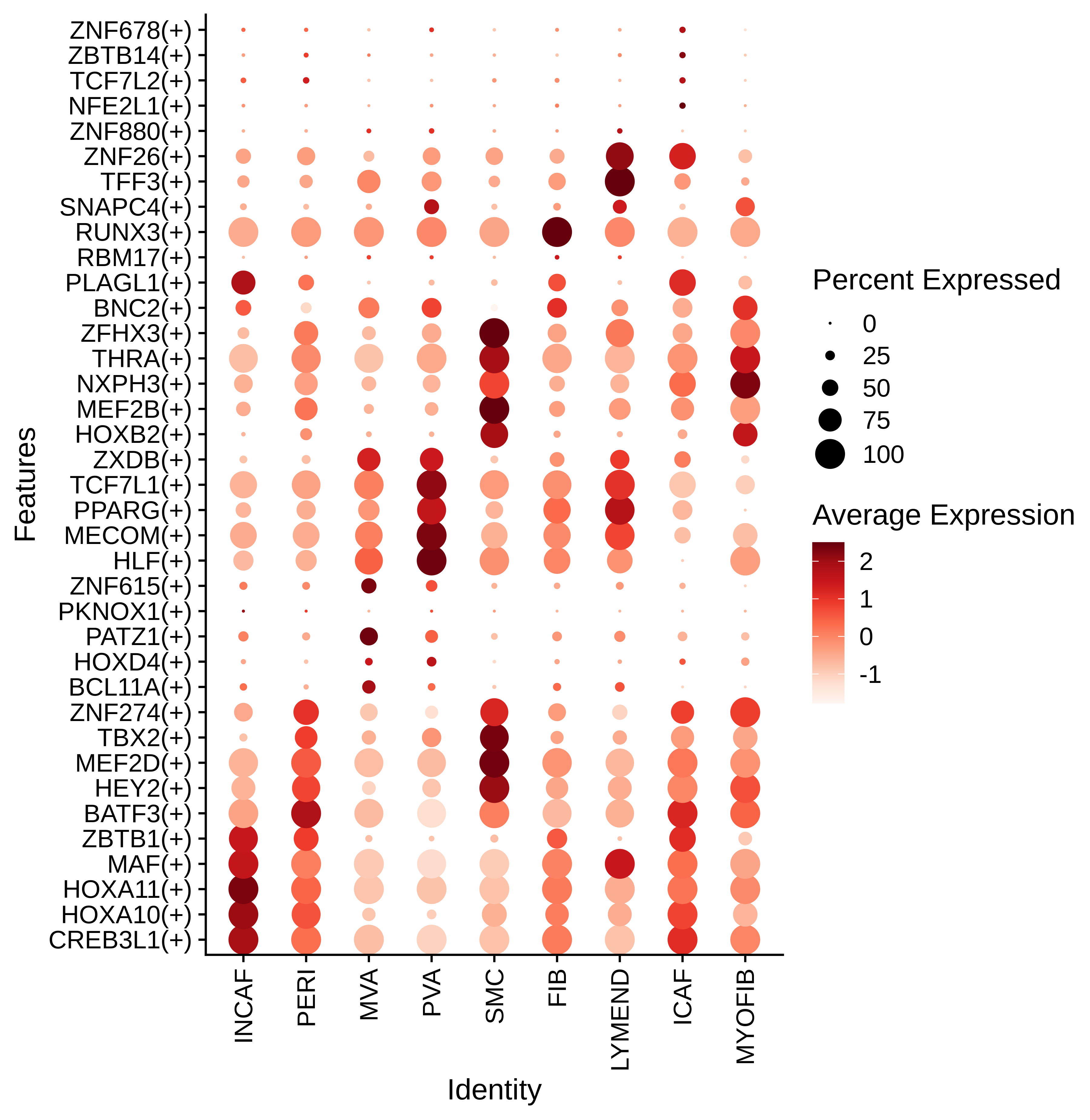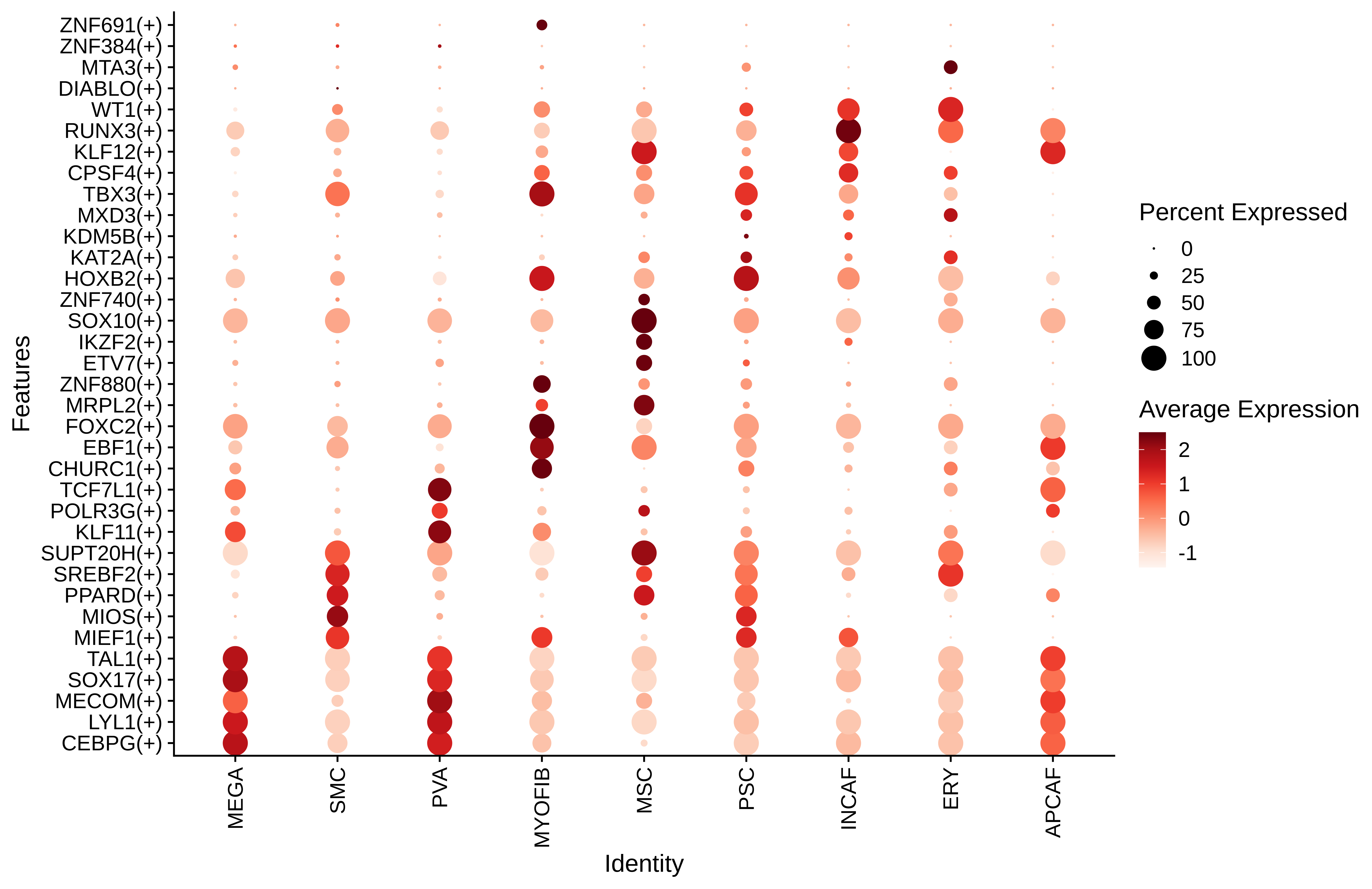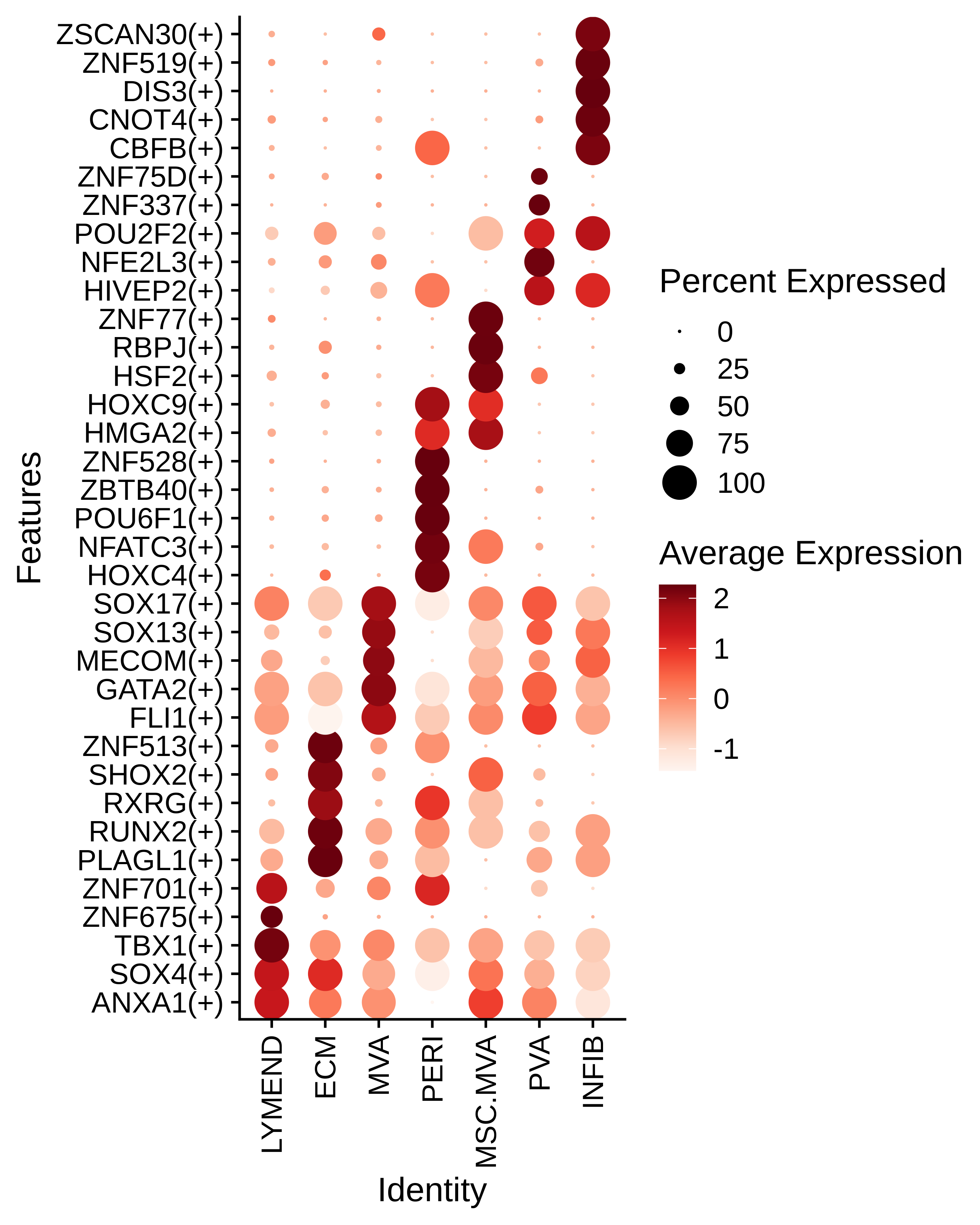| Tissue | Expression Dynamics | Abbreviation |
| Cervix | 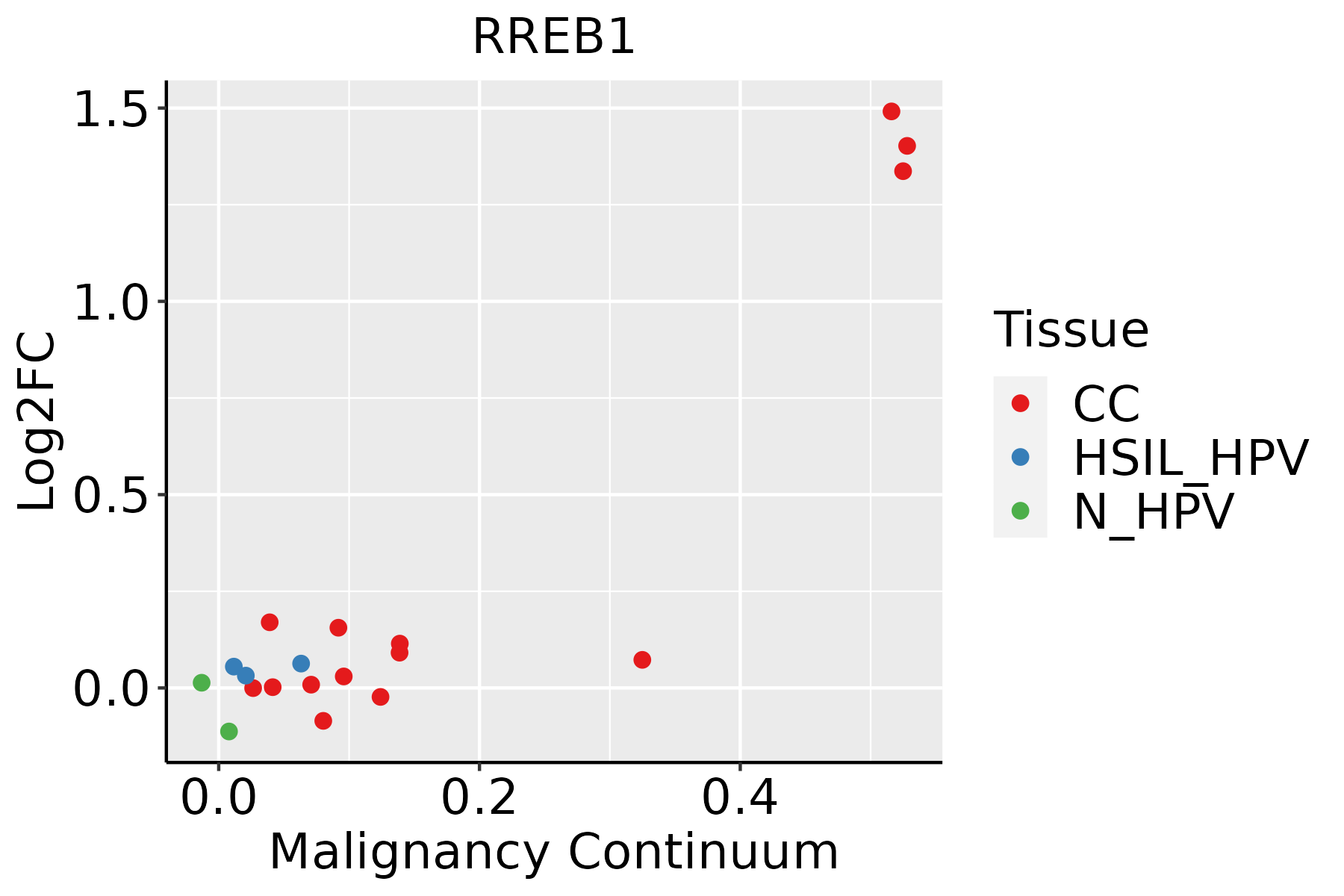 | CC: Cervix cancer |
| HSIL_HPV: HPV-infected high-grade squamous intraepithelial lesions |
| N_HPV: HPV-infected normal cervix |
| Colorectum (GSE201348) |  | FAP: Familial adenomatous polyposis |
| CRC: Colorectal cancer |
| Colorectum (HTA11) |  | AD: Adenomas |
| SER: Sessile serrated lesions |
| MSI-H: Microsatellite-high colorectal cancer |
| MSS: Microsatellite stable colorectal cancer |
| Esophagus |  | ESCC: Esophageal squamous cell carcinoma |
| HGIN: High-grade intraepithelial neoplasias |
| LGIN: Low-grade intraepithelial neoplasias |
| Oral Cavity |  | EOLP: Erosive Oral lichen planus |
| LP: leukoplakia |
| NEOLP: Non-erosive oral lichen planus |
| OSCC: Oral squamous cell carcinoma |
