|
|||||
|
 | |
 | |
 | |
 | |
 | |
 | |
 |
Gene: AK1 |
Gene summary for AK1 |
| Gene information | Species | Human | Gene symbol | AK1 | Gene ID | 203 |
| Gene name | adenylate kinase 1 | |
| Gene Alias | HTL-S-58j | |
| Cytomap | 9q34.11 | |
| Gene Type | protein-coding | GO ID | GO:0006139 | UniProtAcc | P00568 |
Top |
Malignant transformation analysis |
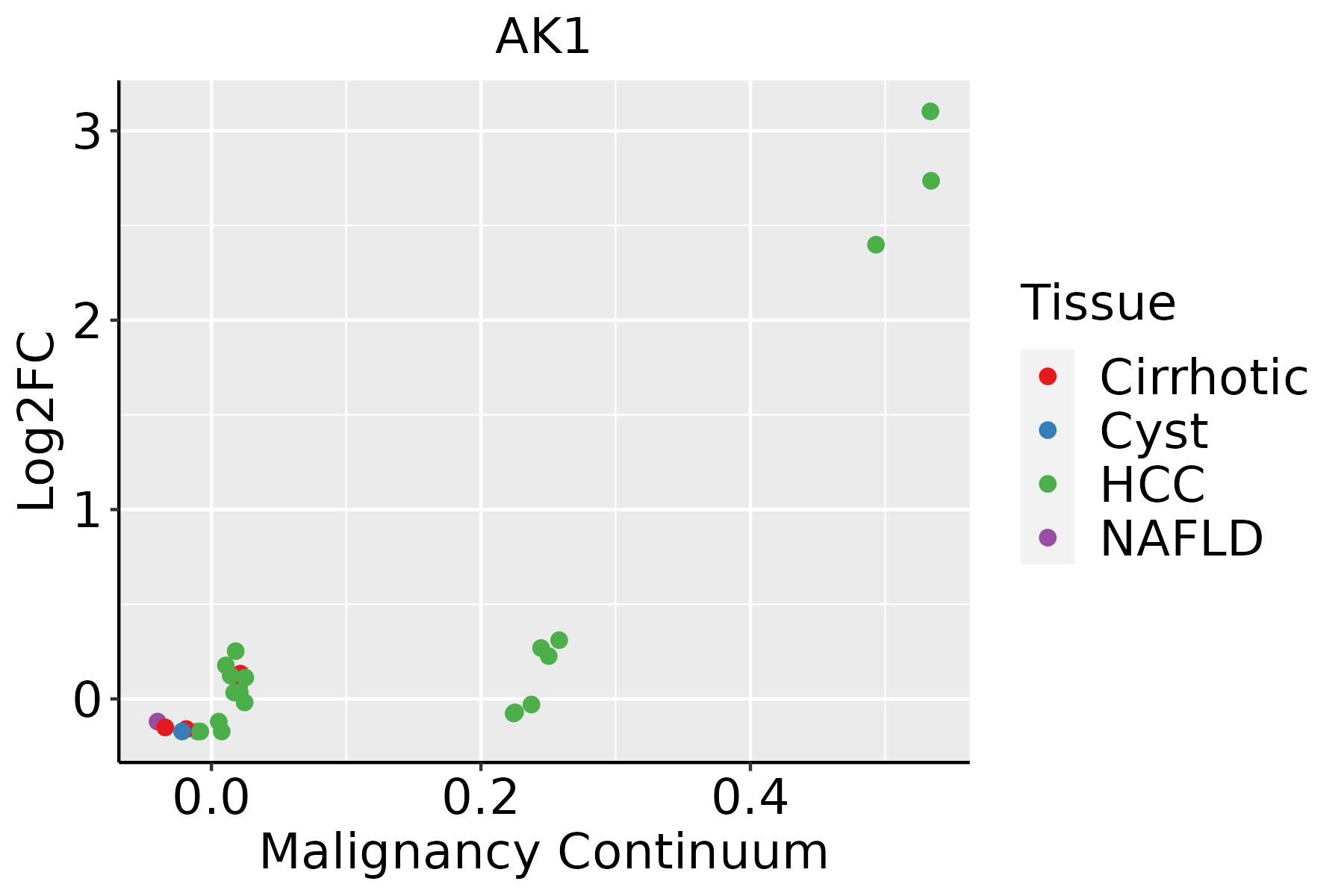
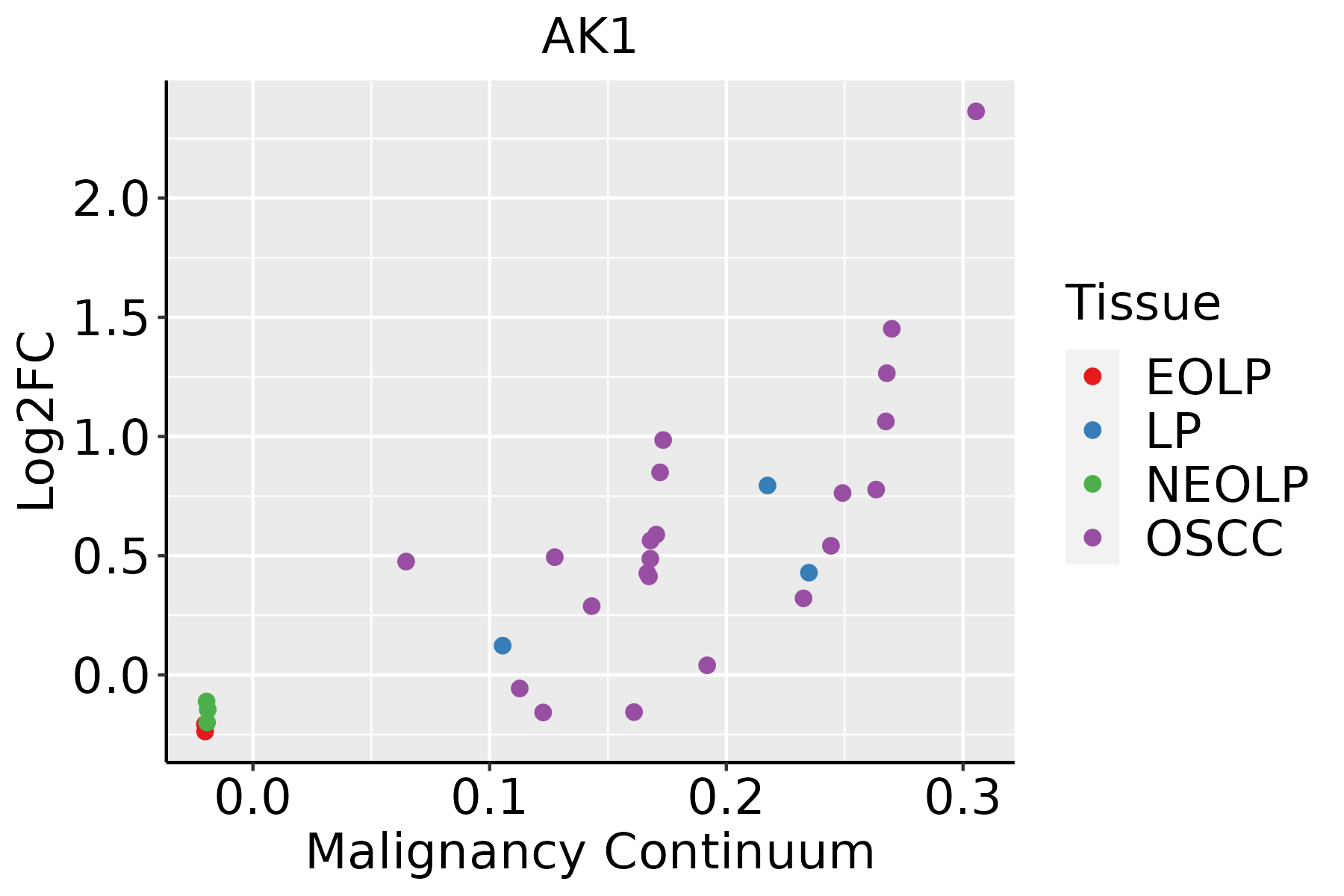
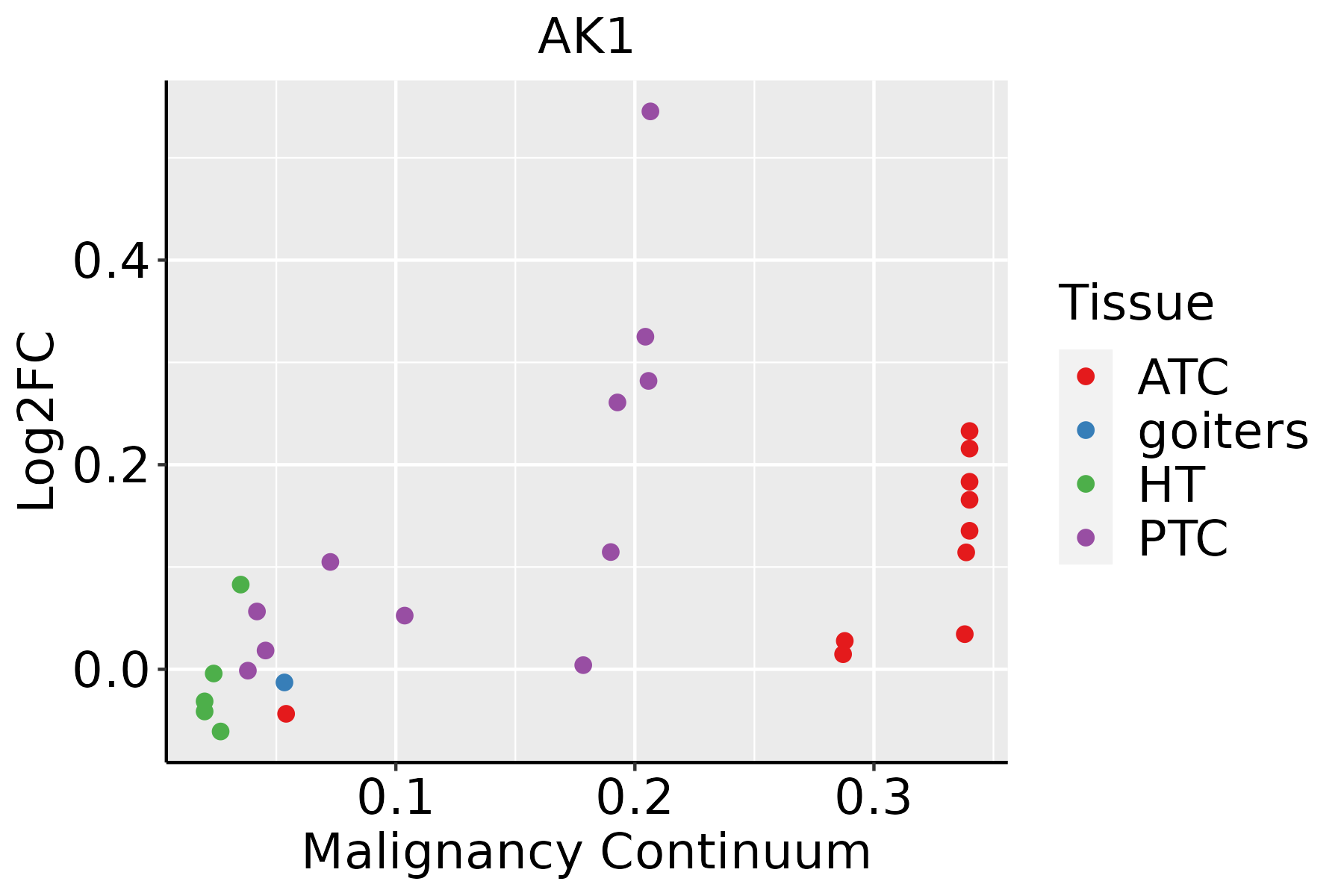
 Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells Identification of the aberrant gene expression in precancerous and cancerous lesions by comparing the gene expression of stem-like cells in diseased tissues with normal stem cells |
| Entrez ID | Symbol | Replicates | Species | Organ | Tissue | Adj P-value | Log2FC | Malignancy |
| 203 | AK1 | LZE4T | Human | Esophagus | ESCC | 1.03e-13 | 3.11e-01 | 0.0811 |
| 203 | AK1 | LZE5T | Human | Esophagus | ESCC | 3.35e-11 | 5.37e-01 | 0.0514 |
| 203 | AK1 | LZE7T | Human | Esophagus | ESCC | 3.17e-12 | 5.95e-01 | 0.0667 |
| 203 | AK1 | LZE8T | Human | Esophagus | ESCC | 1.99e-03 | 1.54e-01 | 0.067 |
| 203 | AK1 | LZE20T | Human | Esophagus | ESCC | 1.09e-12 | 4.75e-01 | 0.0662 |
| 203 | AK1 | LZE22T | Human | Esophagus | ESCC | 2.53e-06 | 3.64e-01 | 0.068 |
| 203 | AK1 | LZE24T | Human | Esophagus | ESCC | 1.08e-15 | 4.48e-01 | 0.0596 |
| 203 | AK1 | LZE21T | Human | Esophagus | ESCC | 1.33e-04 | 3.83e-01 | 0.0655 |
| 203 | AK1 | LZE6T | Human | Esophagus | ESCC | 6.84e-15 | 6.14e-01 | 0.0845 |
| 203 | AK1 | P1T-E | Human | Esophagus | ESCC | 1.49e-10 | 4.23e-01 | 0.0875 |
