| Tissue | Expression Dynamics | Abbreviation |
| Cervix |  | CC: Cervix cancer |
| HSIL_HPV: HPV-infected high-grade squamous intraepithelial lesions |
| N_HPV: HPV-infected normal cervix |
| Colorectum (GSE201348) |  | FAP: Familial adenomatous polyposis |
| CRC: Colorectal cancer |
| Colorectum (HTA11) |  | AD: Adenomas |
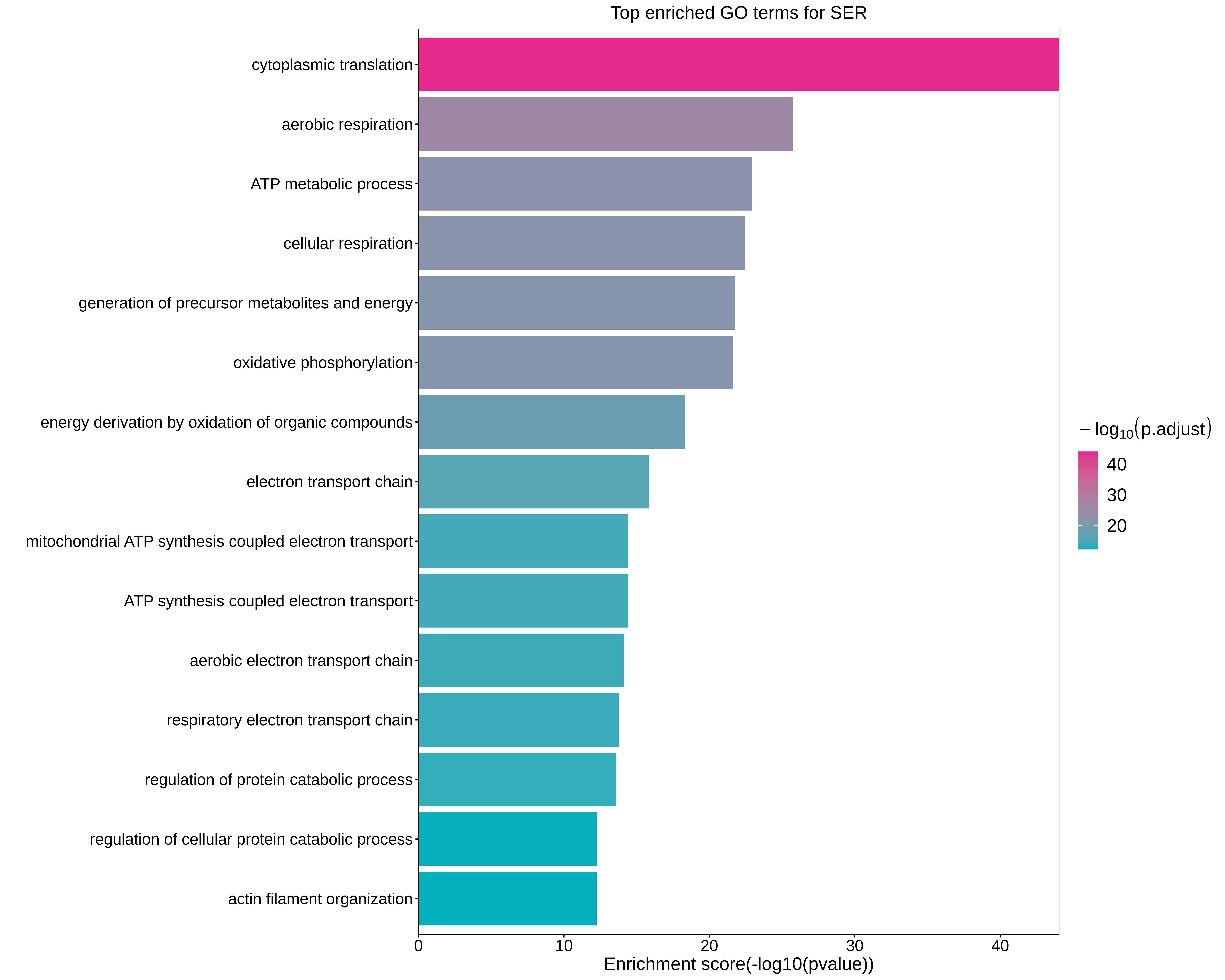
| SER: Sessile serrated lesions |
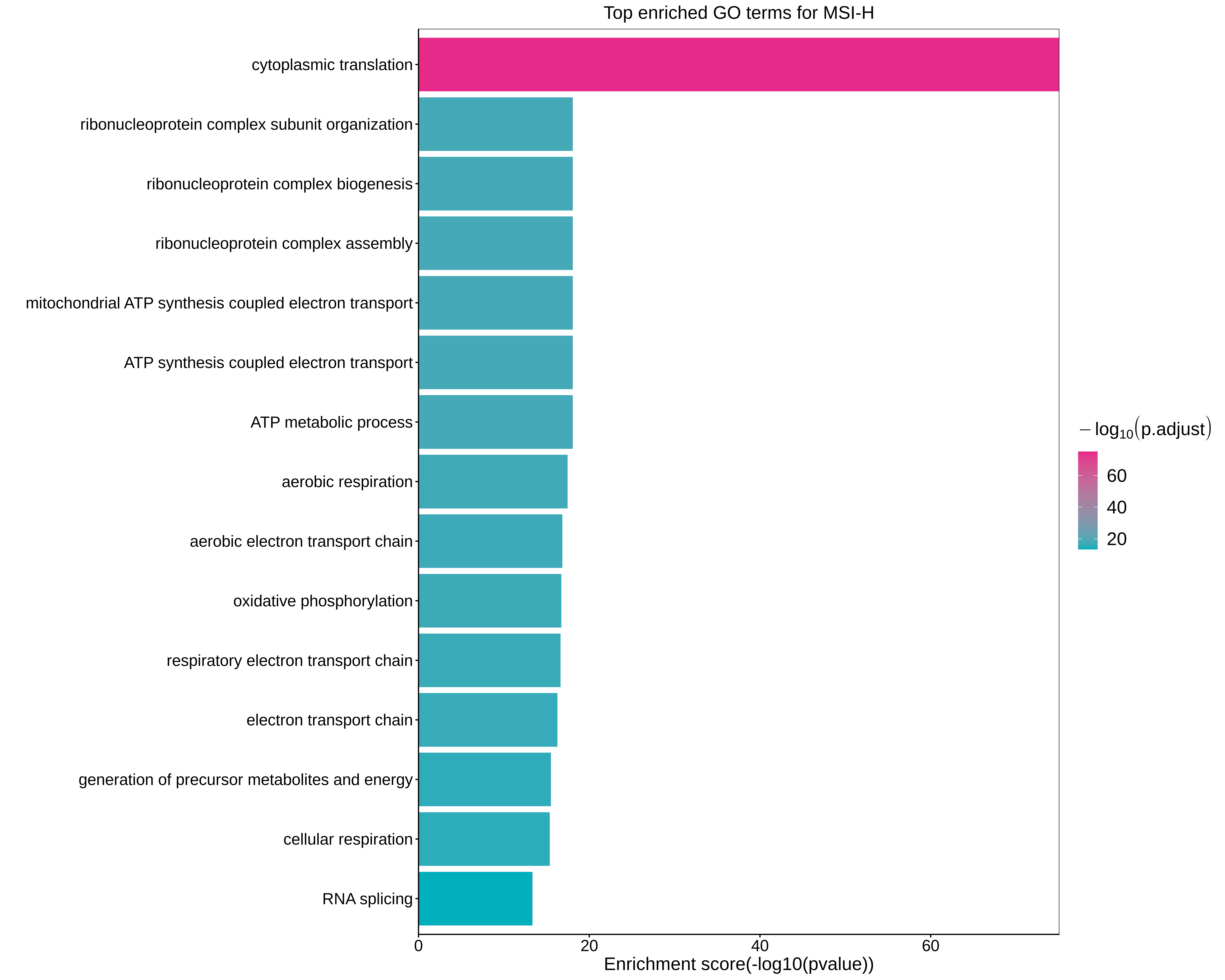
| MSI-H: Microsatellite-high colorectal cancer |
| MSS: Microsatellite stable colorectal cancer |
| Endometrium | 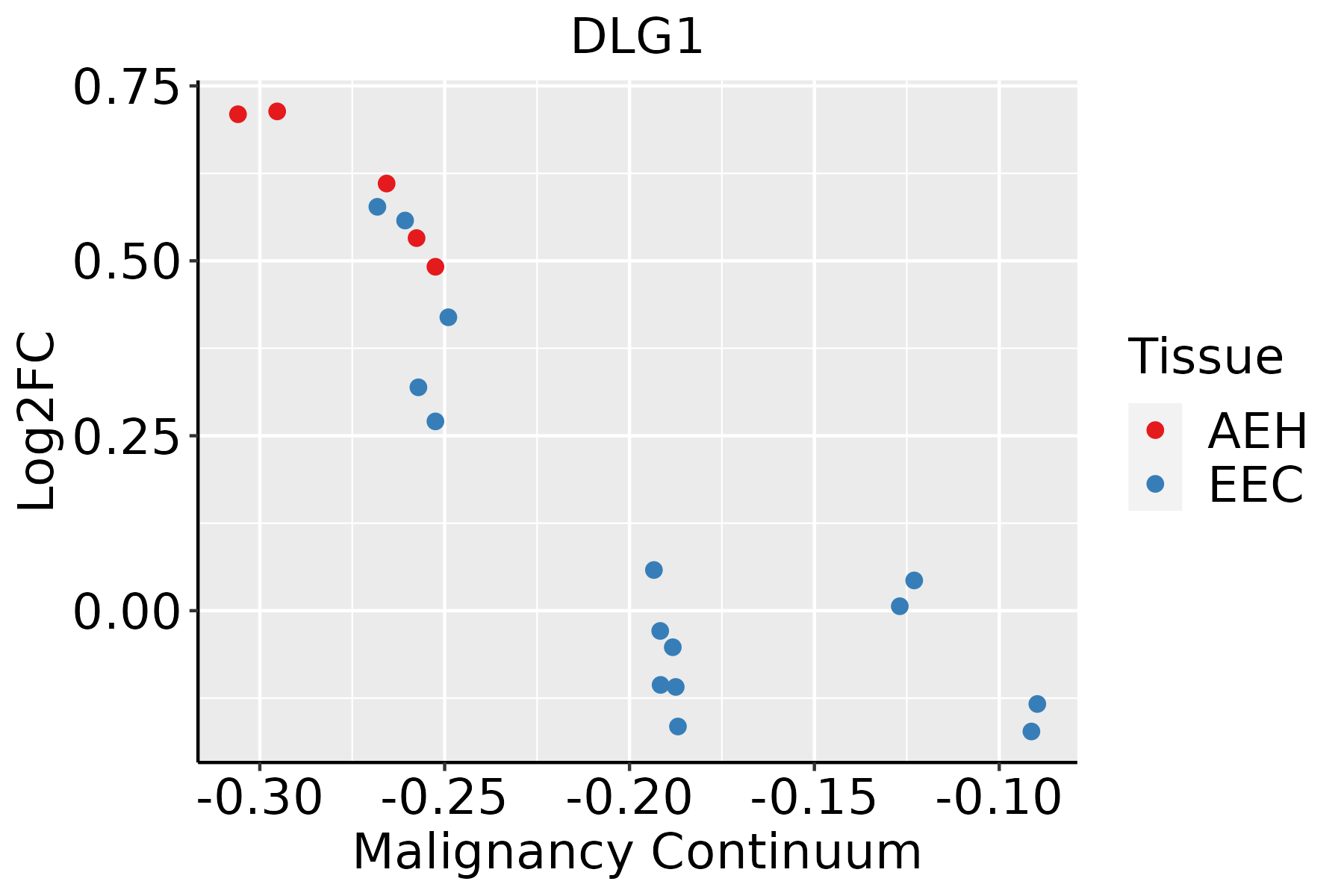 | AEH: Atypical endometrial hyperplasia |
| EEC: Endometrioid Cancer |
| Esophagus | 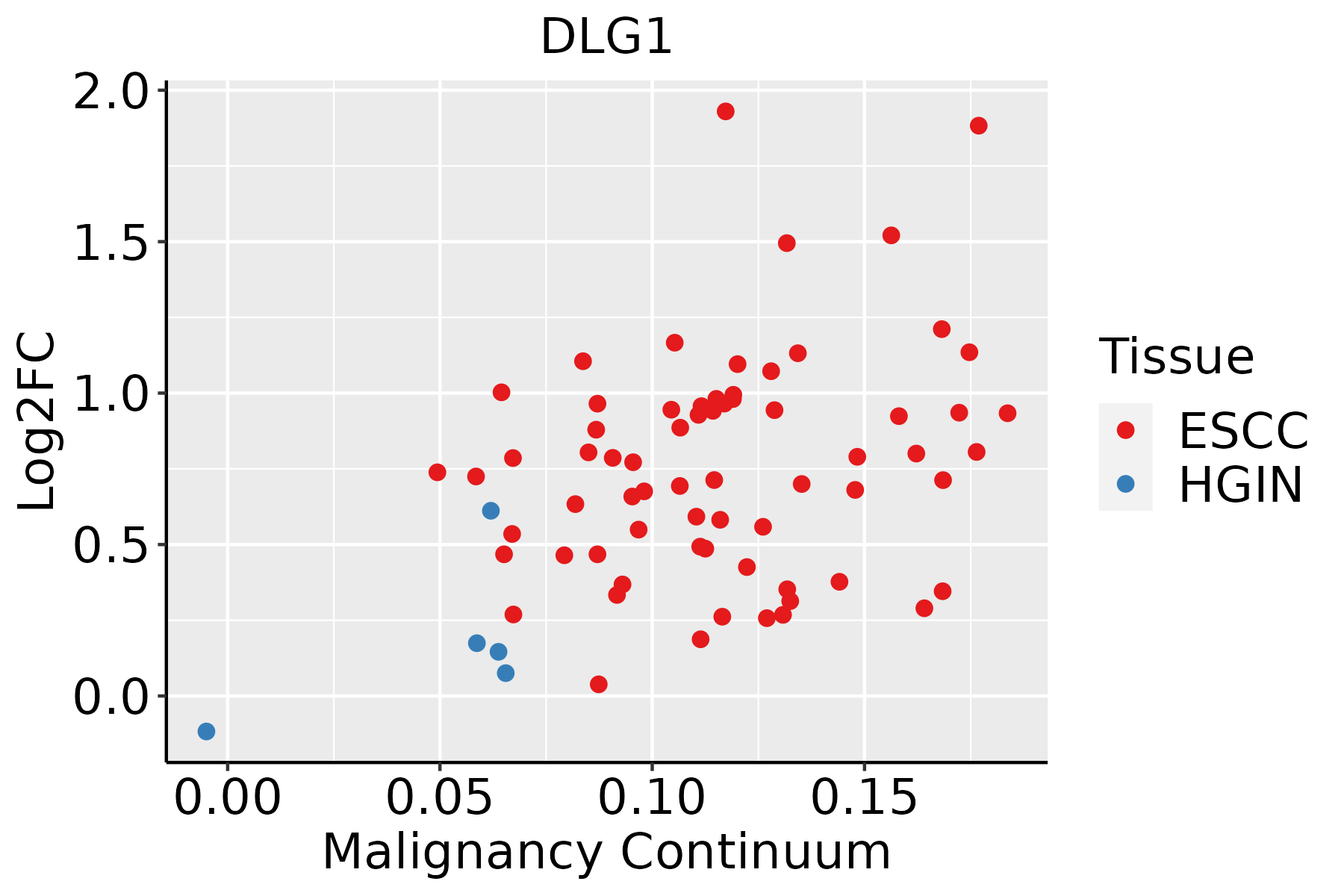 | ESCC: Esophageal squamous cell carcinoma |
| HGIN: High-grade intraepithelial neoplasias |
| LGIN: Low-grade intraepithelial neoplasias |
| GC | 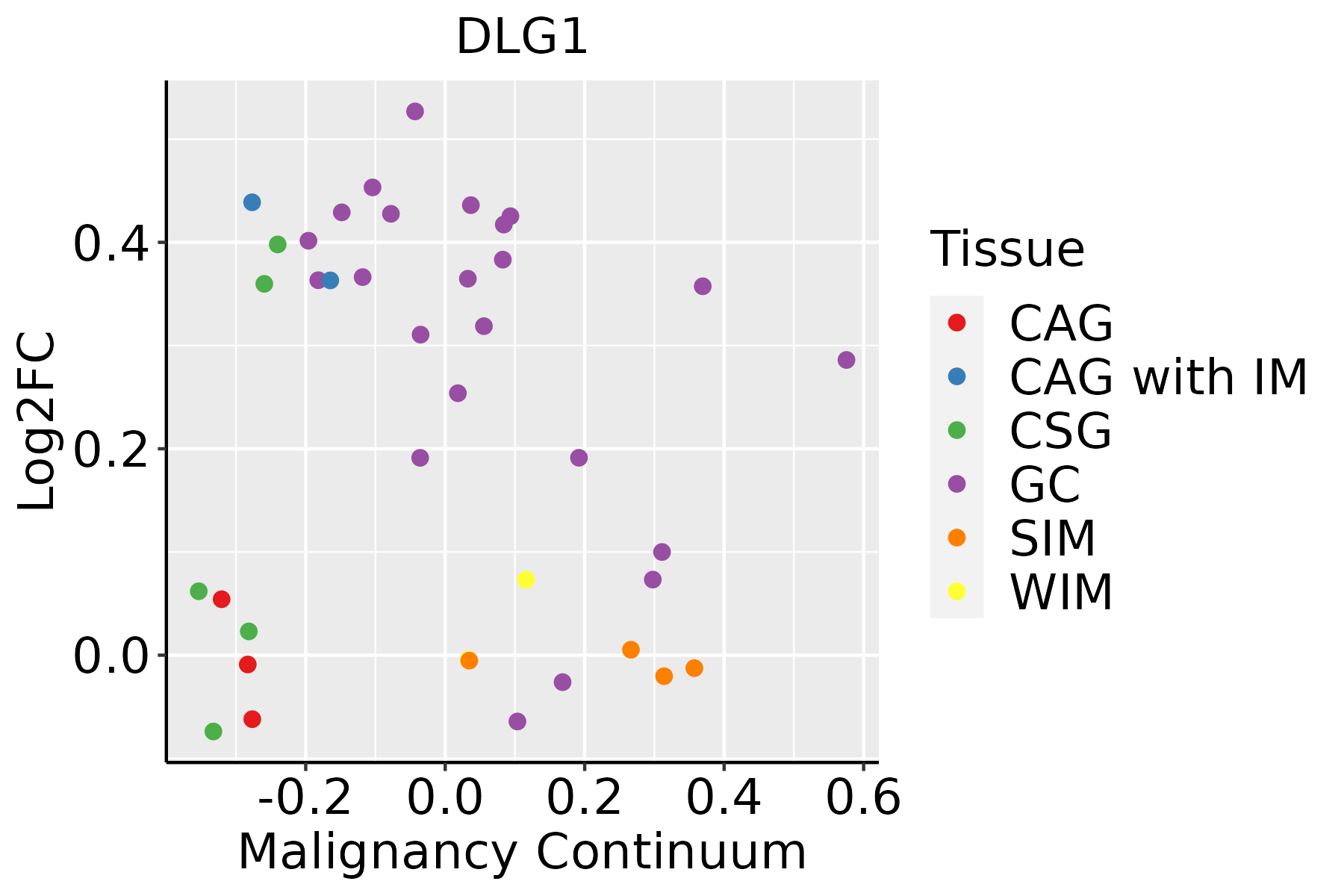 | CAG: Chronic atrophic gastritis |
| CAG with IM: Chronic atrophic gastritis with intestinal metaplasia |
| CSG: Chronic superficial gastritis |
| GC: Gastric cancer |
| SIM: Severe intestinal metaplasia |
| WIM: Wild intestinal metaplasia |
| Liver | 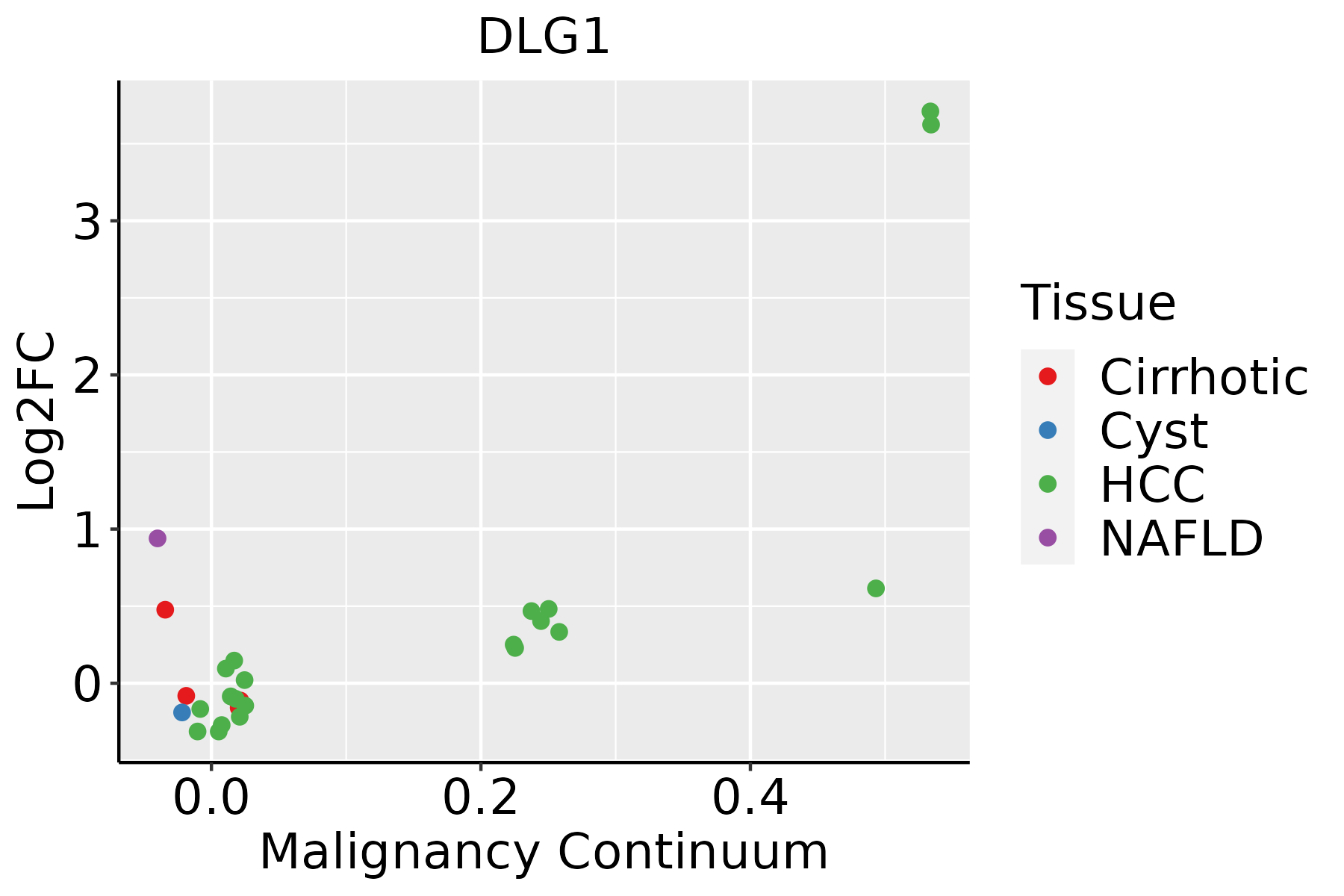 | HCC: Hepatocellular carcinoma |
| NAFLD: Non-alcoholic fatty liver disease |
