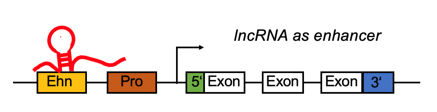
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | TYMP,TNFRSF17,TMEM119,ARHGEF1,SAMHD1,SIRPB2,VAMP1,XRN1,ISG20,MYCBP2,TNFRSF4,DUSP2,LTB,ABHD17A,RAB20,ADAM19,SPATA13,MAST3,BATF3,ROM1,MZF1,RGS10,MAPK8IP3,KLF2,TAP2,N4BP2L2,DMTF1,MKNK1,L3HYPDH,ANKRD44,UNKL,CD69,CALHM6,PARP9,LMBR1L,CLEC10A,RUBCNL,IRAK4,STAT2,RFTN1,COTL1,VMP1,CENATAC,TP53INP1,TMC8,KCTD13,MSANTD2,TUBGCP6,SPI1,CD27,ARAP2,CLK4,GIMAP7,HK3,DENND1C,KCNN4,ZBED5,PRKD2,VASP,INPP5B,CD37,DOK1,SPOCK2,TTC14,SPIB,HCK,ALDH3B1,MOB3A,TNFSF8,TAZ,SPG7,PDZK1IP1,CD74,TMEM106A,GIT2,MLKL,RGS19,CLCN6,ECHDC2,OGT,CCR7,NOD1,RAB4B,PDE7A,KLRB1,RNF19A,CLIC2,POLG2,CXCL10,CHKB,IL15RA,UBD,JAK3,MBD2,BBIP1,FLI1,DHX58,MAP4K1,ABHD14B,OSGEP,USF1,CTC1,MFSD1,CLK1,ATXN7,OTULIN,IL12RB1,CHI3L1,PAOX,OFD1,MOV10,DRAM1,ADAM8,FCHSD1,TREM2,CD84,CC2D1B,GZMM,GMIP,LMO2,SLC7A7,HSH2D,INO80E,HCLS1,C3orf62,GNAI2,APBA3,MAP3K14,HLA-DRB5,APOC1,SAMSN1,DOCK10,ANKRD22,FCER2,BIN2,HLA-DMA,MIIP,ORMDL1,CYBA,FKBP15,AGER,MBD4,AGO3,NFKBIZ,NFAM1,CXCL16,CDK11A,DPYD,TECPR1,DOK3,CMKLR1,PASK,NAGPA,RASGEF1B,FYN,EPSTI1,FADS3,CCDC191,NR2C2,SELP,IL18,IRAG2,IL27RA,TLR1,PPP1R9B,PARP10,UNC13D,CD72,MAN2B1,LAMP3,CCDC69,TMEM273,FAM193B,AOAH,CYRIA,ABCA7,CXCR3,SLC25A28,NR1H3,GLRX,GRK3,ZFC3H1,TAGAP,PILRA,N4BP2L1,HLA-DQB2,HEXD,LPAR5,TIFA,EIF4A1,RBP5,PIP4K2A,FMNL3,HLA-DMB,ERCC5,IKZF3,GCA,MAX,TRIM14,DEF6,HCST,SAT1,CCR1,FCER1G,ATM,PTK2B,NCF4,STK17B,SP140,TNFSF10,LENG8,SH2B1,PPP1R12C,ATP2A3,DGKA,UBE2G2,PPWD1,MLLT6,CYB561A3,C16orf54,UCP2,CRLF3,CNOT6L,LILRB2,RASAL3,RASSF2,CD19,TNIP1,GBP3,BANP,TNFAIP3,C1QB,THOC1,ANKMY1,PML,LPXN,SWAP70,TMEM175,WARS1,AKNA,RIPK3,LRRC25,SDC3,IFNGR1,TMEM176A,C1QC,PSMB9,TNK2,ARHGAP27,PRF1,CD8A,RAPGEF1,FOXP3,NBEAL2,PARP12,CD48,RASGRP2,PIK3AP1,CBLB,SETX,ZNF44,NCKAP1L,SMG1,LCP2,PIDD1,FGR,PLA2