| 203 | AK1 | P2T-E | Human | Esophagus | ESCC | 1.76e-36 | 5.68e-01 | 0.1177 |
| 203 | AK1 | P4T-E | Human | Esophagus | ESCC | 1.28e-33 | 6.76e-01 | 0.1323 |
| 203 | AK1 | P5T-E | Human | Esophagus | ESCC | 3.51e-08 | 2.31e-01 | 0.1327 |
| 203 | AK1 | P8T-E | Human | Esophagus | ESCC | 1.35e-32 | 6.52e-01 | 0.0889 |
| 203 | AK1 | P9T-E | Human | Esophagus | ESCC | 9.00e-17 | 4.76e-01 | 0.1131 |
| 203 | AK1 | P10T-E | Human | Esophagus | ESCC | 1.11e-03 | 1.10e-01 | 0.116 |
| 203 | AK1 | P11T-E | Human | Esophagus | ESCC | 1.05e-28 | 8.02e-01 | 0.1426 |
| 203 | AK1 | P12T-E | Human | Esophagus | ESCC | 6.05e-19 | 3.68e-01 | 0.1122 |
| 203 | AK1 | P15T-E | Human | Esophagus | ESCC | 2.63e-14 | 3.37e-01 | 0.1149 |
| 203 | AK1 | P16T-E | Human | Esophagus | ESCC | 1.53e-13 | 2.65e-01 | 0.1153 |
| Page: 1 2 3 4 5 6 7 8 |
| ∗log2FC in expression of this searched gene in stem-like cells from each diseased tissue sample relative to stem-like cells in normal samples in each tissue plotted against the malignancy continuum. Samples are colored based on if they are from different disease stage. |
Top |
Malignant transformation related pathway analysis |
 Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer Find out the enriched GO biological processes and KEGG pathways involved in transition from healthy to precancer to cancer |
| Tissue | Disease Stage | Enriched GO biological Processes |
| Colorectum | AD |  |
| Colorectum | SER | 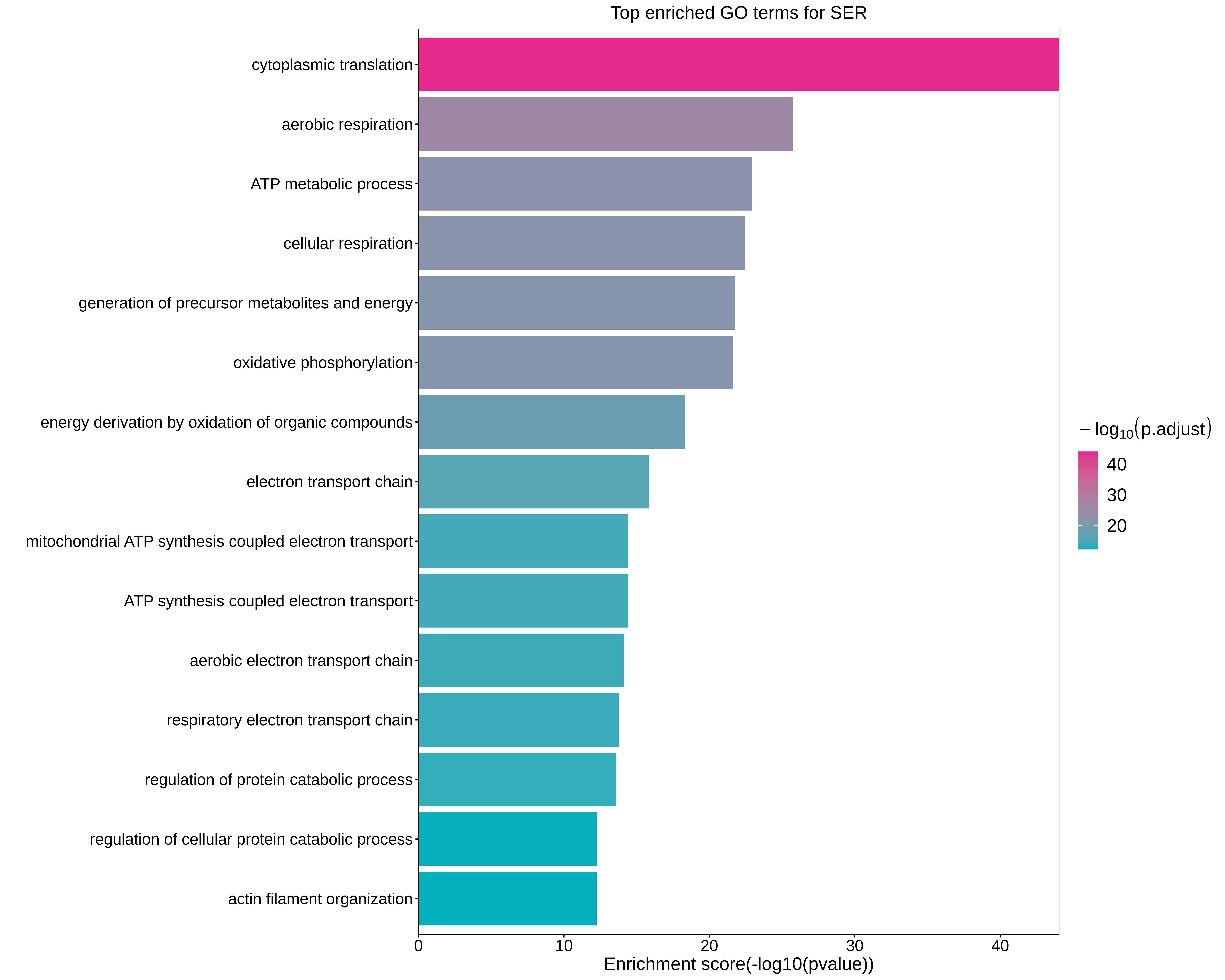 |
| Colorectum | MSS |  |
| Colorectum | MSI-H | 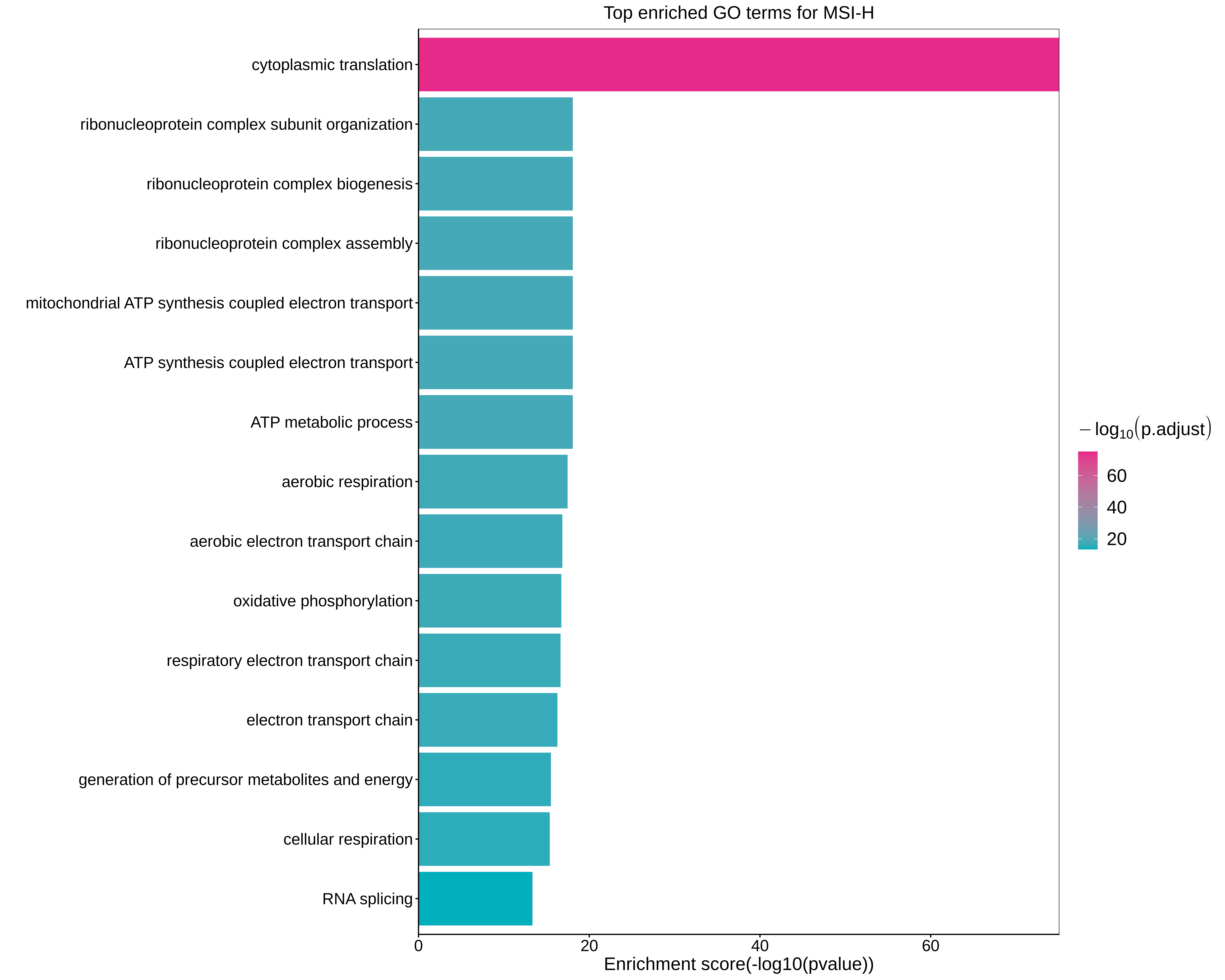 |