| Skin | 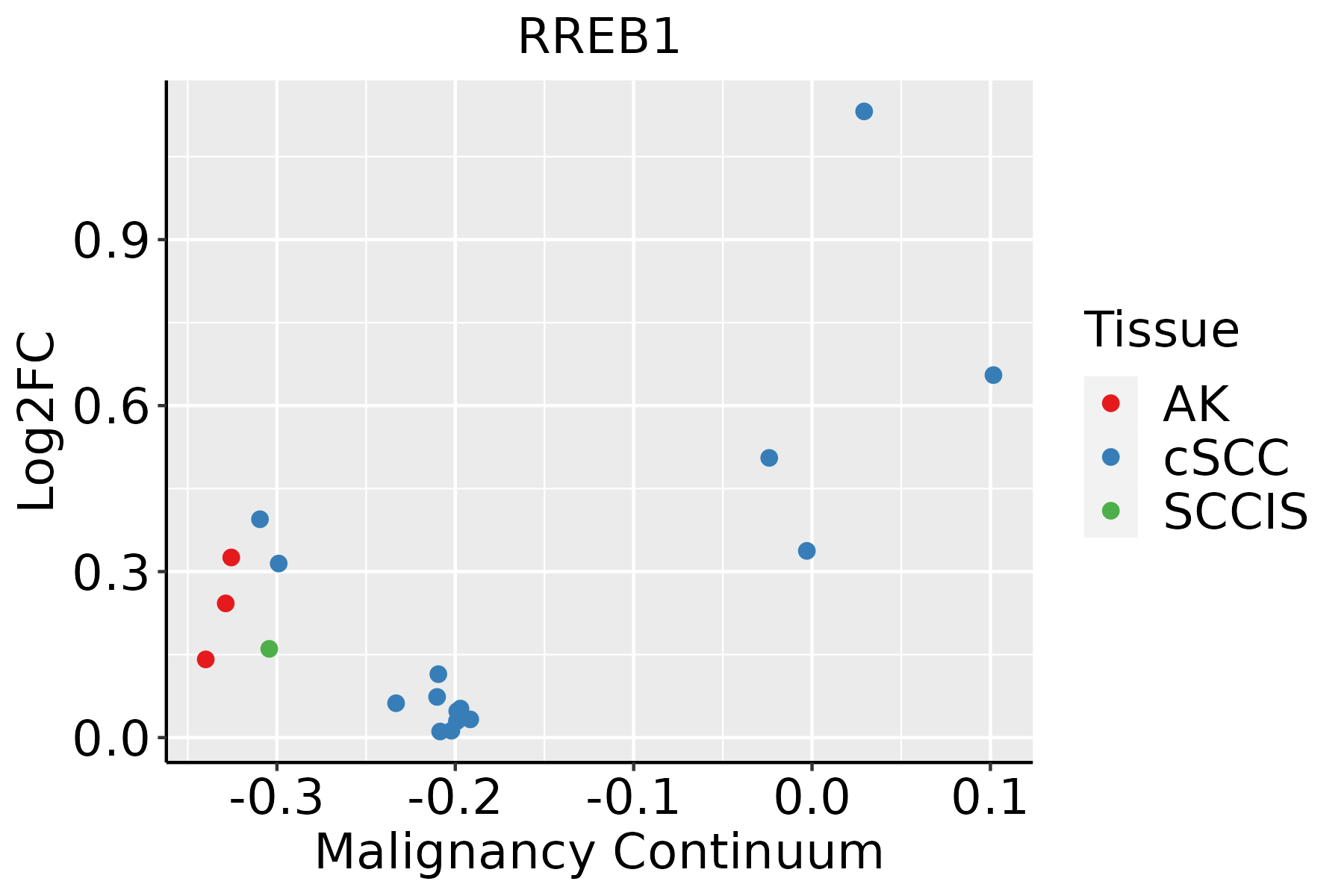 | AK: Actinic keratosis |
| cSCC: Cutaneous squamous cell carcinoma |
| SCCIS:squamous cell carcinoma in situ |
| Thyroid | 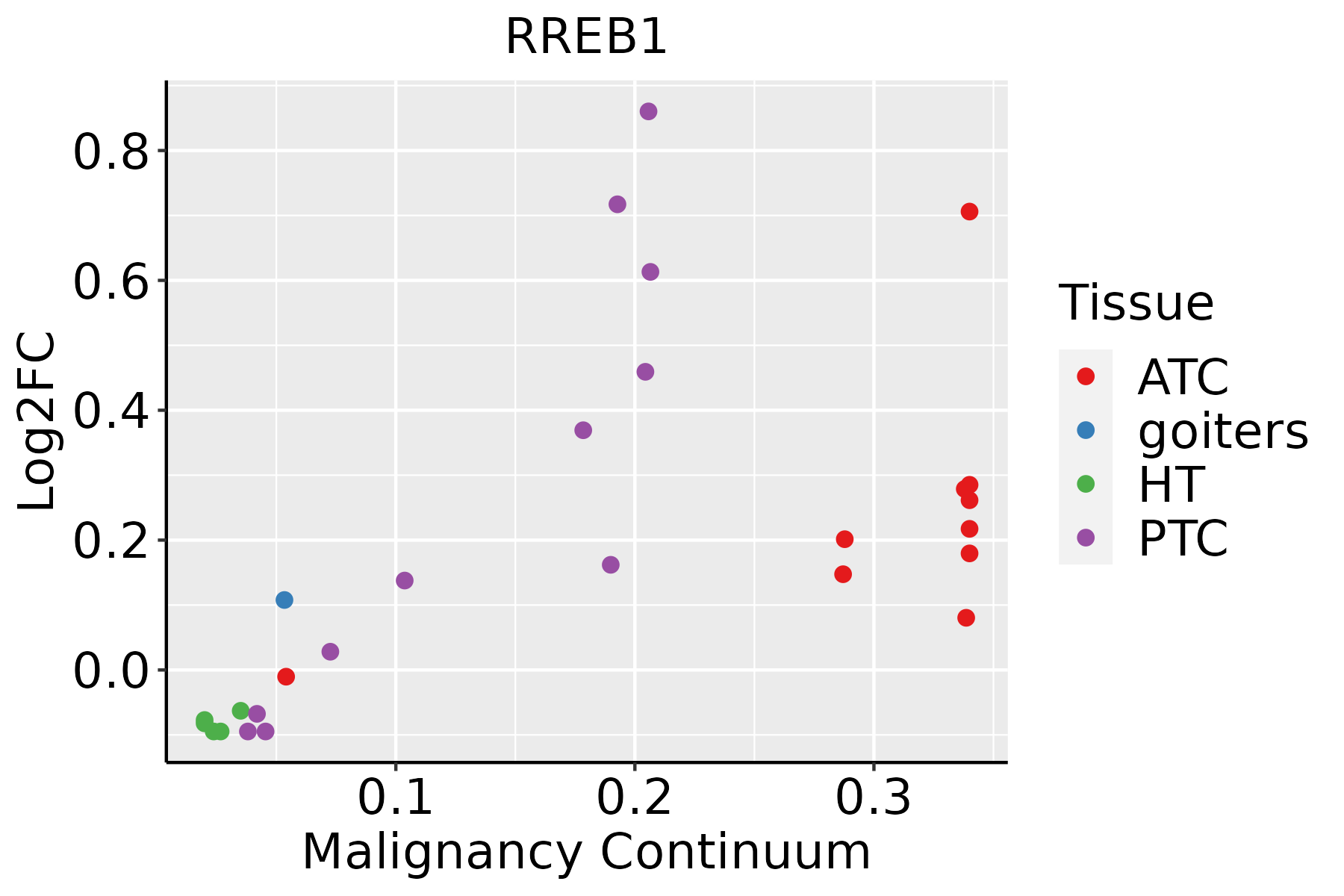 | ATC: Anaplastic thyroid cancer |
| HT: Hashimoto's thyroiditis |
| PTC: Papillary thyroid cancer |
| GO ID | Tissue | Disease Stage | Description | Gene Ratio | Bg Ratio | pvalue | p.adjust | Count |
| GO:00016673 | Colorectum | FAP | ameboidal-type cell migration | 100/2622 | 475/18723 | 1.42e-05 | 3.79e-04 | 100 |
| GO:19027454 | Colorectum | FAP | positive regulation of lamellipodium organization | 16/2622 | 37/18723 | 1.44e-05 | 3.80e-04 | 16 |
| GO:19027434 | Colorectum | FAP | regulation of lamellipodium organization | 20/2622 | 54/18723 | 2.09e-05 | 5.05e-04 | 20 |
| GO:00457854 | Colorectum | FAP | positive regulation of cell adhesion | 92/2622 | 437/18723 | 3.09e-05 | 7.04e-04 | 92 |