G7,POLD4,CD247,SEMA4D,FNBP1,GBGT1,SIGLEC9,CD47,ELMO1,ICAM2,RNF135,ANKDD1A,NFKBID,GSDMD,ABTB1,TASL,SLX4,TRPM2,PDCD1,DTX1,CIITA,SMYD4,IKBKB,TYROBP,CYP4V2,CD79B,MS4A1,XCL2,ARHGAP17,SLC15A3,PIK3R5,ANKRD49,TNFRSF13C,GANC,LY75,CCDC125,SCIMP,UNC93B1,CASP4,ITK,FNBP4,DENND6B,SYNRG,SLC46A3,TRADD,ARHGDIB,PHF11,CYB5R4,RSRP1,RHBDF2,PPM1K,IL11RA,SSH2,CHD2,ST3GAL5,LAG3,PTPN6,CLEC2D,SYNGR2,MCOLN2,APOBEC3H,PSMB10,PSME1,LTB4R,EXOC3L4,ODF3B,CLEC2B,HEMK1,MVP,LINS1,SYTL3,HERC1,PRAM1,GIMAP8,RABEP2,TRIM22,SBNO2,OGFR,APOL2,GSAP,GGA2,DOCK8,NADK,FBXL8,SCARF1,RBM6,DRAM2,IL4I1,PARVG,HLA-B,IL32,KRTCAP2,ZNF862,IL10RB,IDUA,ADPRH,ARRB2,C1R,ORAI3,RASSF5,RAC2,MYL5,CD3D,NLRP1,DOK2,CETP,TNFRSF14,XAF1,TCF25,CAPZB,CASP1,UBXN11,EME2,ZEB2,CROCC,HAVCR2,USF2,PNISR,MS4A6A,TMEM63A,AMT,FHOD1,LCP1,C19orf38,PTPN7,CEPT1,FKBP11,CARD8,SIGLEC10,PLA2G2D,CD300LF,IFFO1,LGALS2,RNF149,CD180,BCL2,TAP1,MYO1F,RASSF1,TRMU,MYO9B,PPP1R18,NAGK,ITGB2,GZMH,IL18R1,PARP15,ETV7,IL4R,ADPGK,FAM214A,PHF19,TFEB,PLEK,MZB1,RNF44,ITGB7,SIGLEC1,FMNL1,ACAP1,LYSMD2,LHFPL2,TBRG1,STING1,CD79A,MEF2C,EML3,KLHDC4,TRPV2,CCNL2,NUAK2,PCED1B,CTSC,RNPC3,ANKRD10,IRF1,TNFRSF1B,SECTM1,LRCH3,SH3BGRL,SNX20,MSL3,CSF1R,DDTL,SGSM3,EHBP1L1,TP53I13,SH2B3,TYK2,STAT4,OSBPL7,SP100,SEPTIN1,KAT2A,PTAFR,TMEM176B,SLC17A9,SIPA1,NIN,BCL2A1,PCSK7,MARCHF1,OSCAR,VAV1,CYBC1,FBXW7,CD7,P2RY8,TAPBP,LNPEP,S1PR2,IRF7,ITSN2,RESF1,ERMARD,SIRPG,FAM53B,BATF2,SRRM2,ERAP2,PSMB8,DPEP2,PIK3IP1,PARP3,DDHD1,MTHFR,HAPLN3,HDAC10,FCGR3A,GMFG,SLC2A5,HSF4,SERPINB9,ANKS3,ARGLU1,C1RL,CCNL1,CEACAM21,RUBCN,IRF2,OSTF1,SLC9A9,PNRC1,VMAC,TCL1A,DBNL,ICOS,CELF1,BTK,MICALL2,AMPD3,CRYGS,PRDM2,TBC1D2B,DENND2D,MFNG,HSD3B7,MOB3C,PBX4,BIRC3,ZNF83,CLEC7A,MCUB,PLCG2,SP110,TXNDC11,TAPBPL,KLHL6,STAT1,C2,PIM2,SH3KBP1,VOPP1,NEIL1,SZT2,TNIP2,DDX5,GRIPAP1,ST8SIA4,CYLD,PLCB2,CD58,MAPK11,PARP8,UBA7,LRRK1,STK4,CXCR6,LYST,IL2RB,RASSF4,IGFLR1,LUC7L,C1S,RAP2B,ADAMDEC1,RUNX3,C1orf54,SAMD9L,TWF2,CASP8,FAM20A,CCL5,TRA2A,NSMCE4A,CKLF,KMT2A,REV1,PREX1,AP1S2,GPR132,APBB3,CTSS,NRIP2,TEPSIN,RBM33,PAG1,WDR37,FES,AGAP6,WRAP73,MYO1G,GABPB2,SH2D2A,CD44,TNFRSF18,CEBPA,SH3BGRL3,ADAMTS10,ODF2L,TNFAIP2,ACKR1,C3,IRF3,ZMYND15,SLFN5,NPC2,CLEC16A,ERICH1,EAF2,BLNK,VASH1,SLA2,RFX5,SNAI3,APOBR,ANKRD13A,HLA-DRB1,ARHGAP30,BIN3,LILRB4,CSAD,RABGAP1L,RINL,DDX39A,TBC1D10C,TRANK1,SAMD10,CD52,DCAF8,CALHM2,KIF21B,PHYKPL,FUCA1,SNX29,LST1,MS4A7,PAN3,PHKG2,ARHGAP4,SELL,CSF3R,DTX3L,TBC1D17,RALGDS,HLA-DOA,ARHGAP9,GRK6,UBE2L6,ADA2,RIPOR2,CENPT,CYTH1,FAS,NFKBIA,EMP3,CARD11,EMB,AIM2,GM2A,ACP5,CXCR4,VNN2,ARRDC2,C2CD2L,SH2B2,TLR5,CYTIP,EXOG,RHOG,TAF1C,ZBP1,TRAF3,SIT1,MYO7A,TRAPPC14,ARHGAP45,ARSA,MAFB,CGAS,HVCN1,CXXC1,EZH1,MRNIP,STK17A,ELMOD3,DDX17,FERMT3,NECAP2,SLC7A6,MED15,APOL3,SENP7,CHFR,SOD2,PXK,GZMK,PGGHG,LAT2,TELO2,WSB1,IL18BP,EFHD2,GAB3,LAPTM5,ANO9,ITGAL,CBLN3,ARPC1B,WASHC1,PSTPIP2,PTGDS,ELK4,KLC1,NINJ1,NMI,GPS2,TRIM52,DERL3,NOL12,JAK1,TTC13,ADGRG5,STAC3,MICAL1,AGAP2,ZMAT1,GOLGA8B,GRB2,GIMAP6,TSC22D3,ALPK1,STRADA,VPS13C,NLRC3,C1QA,CCDC88C,ZNF18,ABI3,GOLGA8A,ITGAE,MPEG1,TCIRG1,B2M,HLA-E,RNPEPL1,TBXAS1,DUSP28,RBM5,FYB1,NKTR,AP1B1,YPEL3,SLCO2B1,PLEKHO1,RILPL2,C19orf71,IL2RG,TTLL3,GNG7,SLC25A45,ETS1,GBP4,RCSD1,STK10,NMRK1,SLAMF6,C3AR1,IFI35,GNGT2,ARHGEF6,ADAP2,LRP5L,DENND4B,TEP1,FXYD5,USPL1,TMC6,EBI3,SFMBT2,WHAMM,NFKBIE,TRAPPC2,SIRT7,STK38 | | Exist in public source | | NA |
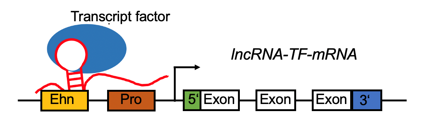