| Colorectum | FAP | 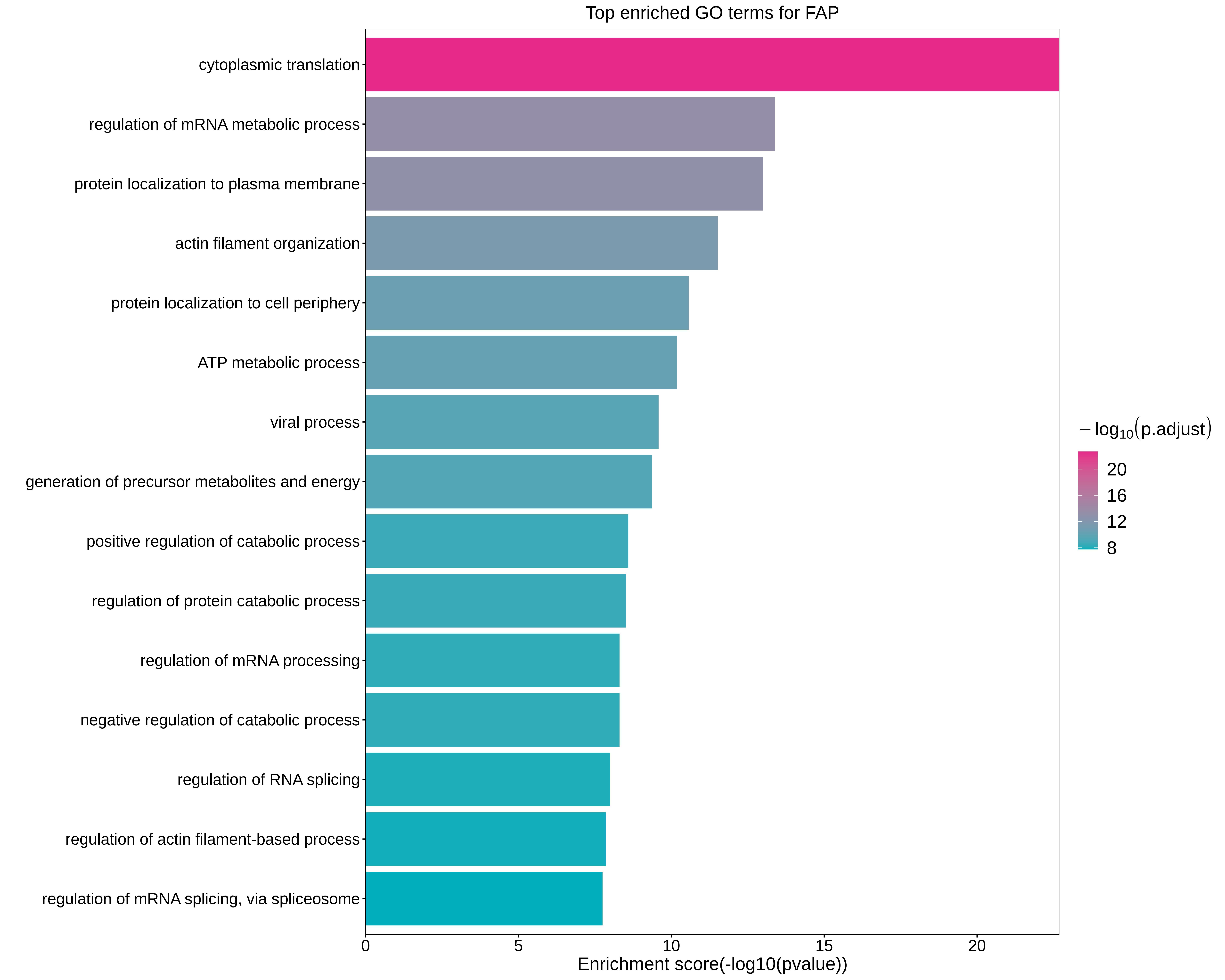 |
| ∗Top 15 enriched GO BP terms are showed in the bar plot of each disease state in each tissue. Each row represents a significant GO biological process which is colored according to the -log10(p.adjust). |
| Page: 1 2 3 4 5 6 7 8 9 |
| GO ID | Tissue | Disease Stage | Description | Gene Ratio | Bg Ratio | pvalue | p.adjust | Count |
| GO:00343295 | Stomach | GC | cell junction assembly | 42/1159 | 420/18723 | 1.50e-03 | 1.62e-02 | 42 |
| GO:00070445 | Stomach | GC | cell-substrate junction assembly | 14/1159 | 95/18723 | 2.07e-03 | 2.09e-02 | 14 |
| GO:00019525 | Stomach | GC | regulation of cell-matrix adhesion | 17/1159 | 128/18723 | 2.33e-03 | 2.25e-02 | 17 |
| GO:00091674 | Stomach | GC | purine ribonucleoside monophosphate metabolic process | 8/1159 | 41/18723 | 3.22e-03 | 2.84e-02 | 8 |
| GO:01501165 | Stomach | GC | regulation of cell-substrate junction organization | 11/1159 | 71/18723 | 4.07e-03 | 3.41e-02 | 11 |
| GO:00071605 | Stomach | GC | cell-matrix adhesion | 25/1159 | 233/18723 | 5.24e-03 | 4.11e-02 | 25 |
| GO:004603411 | Stomach | CAG with IM | ATP metabolic process | 72/1050 | 277/18723 | 6.46e-29 | 1.72e-25 | 72 |
| GO:000914211 | Stomach | CAG with IM | nucleoside triphosphate biosynthetic process | 23/1050 | 85/18723 | 1.62e-10 | 2.98e-08 | 23 |
| GO:000914111 | Stomach | CAG with IM | nucleoside triphosphate metabolic process | 26/1050 | 112/18723 | 4.23e-10 | 6.83e-08 | 26 |
| GO:000915011 | Stomach | CAG with IM | purine ribonucleotide metabolic process | 48/1050 | 368/18723 | 4.36e-08 | 3.32e-06 | 48 |
