCGA-BH-A18G-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| ATP1A2 | SNV | Missense_Mutation | rs768136835 | c.713N>A | p.Arg238His | p.R238H | P50993 | protein_coding | deleterious(0) | benign(0.071) | TCGA-E2-A14U-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Hormone Therapy | arimidex | SD |
| ATP1A2 | insertion | Frame_Shift_Ins | novel | c.2921_2922insATCTATAATGAAACCACCACCAGGAACTGTGGGAGATGCTGAGA | p.Leu975SerfsTer102 | p.L975Sfs*102 | P50993 | protein_coding | | | TCGA-A8-A06Q-01 | Breast | breast invasive carcinoma | Female | <65 | III/IV | Unknown | Unknown | SD |
| ATP1A2 | insertion | Nonsense_Mutation | novel | c.964_965insACTGAGACT | p.Leu322delinsHisTerAspPhe | p.L322delinsH*DF | P50993 | protein_coding | | | TCGA-A8-A090-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| ATP1A2 | insertion | In_Frame_Ins | novel | c.1799_1800insTTCCCAAGCCTCTGCGGCATCCCTGGGGTG | p.Val600_Gly601insSerGlnAlaSerAlaAlaSerLeuGlyTrp | p.V600_G601insSQASAASLGW | P50993 | protein_coding | | | TCGA-BH-A0B8-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Hormone Therapy | arimidex | SD |
| Entrez ID | Symbol | Category | Interaction Types | Drug Claim Name | Drug Name | PMIDs |
| 477 | ATP1A2 | ENZYME, TRANSPORTER, ION CHANNEL, DRUGGABLE GENOME | | olanzapine | OLANZAPINE | |