| Lung | 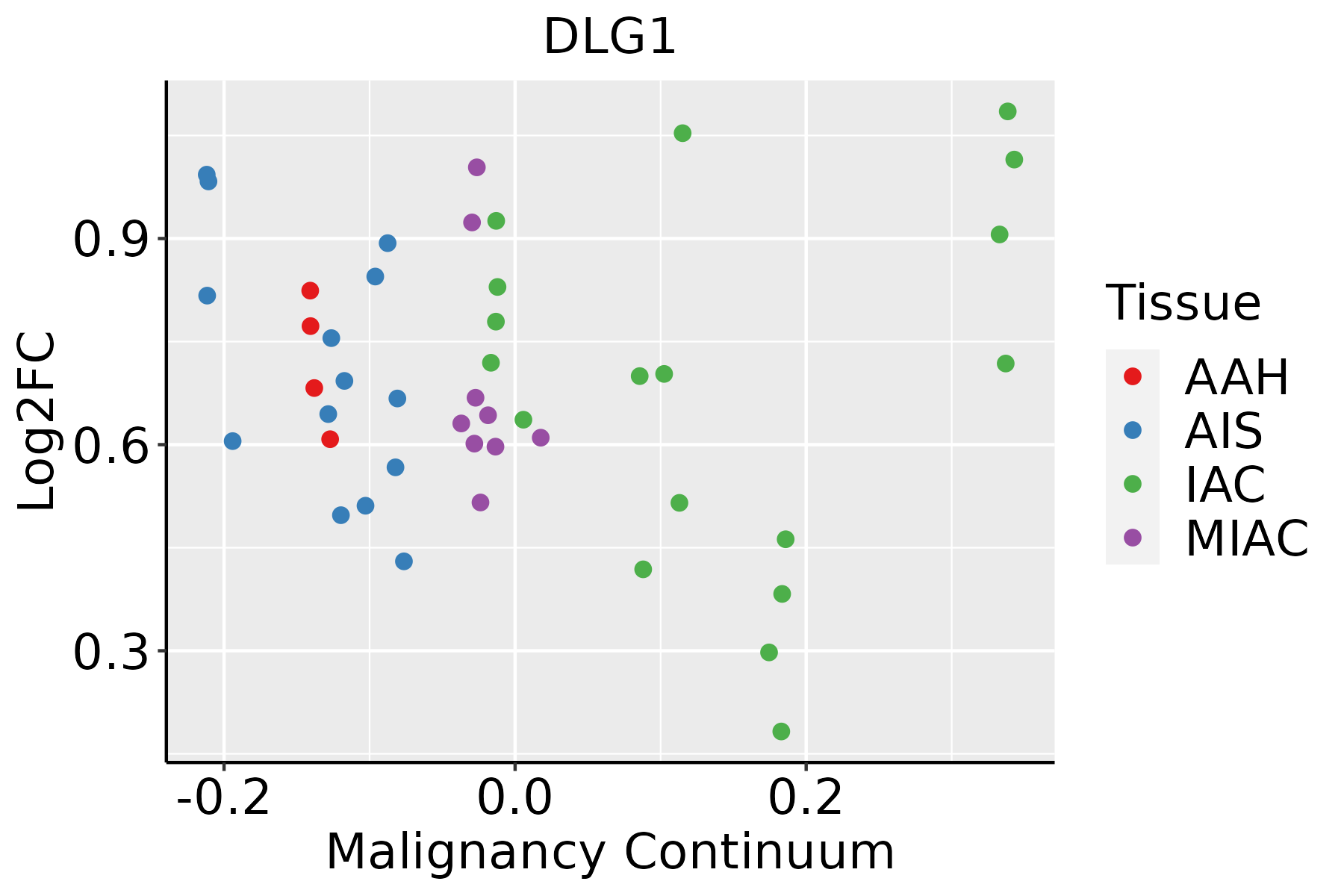 | AAH: Atypical adenomatous hyperplasia |
| AIS: Adenocarcinoma in situ |
| IAC: Invasive lung adenocarcinoma |
| MIA: Minimally invasive adenocarcinoma |
| Oral Cavity | 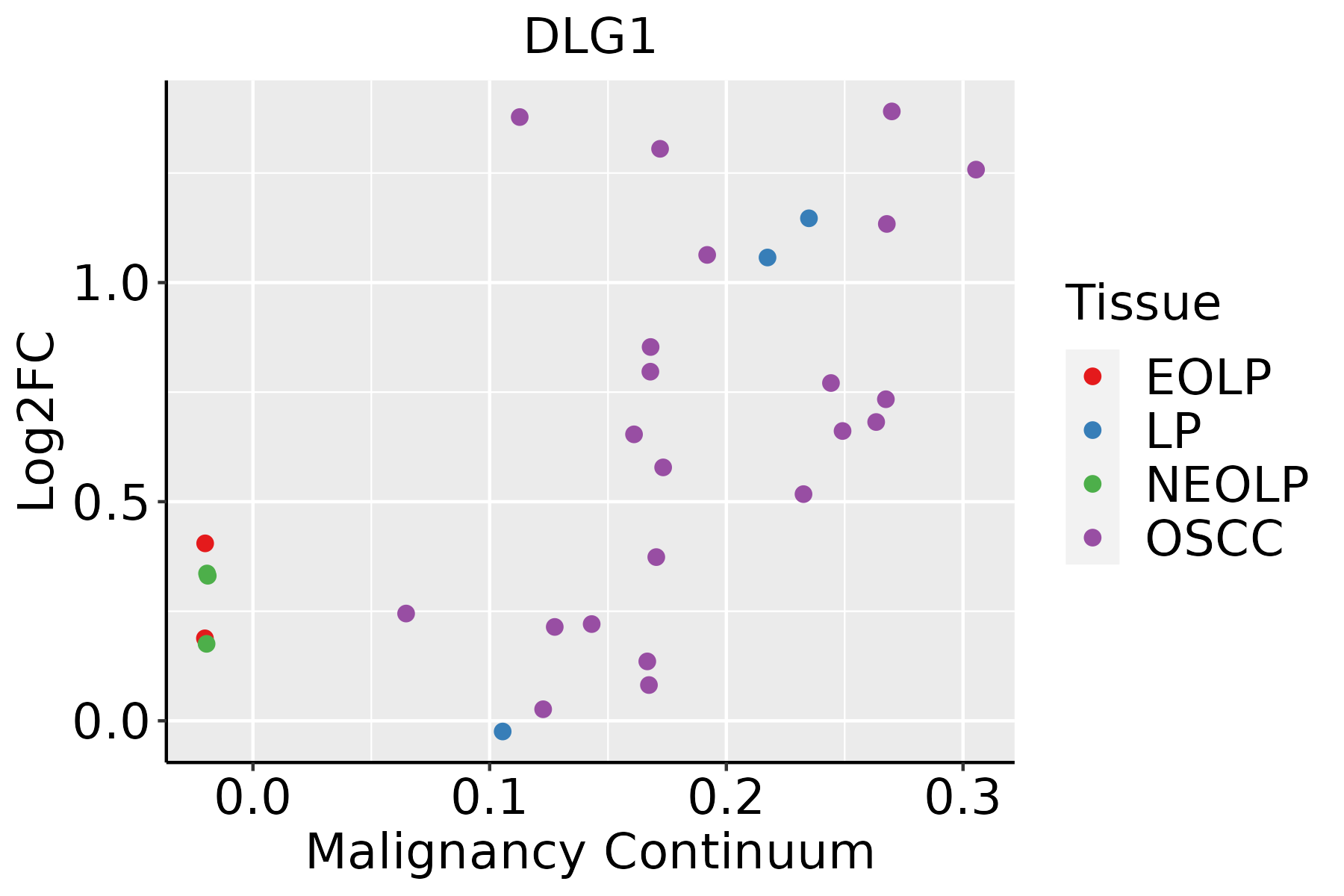 | EOLP: Erosive Oral lichen planus |
| LP: leukoplakia |
| NEOLP: Non-erosive oral lichen planus |
| OSCC: Oral squamous cell carcinoma |
| Prostate | 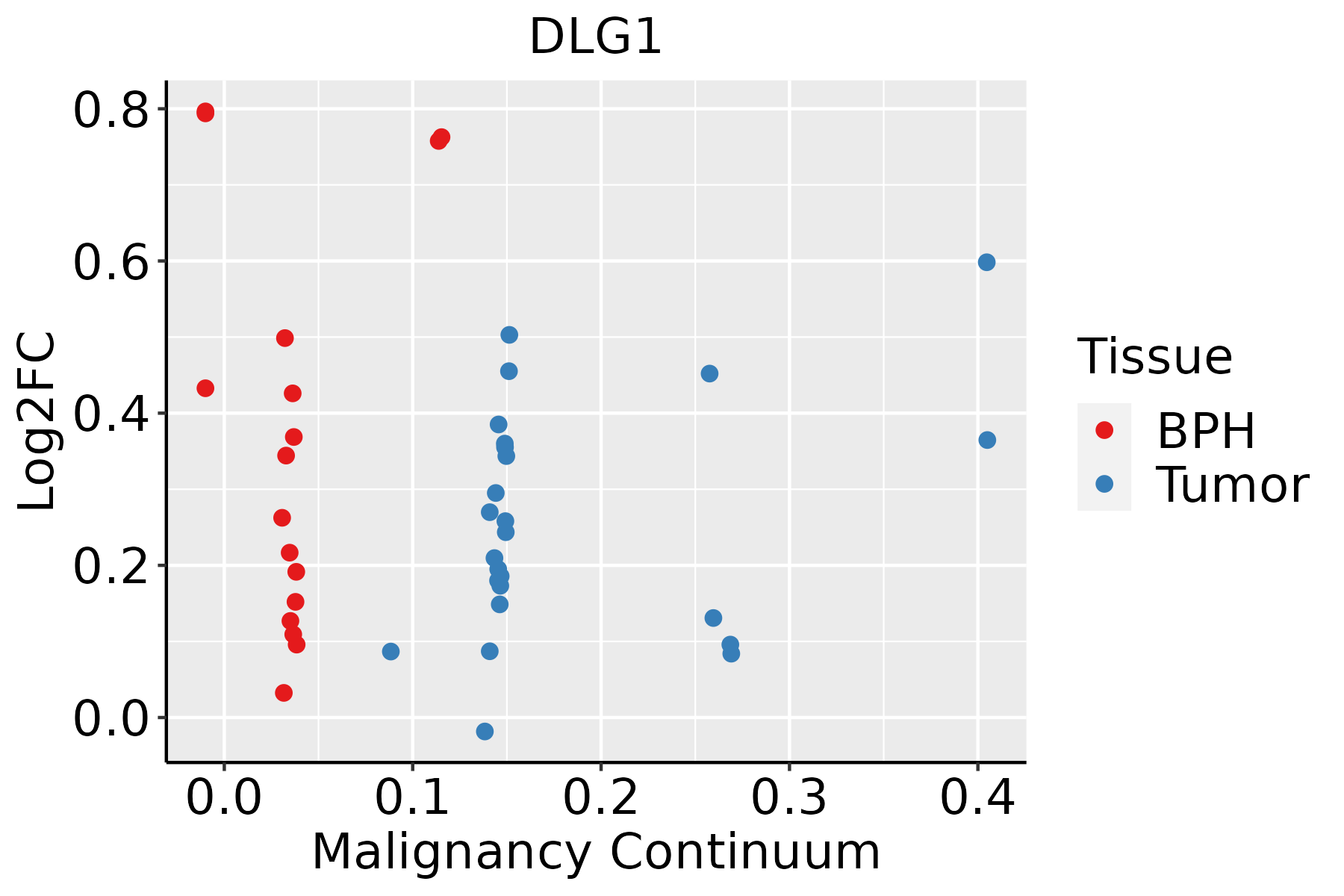 | BPH: Benign Prostatic Hyperplasia |
| Skin | 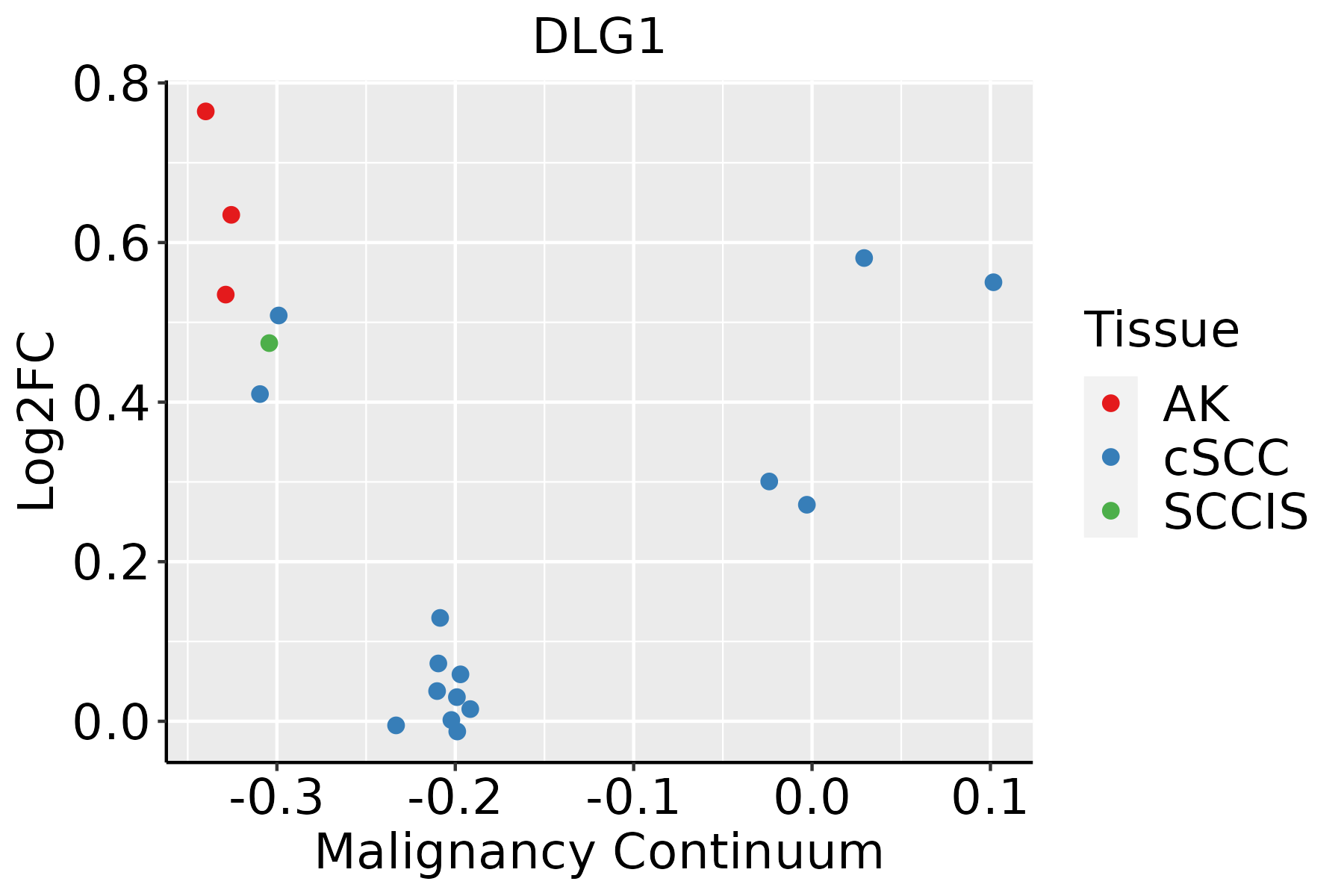 | AK: Actinic keratosis |
| cSCC: Cutaneous squamous cell carcinoma |
| SCCIS:squamous cell carcinoma in situ |
| Thyroid | 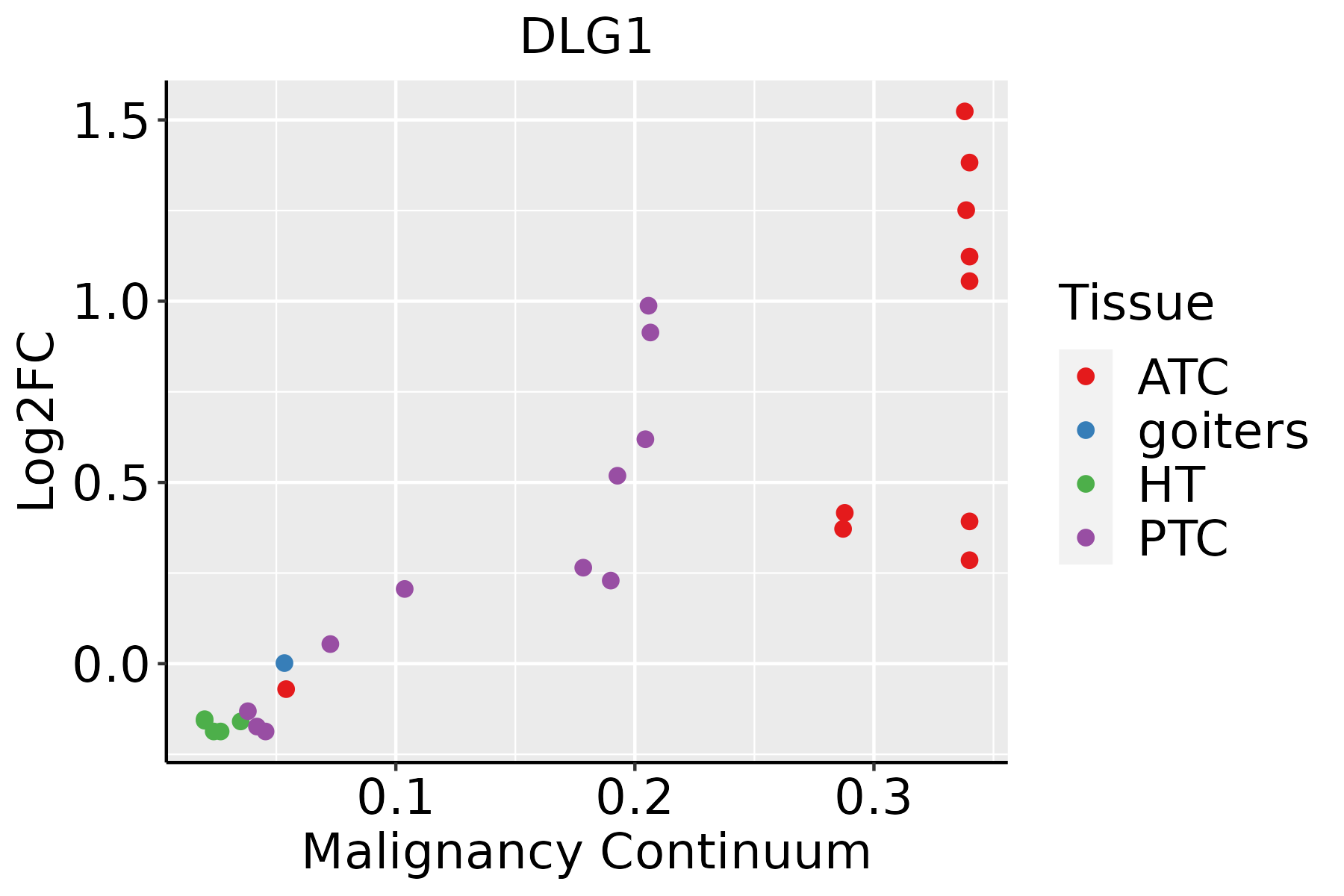 | ATC: Anaplastic thyroid cancer |
| HT: Hashimoto's thyroiditis |
| PTC: Papillary thyroid cancer |
| GO ID | Tissue | Disease Stage | Description | Gene Ratio | Bg Ratio | pvalue | p.adjust | Count |
| GO:007030216 | Oral cavity | LP | regulation of stress-activated protein kinase signaling cascade | 68/4623 | 195/18723 | 8.92e-04 | 7.64e-03 | 68 |
| GO:0110053110 | Oral cavity | LP | regulation of actin filament organization | 92/4623 | 278/18723 | 9.25e-04 | 7.85e-03 | 92 |
| GO:190437717 | Oral cavity | LP | positive regulation of protein localization to cell periphery | 29/4623 | 69/18723 | 1.15e-03 | 9.40e-03 | 29 |