| GO:000616311 | Stomach | CAG with IM | purine nucleotide metabolic process | 50/1050 | 396/18723 | 6.52e-08 | 4.63e-06 | 50 |
| GO:000925911 | Stomach | CAG with IM | ribonucleotide metabolic process | 48/1050 | 385/18723 | 1.76e-07 | 1.16e-05 | 48 |
| GO:000915211 | Stomach | CAG with IM | purine ribonucleotide biosynthetic process | 28/1050 | 169/18723 | 2.35e-07 | 1.51e-05 | 28 |
| GO:007252111 | Stomach | CAG with IM | purine-containing compound metabolic process | 50/1050 | 416/18723 | 3.04e-07 | 1.82e-05 | 50 |
| GO:001969311 | Stomach | CAG with IM | ribose phosphate metabolic process | 48/1050 | 396/18723 | 4.08e-07 | 2.34e-05 | 48 |
| GO:000926011 | Stomach | CAG with IM | ribonucleotide biosynthetic process | 28/1050 | 182/18723 | 1.11e-06 | 5.44e-05 | 28 |
| GO:000911711 | Stomach | CAG with IM | nucleotide metabolic process | 54/1050 | 489/18723 | 1.52e-06 | 7.14e-05 | 54 |
| GO:000675311 | Stomach | CAG with IM | nucleoside phosphate metabolic process | 54/1050 | 497/18723 | 2.49e-06 | 1.07e-04 | 54 |
| GO:004639011 | Stomach | CAG with IM | ribose phosphate biosynthetic process | 28/1050 | 190/18723 | 2.66e-06 | 1.12e-04 | 28 |
| GO:000616411 | Stomach | CAG with IM | purine nucleotide biosynthetic process | 28/1050 | 191/18723 | 2.95e-06 | 1.21e-04 | 28 |
| Page: 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 |
| Pathway ID | Tissue | Disease Stage | Description | Gene Ratio | Bg Ratio | pvalue | p.adjust | qvalue | Count |
| hsa012325 | Esophagus | ESCC | Nucleotide metabolism | 59/4205 | 85/8465 | 1.67e-04 | 6.58e-04 | 3.37e-04 | 59 |