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | DCAF8,PRPF38B,SLC11A1,STAP1,SGSM3,TNIP2,GM2A,ACSL5,RUNX3,DGKA,MAST3,HLA-DRA,CCDC88B,ZNF18,RAB20,CD19,S1PR2,SLA2,DERL3,IL12RB1,TNFSF10,XAF1,ENTPD1,RGS10,DRAM1,CHI3L1,DTX2,ZMAT1,DTX3L,VPS13C,BIRC3,CCDC125,SLAMF6,RILPL2,FGR,ARRB2,FCHO1,PLA2G2D,ALPK1,BCL2,AOAH,SMG1,TIFA,DOCK2,CD8A,FDCSP,CTC1,CXCL10,HSH2D,STK38,TMEM176B,CCDC109B,ERAP2,TBC1D10C,IGFLR1,NFKBID,TRMU,TNFSF12,HLA-DQA1,RALGDS,TLR1,RARRES3,HEMK1,ZMIZ2,RAB37,PPP1R9B,DDX17,NMRK1,TNFAIP8L2,UBE2L6,UBA7,RBM5,E4F1,ACP5,MB21D1,IRF5,KLHDC4,IER5,MBNL1,TAZ,AGAP2,IL3RA,STAT2,APOBEC3D,ACAP1,DAPP1,ARHGAP9,HAVCR2,KLF2,CD180,UCP2,ARHGAP25,GPR65,LSP1,GLIPR1,CD79A,ZNF83,C1QA,KIAA1551,ADAM19,EVL,IFNAR2,C16orf54,APOBR,STAC3,RASAL3,CD40,PIK3IP1,FCGR2A,AGO4,USF2,KLRB1,APOBEC3H,C7orf43,TMEM176A,CELF1,SH2D2A,TRIM14,VPREB3,ZC3H6,CARD11,CXorf21,TRIM69,FCER2,PIK3CD,CLASRP,HERPUD1,LUC7L,LILRB4,MAPK8IP3,MIIP,DPEP2,ELMO1,SNX20,COTL1,CCDC66,MZF1,HCST,RNF149,GZMK,ICAM2,C3orf62,FAM160B2,ITGA4,TWF2,VAMP1,PSTPIP2,PTGER4,GSTK1,FCRLA,MYL5,RAPGEF1,PAPLN,IL10RA,SH3KBP1,RNF135,UBXN11,IL2RA,STRADA,PRR14,TRAF5,CARD16,GIMAP6,GBP1,LYST,RELB,GBP3,GBP4,ACCS,RABEP2,CHFR,ENGASE,ZEB2,GRIPAP1,TEP1,OSTF1,PBX4,IL18R1,GIT2,GGA1,WSB1,CCDC130,TNFRSF13C,PARP9,FBXL8,AMPD3,LCP1,CD14,HLA-DOA,SCNM1,PIK3R5,ARRDC2,ARAP2,C5AR1,ABHD14B,RASGEF1B,TAGAP,COMMD3,CD48,VAV1,CD22,CCNL1,HLA-DOB,CTSC,LY96,PHF19,CEPT1,RNASE6,CTSW,IFNGR2,GZMM,CFLAR,ITGAL,ZDHHC18,LCP2,TRIM22,CD52,GZMA,APOBEC3C,SLX4,SYNGR2,CARD8,RBM6,TYROBP,FAM65B,SH3BGRL,PLCB2,ARHGAP30,DRAM2,TRADD,MAP3K1,SEMA4D,ARGLU1,CCL21,BTN2A2,FGD2,AP4B1,MDM4,FLI1,CASP4,SLFN5,CCDC69 | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
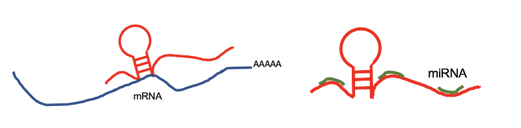
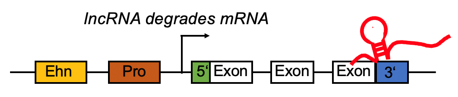
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | PRMT5,PSMB5,ATP5F1B,YWHAE,PEX19,STRAP | | Exist in public source | | NA |
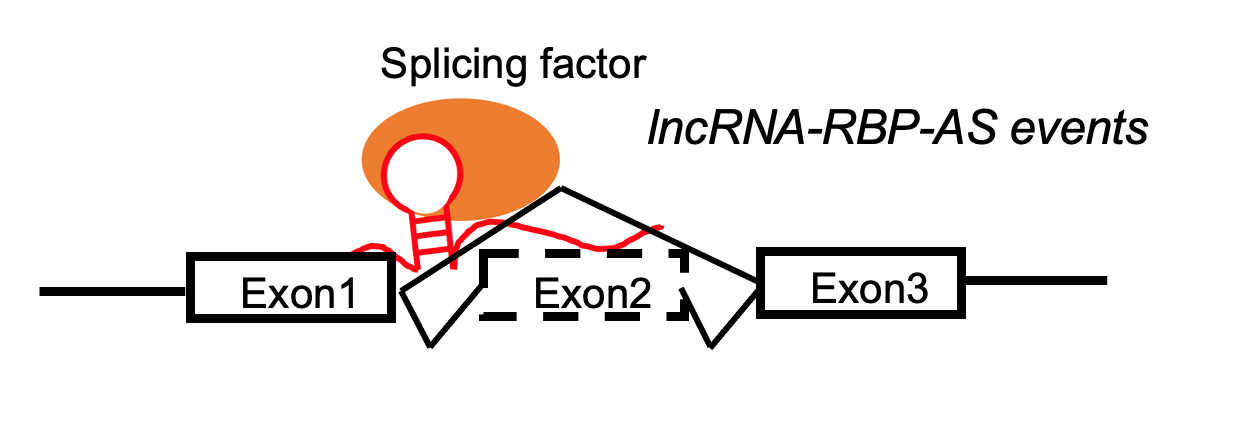