| 477 | ATP1A2 | ENZYME, TRANSPORTER, ION CHANNEL, DRUGGABLE GENOME | inhibitor | CHEMBL3545057 | ACETYLDIGITOXIN | |
| 477 | ATP1A2 | ENZYME, TRANSPORTER, ION CHANNEL, DRUGGABLE GENOME | inhibitor | CHEMBL254219 | DIGITOXIN | |
| 477 | ATP1A2 | ENZYME, TRANSPORTER, ION CHANNEL, DRUGGABLE GENOME | inhibitor | CHEMBL1751 | DIGOXIN | |
| 477 | ATP1A2 | ENZYME, TRANSPORTER, ION CHANNEL, DRUGGABLE GENOME | inhibitor | CHEMBL1614 | DESLANOSIDE | |







 Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells
Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells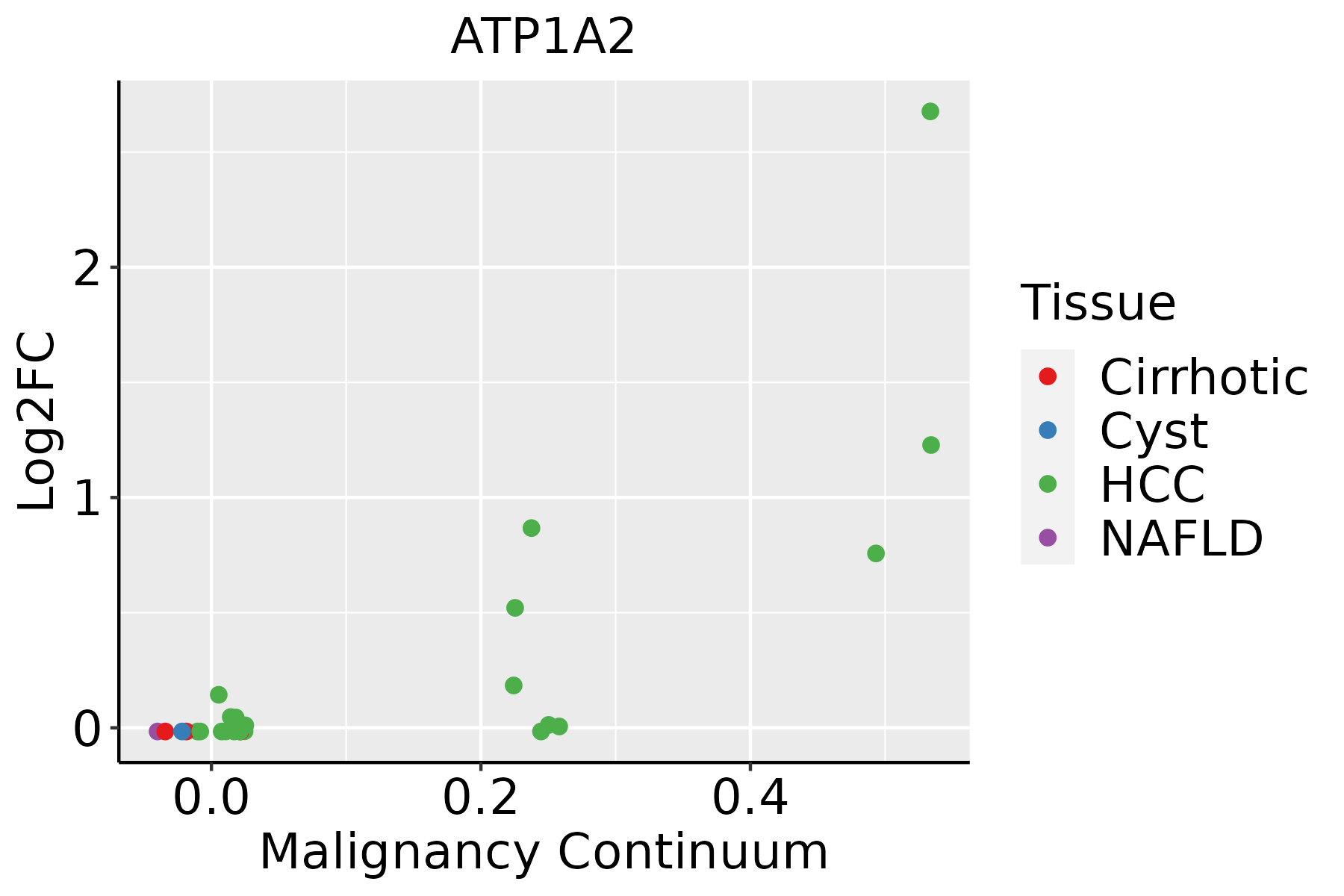
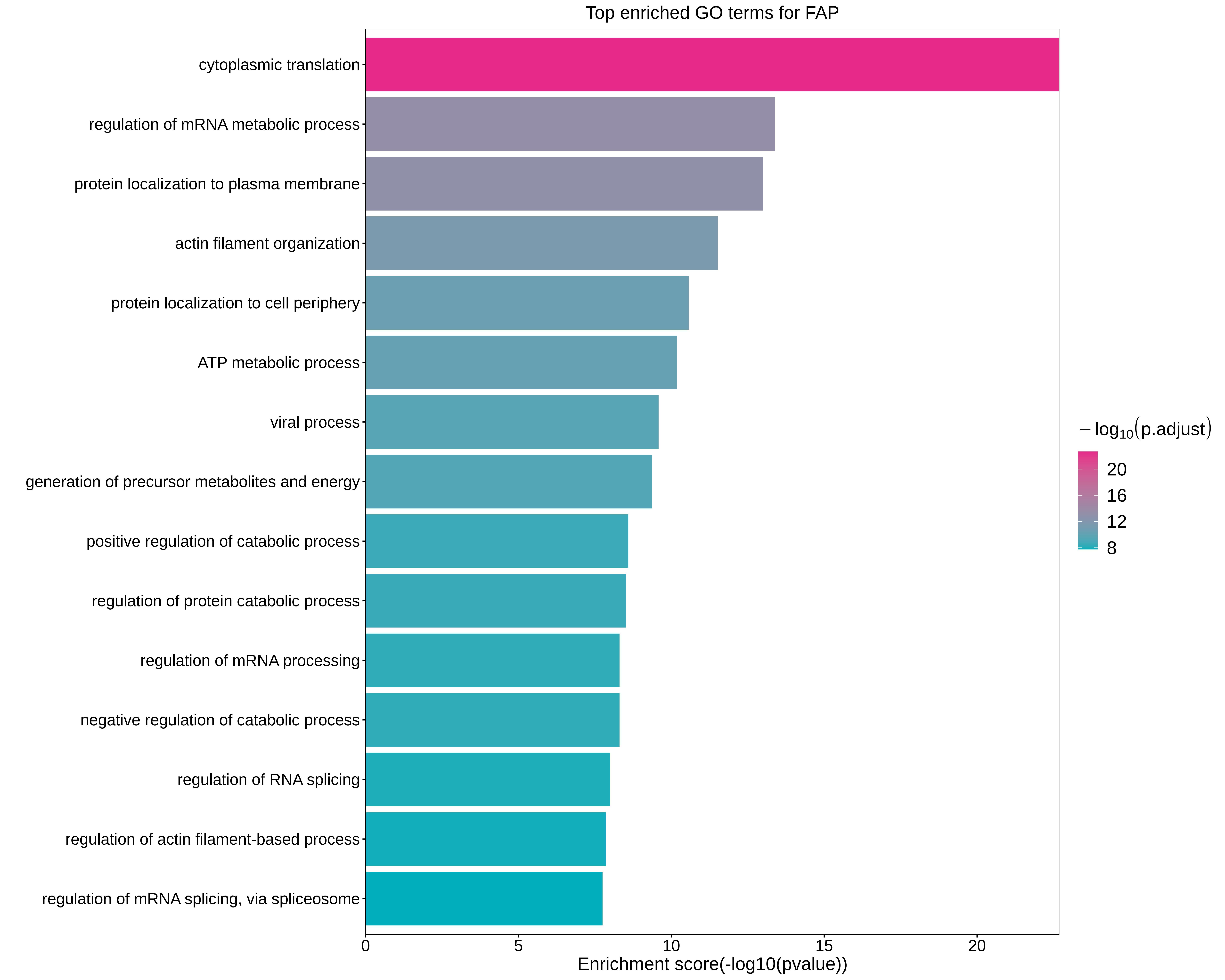
 Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer
Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer
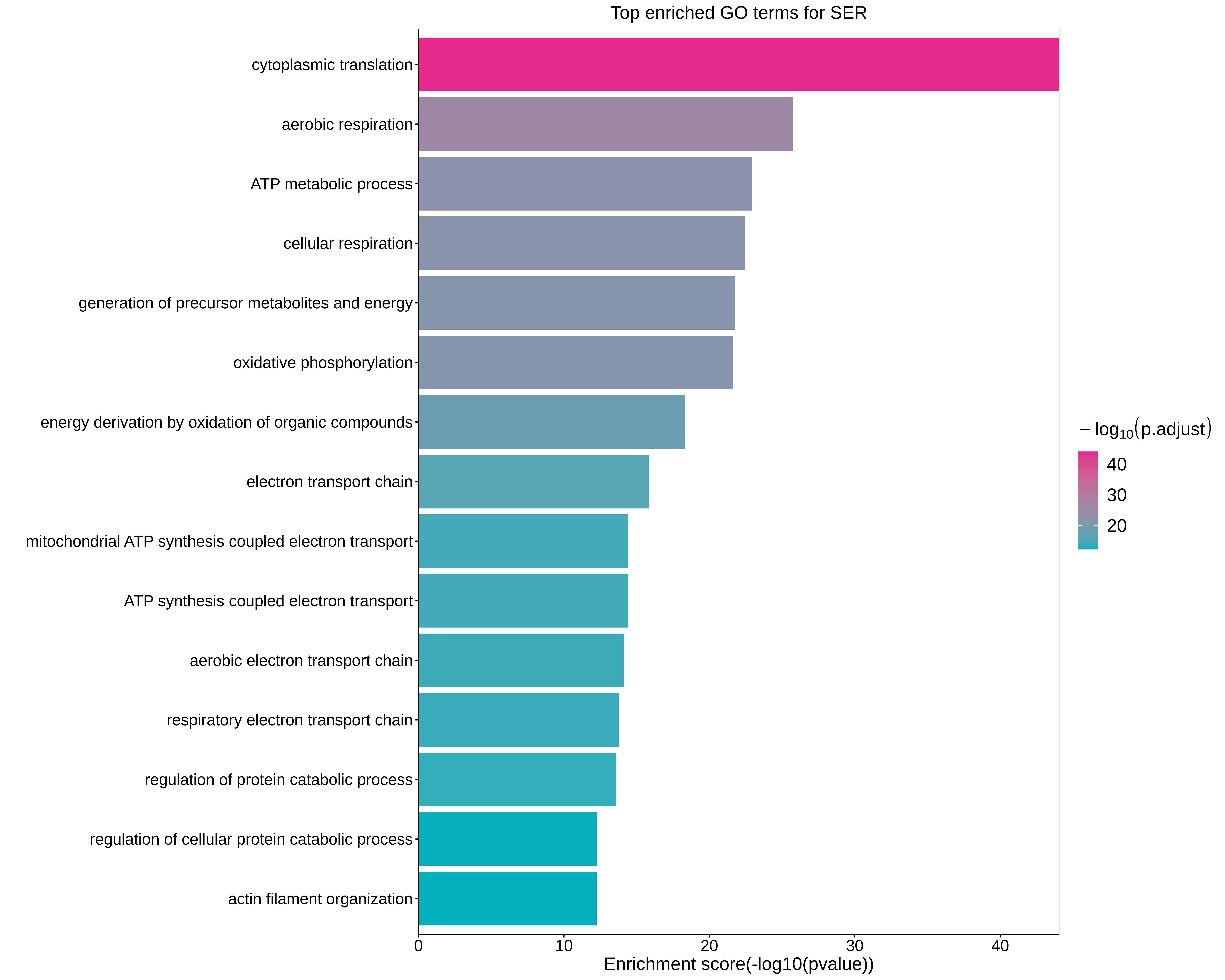

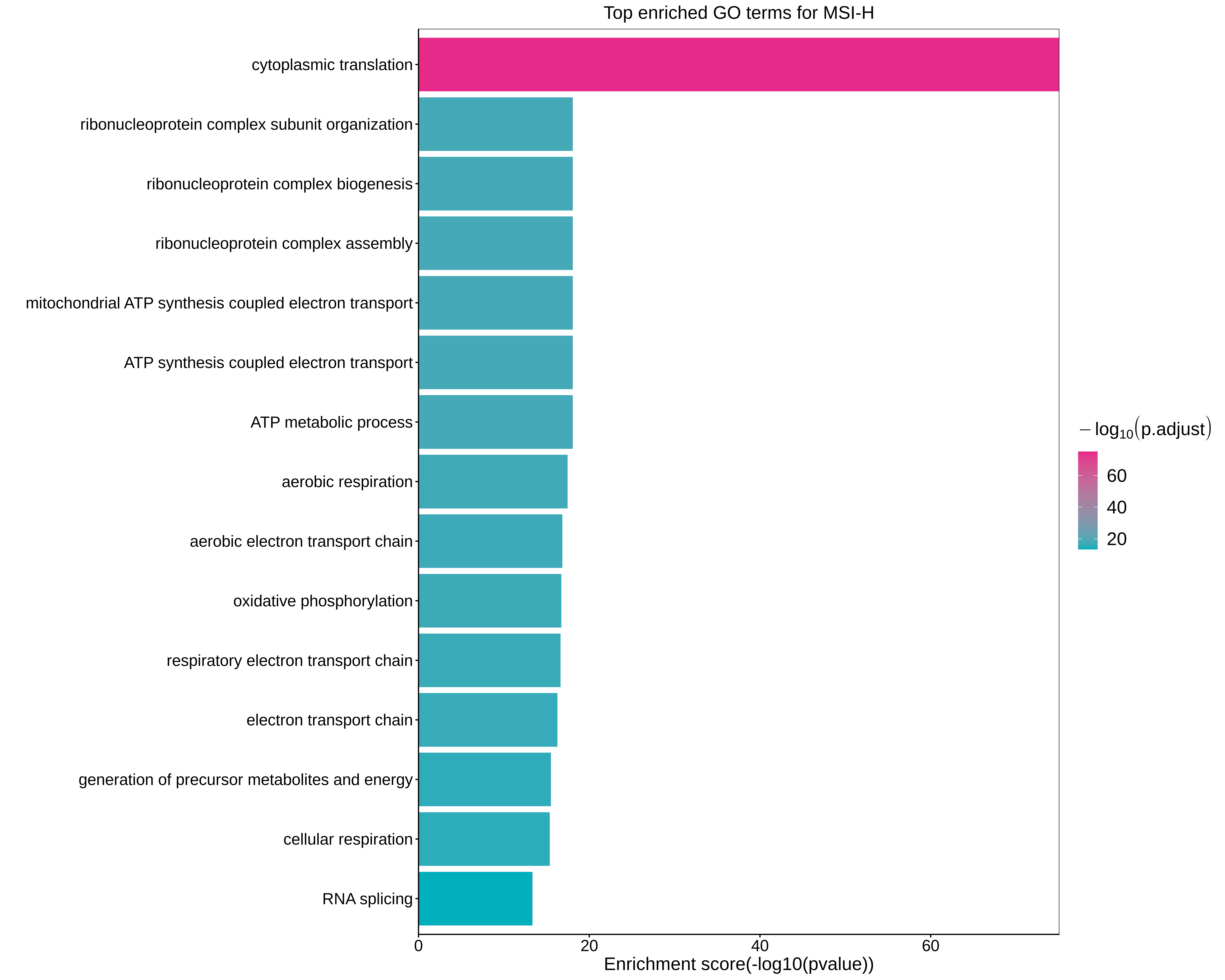
 Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states
Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states
Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states Annotation of somatic variants for genes involved in malignant transformation
Annotation of somatic variants for genes involved in malignant transformation Identification of chemicals and drugs interact with genes involved in malignant transfromation
Identification of chemicals and drugs interact with genes involved in malignant transfromation