| hsa012405 | Esophagus | ESCC | Biosynthesis of cofactors | 97/4205 | 153/8465 | 3.88e-04 | 1.35e-03 | 6.94e-04 | 97 |
| hsa0123212 | Esophagus | ESCC | Nucleotide metabolism | 59/4205 | 85/8465 | 1.67e-04 | 6.58e-04 | 3.37e-04 | 59 |
| hsa0124012 | Esophagus | ESCC | Biosynthesis of cofactors | 97/4205 | 153/8465 | 3.88e-04 | 1.35e-03 | 6.94e-04 | 97 |
| hsa01240 | Liver | Cirrhotic | Biosynthesis of cofactors | 66/2530 | 153/8465 | 3.11e-04 | 1.99e-03 | 1.23e-03 | 66 |
| hsa01232 | Liver | Cirrhotic | Nucleotide metabolism | 39/2530 | 85/8465 | 1.27e-03 | 6.73e-03 | 4.15e-03 | 39 |
| hsa012401 | Liver | Cirrhotic | Biosynthesis of cofactors | 66/2530 | 153/8465 | 3.11e-04 | 1.99e-03 | 1.23e-03 | 66 |
| hsa012321 | Liver | Cirrhotic | Nucleotide metabolism | 39/2530 | 85/8465 | 1.27e-03 | 6.73e-03 | 4.15e-03 | 39 |
| hsa012402 | Liver | HCC | Biosynthesis of cofactors | 103/4020 | 153/8465 | 4.67e-07 | 5.05e-06 | 2.81e-06 | 103 |
| hsa012322 | Liver | HCC | Nucleotide metabolism | 59/4020 | 85/8465 | 3.30e-05 | 1.88e-04 | 1.04e-04 | 59 |
| hsa012403 | Liver | HCC | Biosynthesis of cofactors | 103/4020 | 153/8465 | 4.67e-07 | 5.05e-06 | 2.81e-06 | 103 |
| hsa012323 | Liver | HCC | Nucleotide metabolism | 59/4020 | 85/8465 | 3.30e-05 | 1.88e-04 | 1.04e-04 | 59 |
| hsa012324 | Oral cavity | OSCC | Nucleotide metabolism | 54/3704 | 85/8465 | 1.78e-04 | 5.95e-04 | 3.03e-04 | 54 |
| hsa012404 | Oral cavity | OSCC | Biosynthesis of cofactors | 88/3704 | 153/8465 | 3.84e-04 | 1.20e-03 | 6.12e-04 | 88 |
| hsa0123211 | Oral cavity | OSCC | Nucleotide metabolism | 54/3704 | 85/8465 | 1.78e-04 | 5.95e-04 | 3.03e-04 | 54 |