| GO:00106343 | Colorectum | FAP | positive regulation of epithelial cell migration | 44/2622 | 176/18723 | 7.02e-05 | 1.31e-03 | 44 |
| GO:00901324 | Colorectum | FAP | epithelium migration | 77/2622 | 360/18723 | 7.70e-05 | 1.38e-03 | 77 |
| GO:00107694 | Colorectum | FAP | regulation of cell morphogenesis involved in differentiation | 28/2622 | 96/18723 | 8.70e-05 | 1.55e-03 | 28 |
| GO:00106314 | Colorectum | FAP | epithelial cell migration | 76/2622 | 357/18723 | 1.00e-04 | 1.69e-03 | 76 |
| GO:00901304 | Colorectum | FAP | tissue migration | 77/2622 | 365/18723 | 1.23e-04 | 2.00e-03 | 77 |
| GO:00344464 | Colorectum | FAP | substrate adhesion-dependent cell spreading | 30/2622 | 108/18723 | 1.32e-04 | 2.12e-03 | 30 |
| GO:00106324 | Colorectum | FAP | regulation of epithelial cell migration | 64/2622 | 292/18723 | 1.45e-04 | 2.26e-03 | 64 |
| GO:00610411 | Colorectum | FAP | regulation of wound healing | 35/2622 | 134/18723 | 1.48e-04 | 2.29e-03 | 35 |
| GO:19000244 | Colorectum | FAP | regulation of substrate adhesion-dependent cell spreading | 19/2622 | 57/18723 | 1.73e-04 | 2.58e-03 | 19 |
| GO:19030341 | Colorectum | FAP | regulation of response to wounding | 41/2622 | 167/18723 | 1.86e-04 | 2.74e-03 | 41 |
| GO:00107704 | Colorectum | FAP | positive regulation of cell morphogenesis involved in differentiation | 23/2622 | 79/18723 | 3.70e-04 | 4.63e-03 | 23 |
| GO:19000264 | Colorectum | FAP | positive regulation of substrate adhesion-dependent cell spreading | 14/2622 | 41/18723 | 9.20e-04 | 9.24e-03 | 14 |
| GO:00308793 | Colorectum | FAP | mammary gland development | 33/2622 | 137/18723 | 1.07e-03 | 1.04e-02 | 33 |
| GO:00506732 | Colorectum | FAP | epithelial cell proliferation | 84/2622 | 437/18723 | 1.39e-03 | 1.25e-02 | 84 |