| GO:0072521110 | Oral cavity | LP | purine-containing compound metabolic process | 130/4623 | 416/18723 | 1.30e-03 | 1.05e-02 | 130 |
| GO:000913218 | Oral cavity | LP | nucleoside diphosphate metabolic process | 46/4623 | 124/18723 | 1.39e-03 | 1.10e-02 | 46 |
| GO:004298211 | Oral cavity | LP | amyloid precursor protein metabolic process | 37/4623 | 95/18723 | 1.45e-03 | 1.14e-02 | 37 |
| GO:190280612 | Oral cavity | LP | regulation of cell cycle G1/S phase transition | 59/4623 | 168/18723 | 1.55e-03 | 1.20e-02 | 59 |
| GO:003287216 | Oral cavity | LP | regulation of stress-activated MAPK cascade | 66/4623 | 192/18723 | 1.60e-03 | 1.23e-02 | 66 |
| GO:190199112 | Oral cavity | LP | negative regulation of mitotic cell cycle phase transition | 62/4623 | 179/18723 | 1.76e-03 | 1.34e-02 | 62 |
| GO:001094811 | Oral cavity | LP | negative regulation of cell cycle process | 95/4623 | 294/18723 | 1.80e-03 | 1.37e-02 | 95 |
| GO:200004512 | Oral cavity | LP | regulation of G1/S transition of mitotic cell cycle | 51/4623 | 142/18723 | 1.81e-03 | 1.37e-02 | 51 |
| GO:190307816 | Oral cavity | LP | positive regulation of protein localization to plasma membrane | 26/4623 | 62/18723 | 2.11e-03 | 1.54e-02 | 26 |
| GO:0009259110 | Oral cavity | LP | ribonucleotide metabolic process | 120/4623 | 385/18723 | 2.15e-03 | 1.57e-02 | 120 |