| hsa0124011 | Oral cavity | OSCC | Biosynthesis of cofactors | 88/3704 | 153/8465 | 3.84e-04 | 1.20e-03 | 6.12e-04 | 88 |
| hsa0123221 | Oral cavity | LP | Nucleotide metabolism | 42/2418 | 85/8465 | 3.62e-05 | 2.36e-04 | 1.52e-04 | 42 |
| hsa0124021 | Oral cavity | LP | Biosynthesis of cofactors | 57/2418 | 153/8465 | 1.17e-02 | 3.91e-02 | 2.52e-02 | 57 |
| hsa0123231 | Oral cavity | LP | Nucleotide metabolism | 42/2418 | 85/8465 | 3.62e-05 | 2.36e-04 | 1.52e-04 | 42 |
| hsa0124031 | Oral cavity | LP | Biosynthesis of cofactors | 57/2418 | 153/8465 | 1.17e-02 | 3.91e-02 | 2.52e-02 | 57 |
| Page: 1 |
Top |
Cell-cell communication analysis |
 Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states Identification of potential cell-cell interactions between two cell types and their ligand-receptor pairs for different disease states |
| Ligand | Receptor | LRpair | Pathway | Tissue | Disease Stage |
| Page: 1 |
Top |
Single-cell gene regulatory network inference analysis |
 Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states Find out the significant the regulons (TFs) and the target genes of each regulon across cell types for different disease states |
| TF | Cell Type | Tissue | Disease Stage | Target Gene | RSS | Regulon Activity |
| ∗The dot plots of a searched regulon are shown for all cell subpopulations in each disease state of each tissue based on the regulon specific score inferred using pySCENIC and by calculating the average expression. |
| Page: 1 |
Top |
Somatic mutation of malignant transformation related genes |