| GO:0090303 | Colorectum | FAP | positive regulation of wound healing | 17/2622 | 59/18723 | 2.33e-03 | 1.87e-02 | 17 |
| GO:00506781 | Colorectum | FAP | regulation of epithelial cell proliferation | 71/2622 | 381/18723 | 6.67e-03 | 4.12e-02 | 71 |
| Hugo Symbol | Variant Class | Variant Classification | dbSNP RS | HGVSc | HGVSp | HGVSp Short | SWISSPROT | BIOTYPE | SIFT | PolyPhen | Tumor Sample Barcode | Tissue | Histology | Sex | Age | Stage | Therapy Types | Drugs | Outcome |
| RREB1 | SNV | Missense_Mutation | novel | c.4799N>C | p.Cys1600Ser | p.C1600S | Q92766 | protein_coding | deleterious(0) | probably_damaging(0.98) | TCGA-5L-AAT0-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Hormone Therapy | tamoxiphen | SD |
| RREB1 | SNV | Missense_Mutation | novel | c.953G>T | p.Cys318Phe | p.C318F | Q92766 | protein_coding | deleterious(0) | probably_damaging(0.999) | TCGA-5T-A9QA-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Chemotherapy | taxol | SD |
| RREB1 | SNV | Missense_Mutation | novel | c.4672G>A | p.Val1558Met | p.V1558M | Q92766 | protein_coding | tolerated(0.13) | benign(0.023) | TCGA-A2-A0CT-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Chemotherapy | cytoxan | SD |
| RREB1 | SNV | Missense_Mutation | novel | c.220N>G | p.Ile74Val | p.I74V | Q92766 | protein_coding | tolerated(0.42) | benign(0.001) | TCGA-A8-A06P-01 | Breast | breast invasive carcinoma | Female | <65 | III/IV | Unspecific | | SD |
| RREB1 | SNV | Missense_Mutation | | c.4658N>G | p.Asp1553Gly | p.D1553G | Q92766 | protein_coding | tolerated(0.12) | benign(0.42) | TCGA-A8-A09Z-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |
| RREB1 | SNV | Missense_Mutation | | c.694G>A | p.Asp232Asn | p.D232N | Q92766 | protein_coding | tolerated(0.07) | possibly_damaging(0.892) | TCGA-AN-A0FY-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Unknown | Unknown | SD |