| GO:003287316 | Oral cavity | LP | negative regulation of stress-activated MAPK cascade | 22/4623 | 51/18723 | 2.96e-03 | 2.03e-02 | 22 |
| GO:007030316 | Oral cavity | LP | negative regulation of stress-activated protein kinase signaling cascade | 22/4623 | 51/18723 | 2.96e-03 | 2.03e-02 | 22 |
| GO:003157914 | Oral cavity | LP | membrane raft organization | 13/4623 | 25/18723 | 2.98e-03 | 2.04e-02 | 13 |
| GO:0006163110 | Oral cavity | LP | purine nucleotide metabolic process | 122/4623 | 396/18723 | 3.10e-03 | 2.11e-02 | 122 |
| GO:0009150110 | Oral cavity | LP | purine ribonucleotide metabolic process | 114/4623 | 368/18723 | 3.41e-03 | 2.28e-02 | 114 |
| GO:000918519 | Oral cavity | LP | ribonucleoside diphosphate metabolic process | 39/4623 | 106/18723 | 3.64e-03 | 2.42e-02 | 39 |
| GO:004211017 | Oral cavity | LP | T cell activation | 146/4623 | 487/18723 | 4.13e-03 | 2.66e-02 | 146 |
| Pathway ID | Tissue | Disease Stage | Description | Gene Ratio | Bg Ratio | pvalue | p.adjust | qvalue | Count |
| hsa0516620 | Cervix | CC | Human T-cell leukemia virus 1 infection | 61/1267 | 222/8465 | 8.13e-07 | 7.98e-06 | 4.72e-06 | 61 |
| hsa0453020 | Cervix | CC | Tight junction | 49/1267 | 169/8465 | 1.87e-06 | 1.78e-05 | 1.05e-05 | 49 |
| hsa051657 | Cervix | CC | Human papillomavirus infection | 74/1267 | 331/8465 | 1.70e-04 | 1.02e-03 | 6.03e-04 | 74 |