 Annotation of somatic variants for genes involved in malignant transformation Annotation of somatic variants for genes involved in malignant transformation |
| Hugo Symbol | Variant Class | Variant Classification | dbSNP RS | HGVSc | HGVSp | HGVSp Short | SWISSPROT | BIOTYPE | SIFT | PolyPhen | Tumor Sample Barcode | Tissue | Histology | Sex | Age | Stage | Therapy Types | Drugs | Outcome |
| AK1 | SNV | Missense_Mutation | c.335N>A | p.Pro112Gln | p.P112Q | protein_coding | deleterious(0) | probably_damaging(1) | TCGA-A2-A25A-01 | Breast | breast invasive carcinoma | Female | <65 | I/II | Unspecific | Cytoxan | SD | ||
| AK1 | SNV | Missense_Mutation | rs150701679 | c.133N>A | p.Val45Met | p.V45M | protein_coding | deleterious(0) | possibly_damaging(0.584) | TCGA-5M-AAT6-01 | Colorectum | colon adenocarcinoma | Female | <65 | III/IV | Unknown | Unknown | PD | |
| AK1 | SNV | Missense_Mutation | c.371G>T | p.Arg124Leu | p.R124L | protein_coding | tolerated(0.06) | benign(0.012) | TCGA-CM-4743-01 | Colorectum | colon adenocarcinoma | Male | >=65 | I/II | Chemotherapy | capecitabine | SD | ||
| AK1 | SNV | Missense_Mutation | rs201971818 | c.526N>A | p.Val176Ile | p.V176I | protein_coding | tolerated(0.31) | possibly_damaging(0.518) | TCGA-F4-6856-01 | Colorectum | colon adenocarcinoma | Male | <65 | I/II | Ancillary | leucovorin | CR | |
| AK1 | SNV | Missense_Mutation | novel | c.559N>G | p.Arg187Gly | p.R187G | protein_coding | tolerated(0.32) | benign(0.414) | TCGA-AG-3726-01 | Colorectum | rectum adenocarcinoma | Female | <65 | I/II | Unknown | Unknown | SD | |