| RREB1 | SNV | Missense_Mutation | | c.1636C>T | p.Pro546Ser | p.P546S | Q92766 | protein_coding | tolerated(0.11) | benign(0.037) | TCGA-B6-A0RE-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Unknown | Unknown | SD |
| RREB1 | SNV | Missense_Mutation | | c.601N>A | p.Glu201Lys | p.E201K | Q92766 | protein_coding | tolerated(0.16) | benign(0.234) | TCGA-C8-A138-01 | Breast | breast invasive carcinoma | Female | <65 | III/IV | Unknown | Unknown | SD |
| RREB1 | SNV | Missense_Mutation | | c.4854N>C | p.Gln1618His | p.Q1618H | Q92766 | protein_coding | deleterious(0) | probably_damaging(0.994) | TCGA-C8-A27B-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Chemotherapy | 5-fluorouracil | CR |
| RREB1 | SNV | Missense_Mutation | | c.2088N>C | p.Glu696Asp | p.E696D | Q92766 | protein_coding | deleterious(0) | probably_damaging(0.984) | TCGA-D8-A1XQ-01 | Breast | breast invasive carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD |







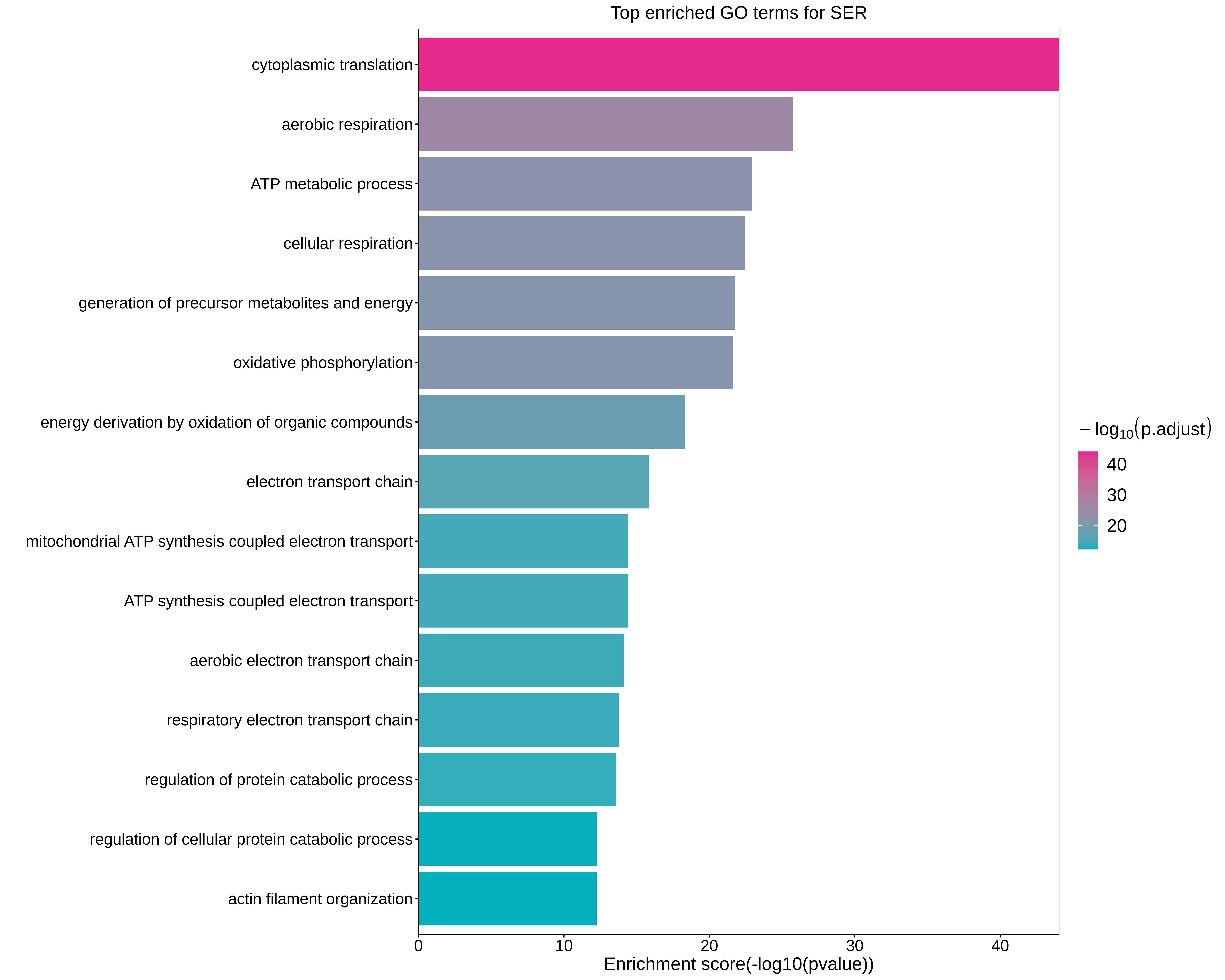
 Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells
Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer
Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer

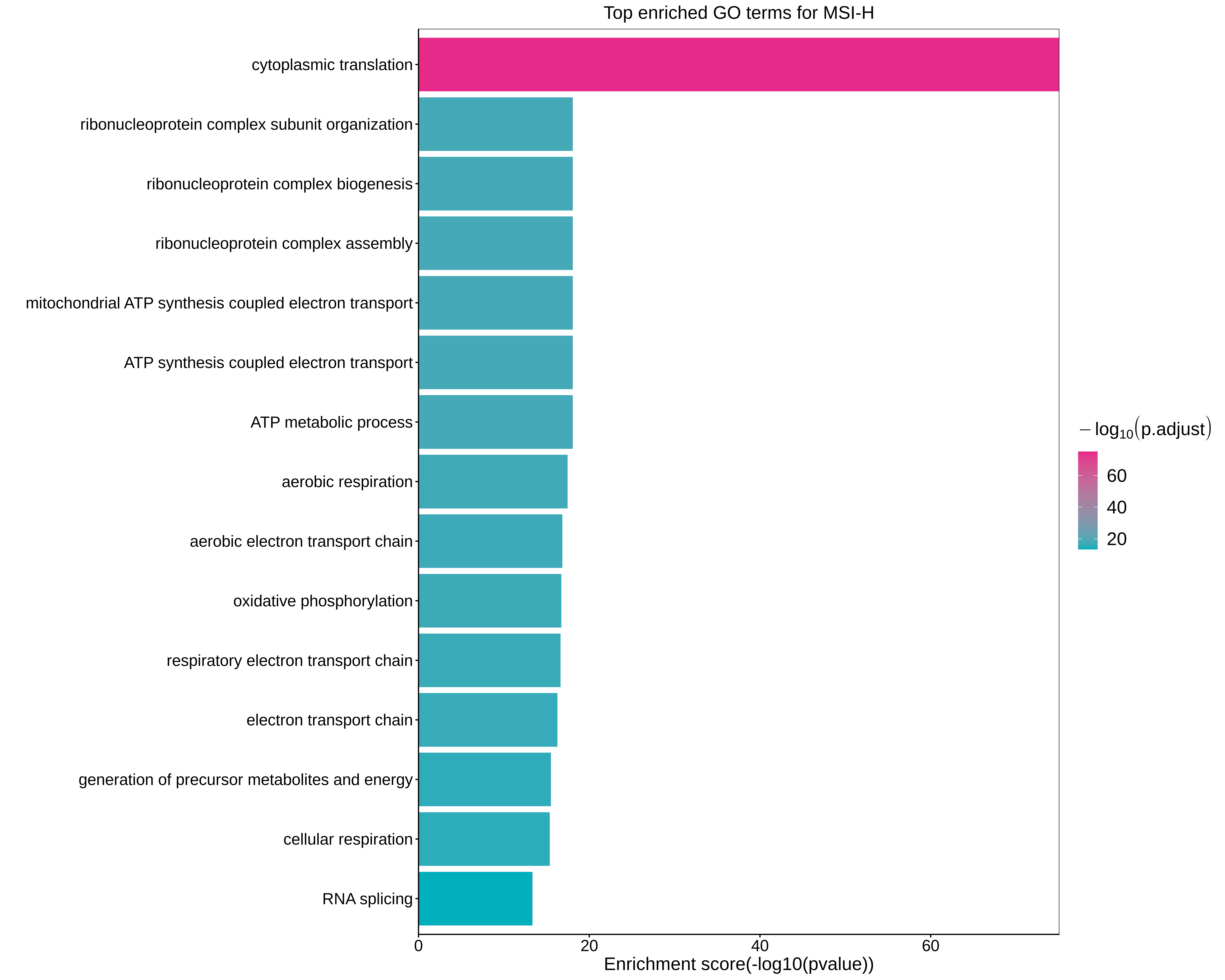
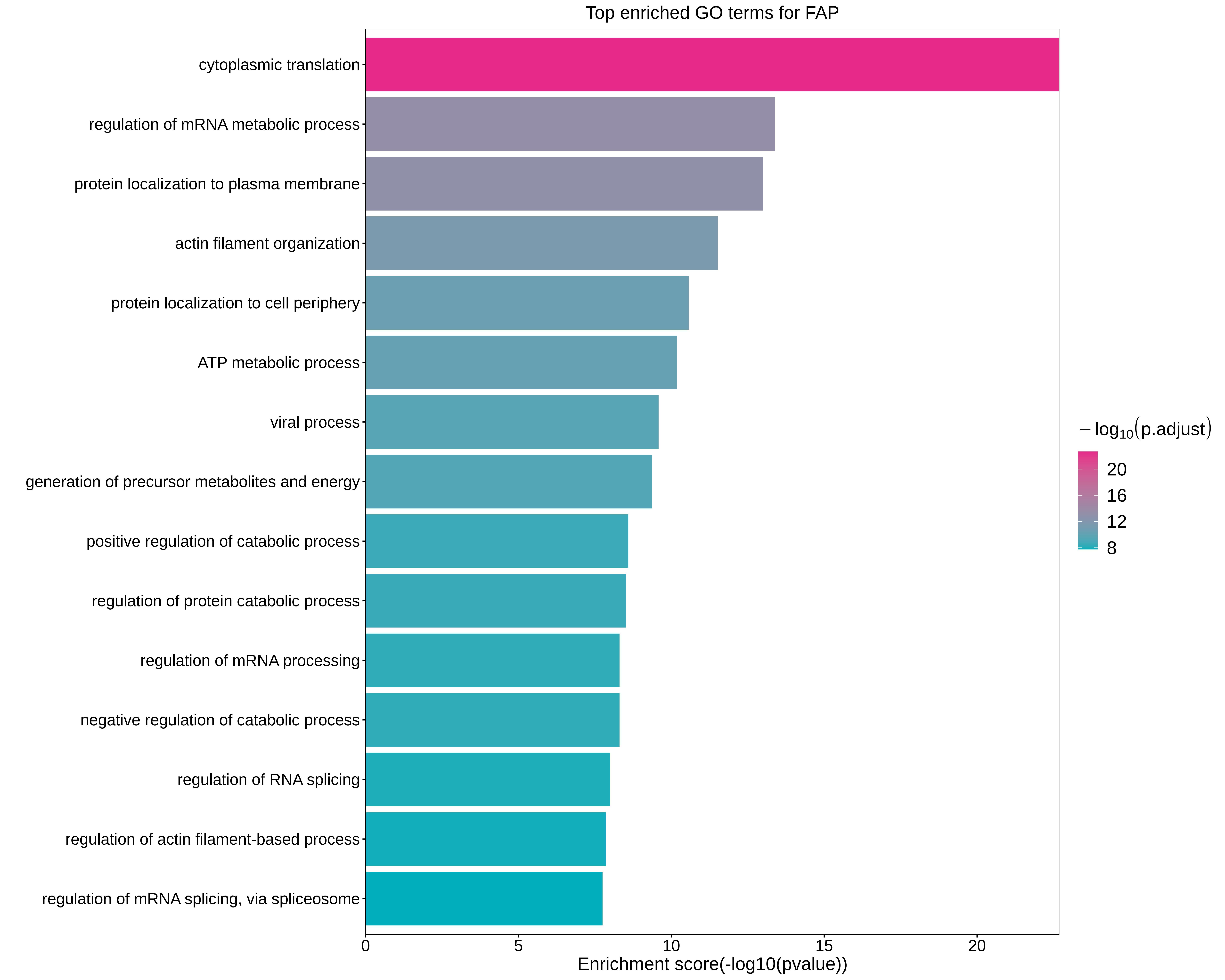
 Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states
Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states
Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states Annotation of somatic variants for genes involved in malignant transformation
Annotation of somatic variants for genes involved in malignant transformation Identification of chemicals and drugs interact with genes involved in malignant transfromation
Identification of chemicals and drugs interact with genes involved in malignant transfromation