| hsa043908 | Cervix | CC | Hippo signaling pathway | 40/1267 | 157/8465 | 3.64e-04 | 1.82e-03 | 1.07e-03 | 40 |
| hsa046604 | Cervix | CC | T cell receptor signaling pathway | 25/1267 | 104/8465 | 9.52e-03 | 2.94e-02 | 1.74e-02 | 25 |
| hsa05166110 | Cervix | CC | Human T-cell leukemia virus 1 infection | 61/1267 | 222/8465 | 8.13e-07 | 7.98e-06 | 4.72e-06 | 61 |
| hsa04530110 | Cervix | CC | Tight junction | 49/1267 | 169/8465 | 1.87e-06 | 1.78e-05 | 1.05e-05 | 49 |
| hsa0516512 | Cervix | CC | Human papillomavirus infection | 74/1267 | 331/8465 | 1.70e-04 | 1.02e-03 | 6.03e-04 | 74 |
| hsa0439013 | Cervix | CC | Hippo signaling pathway | 40/1267 | 157/8465 | 3.64e-04 | 1.82e-03 | 1.07e-03 | 40 |
| hsa0466011 | Cervix | CC | T cell receptor signaling pathway | 25/1267 | 104/8465 | 9.52e-03 | 2.94e-02 | 1.74e-02 | 25 |
| hsa04530 | Colorectum | AD | Tight junction | 76/2092 | 169/8465 | 5.49e-09 | 9.69e-08 | 6.18e-08 | 76 |
| hsa05166 | Colorectum | AD | Human T-cell leukemia virus 1 infection | 72/2092 | 222/8465 | 5.24e-03 | 2.44e-02 | 1.55e-02 | 72 |
| hsa045301 | Colorectum | AD | Tight junction | 76/2092 | 169/8465 | 5.49e-09 | 9.69e-08 | 6.18e-08 | 76 |
| hsa051661 | Colorectum | AD | Human T-cell leukemia virus 1 infection | 72/2092 | 222/8465 | 5.24e-03 | 2.44e-02 | 1.55e-02 | 72 |