| AK1 | SNV | Missense_Mutation | rs761271981 | c.469N>A | p.Asp157Asn | p.D157N | protein_coding | deleterious(0) | probably_damaging(0.998) | TCGA-DC-5337-01 | Colorectum | rectum adenocarcinoma | Male | >=65 | I/II | Unknown | Unknown | SD | |
| AK1 | SNV | Missense_Mutation | rs150701679 | c.133N>A | p.Val45Met | p.V45M | protein_coding | deleterious(0) | possibly_damaging(0.584) | TCGA-A5-A0G1-01 | Endometrium | uterine corpus endometrioid carcinoma | Female | >=65 | I/II | Unknown | Unknown | SD | |
| AK1 | SNV | Missense_Mutation | c.371N>C | p.Arg124Pro | p.R124P | protein_coding | deleterious(0.03) | benign(0.44) | TCGA-AX-A0IW-01 | Endometrium | uterine corpus endometrioid carcinoma | Female | >=65 | III/IV | Unspecific | Carboplatin & Paclitaxel | SD | ||
| AK1 | SNV | Missense_Mutation | rs375517545 | c.559N>T | p.Arg187Cys | p.R187C | protein_coding | tolerated(0.2) | benign(0.034) | TCGA-AX-A0IZ-01 | Endometrium | uterine corpus endometrioid carcinoma | Female | <65 | I/II | Unknown | Unknown | SD | |
| AK1 | SNV | Missense_Mutation | novel | c.50C>T | p.Ala17Val | p.A17V | protein_coding | tolerated(0.69) | benign(0) | TCGA-BG-A3EW-01 | Endometrium | uterine corpus endometrioid carcinoma | Female | <65 | III/IV | Unknown | Unknown | SD |
| Page: 1 2 |
Top |
Related drugs of malignant transformation related genes |
 Identification of chemicals and drugs interact with genes involved in malignant transfromation Identification of chemicals and drugs interact with genes involved in malignant transfromation |
| (DGIdb 4.0) |
| Entrez ID | Symbol | Category | Interaction Types | Drug Claim Name | Drug Name | PMIDs |
| Page: 1 |