| hsa045302 | Colorectum | SER | Tight junction | 59/1580 | 169/8465 | 3.24e-07 | 5.98e-06 | 4.34e-06 | 59 |
| hsa045303 | Colorectum | SER | Tight junction | 59/1580 | 169/8465 | 3.24e-07 | 5.98e-06 | 4.34e-06 | 59 |
| hsa045304 | Colorectum | MSS | Tight junction | 66/1875 | 169/8465 | 4.10e-07 | 6.25e-06 | 3.83e-06 | 66 |
| hsa051662 | Colorectum | MSS | Human T-cell leukemia virus 1 infection | 68/1875 | 222/8465 | 1.84e-03 | 9.61e-03 | 5.89e-03 | 68 |
| hsa04390 | Colorectum | MSS | Hippo signaling pathway | 48/1875 | 157/8465 | 8.32e-03 | 3.10e-02 | 1.90e-02 | 48 |
| hsa045305 | Colorectum | MSS | Tight junction | 66/1875 | 169/8465 | 4.10e-07 | 6.25e-06 | 3.83e-06 | 66 |
| Hugo Symbol | Variant Class | Variant Classification | dbSNP RS | HGVSc | HGVSp | HGVSp Short | SWISSPROT | BIOTYPE | SIFT | PolyPhen | Tumor Sample Barcode | Tissue | Histology | Sex | Age | Stage | Therapy Types | Drugs | Outcome |
| DLG1 | SNV | Missense_Mutation | novel | c.301N>A | p.Leu101Ile | p.L101I | Q12959 | protein_coding | tolerated_low_confidence(0.17) | possibly_damaging(0.689) | TCGA-A2-A4S1-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| DLG1 | SNV | Missense_Mutation | | c.731N>C | p.Asn244Thr | p.N244T | Q12959 | protein_coding | deleterious(0.03) | probably_damaging(0.999) | TCGA-A8-A083-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| DLG1 | SNV | Missense_Mutation | novel | c.1434N>G | p.Phe478Leu | p.F478L | Q12959 | protein_coding | deleterious(0.01) | probably_damaging(0.999) | TCGA-AC-A5XS-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Hormone Therapy | femara | SD |
| DLG1 | SNV | Missense_Mutation | | c.16N>G | p.Gln6Glu | p.Q6E | Q12959 | protein_coding | tolerated(0.38) | benign(0) | TCGA-AN-A0FW-01 | Breast | breast invasive carcinoma | Female | >=65 | III/IV | Unknown | Unknown | SD |
| DLG1 | SNV | Missense_Mutation | | c.1186N>T | p.Asp396Tyr | p.D396Y | Q12959 | protein_coding | deleterious(0) | probably_damaging(0.987) | TCGA-BH-A0AZ-01 | Breast | breast invasive carcinoma | Female | <65 | III/IV | Chemotherapy | doxorubicin | CR |
| DLG1 | SNV | Missense_Mutation | novel | c.2516N>C | p.Arg839Thr | p.R839T | Q12959 | protein_coding | deleterious(0) | probably_damaging(0.999) | TCGA-BH-A209-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| DLG1 | SNV | Missense_Mutation | | c.2251G>T | p.Asp751Tyr | p.D751Y | Q12959 | protein_coding | deleterious(0) | probably_damaging(0.978) | TCGA-C8-A26Y-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| DLG1 | SNV | Missense_Mutation | | c.1692G>A | p.Met564Ile | p.M564I | Q12959 | protein_coding | tolerated(0.07) | benign(0.406) | TCGA-D8-A1JJ-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Chemotherapy | doxorubicine | SD |
| DLG1 | SNV | Missense_Mutation | novel | c.2013N>A | p.Phe671Leu | p.F671L | Q12959 | protein_coding | tolerated(0.92) | benign(0) | TCGA-E2-A1LG-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Chemotherapy | doxorubicin | SD |
| DLG1 | SNV | Missense_Mutation | novel | c.2432N>G | p.Tyr811Cys | p.Y811C | Q12959 | protein_coding | deleterious(0) | probably_damaging(1) | TCGA-LL-A740-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Chemotherapy | adriamycin | CR |







 Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells
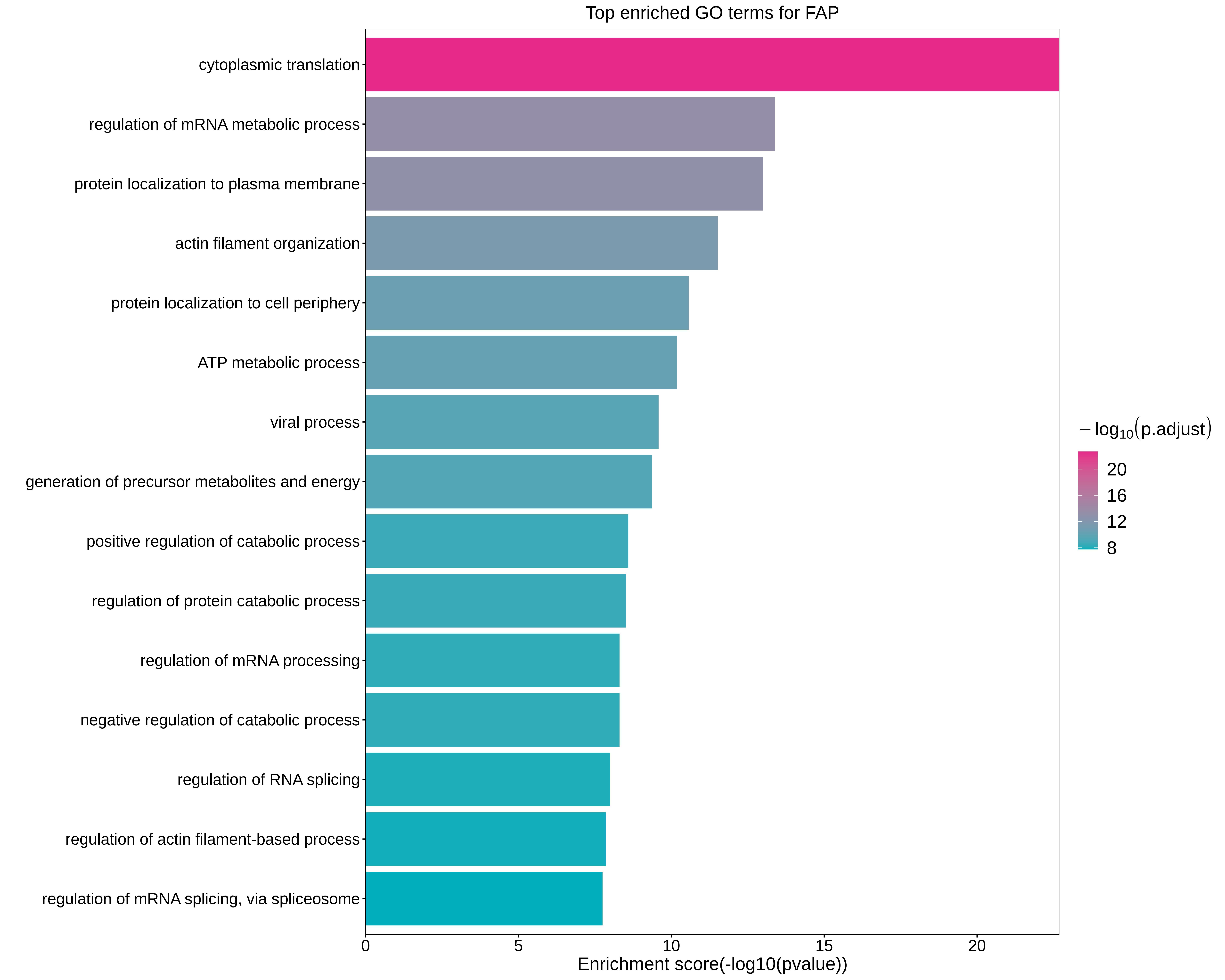
Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer
Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer

 Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states
Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states
Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states Annotation of somatic variants for genes involved in malignant transformation
Annotation of somatic variants for genes involved in malignant transformation Identification of chemicals and drugs interact with genes involved in malignant transfromation
Identification of chemicals and drugs interact with genes involved in malignant transfromation