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000280153 | ENST00000625054 | exon_skip_75398 | chr11:72692752-72692785 | -11.65 | -0.6472 | ARAP1 | ENST00000393605 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_81540 | chr12:42384939-42384996 | -9.06 | -0.3236 | PPHLN1 | ENST00000395568 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_482144 | chr8:22539468-22539498 | -11.05 | -0.5023 | PPP3CC | ENST00000240139 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_285471 | chr17:3723287-3723383 | -14.11 | -0.1485 | ITGAE | ENST00000263087 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000280153 | ENST00000625054 | exon_skip_48296 | chr10:26755654-26755741 | -17.73 | -0.2814 | ABI1 | ENST00000376142 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_95025 | chr12:79805591-79805768 | -18.90 | -0.1112 | PPP1R12A | ENST00000261207,ENST00000450142 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_143275 | chr16:24939363-24939597 | -32.21 | -0.1419 | ARHGAP17 | ENST00000289968 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_378912 | chr3:152446703-152446757 | -8.70 | -0.3955 | MBNL1 | ENST00000282486,ENST00000463374 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_482403 | chr8:27445793-27445919 | -13.97 | -0.1486 | PTK2B | ENST00000346049,ENST00000397501 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_41813 | chr10:68337857-68337996 | -16.35 | -0.1220 | HNRNPH3 | ENST00000265866 | Frame-shift | | ENSG00000280153 | ENST00000625054 | exon_skip_53559 | chr10:101793912-101793998 | -10.93 | -0.1584 | OGA | ENST00000361464 | Frame-shift | | ENSG00000280153 | ENST00000625054 | exon_skip_382458 | chr3:38131595-38131638 | -12.22 | -0.2980 | ACAA1 | ENST00000333167 | Frame-shift | | ENSG00000280153 | ENST00000625054 | exon_skip_423795 | chr4:55888887-55888932 | -7.38 | -0.2838 | EXOC1 | ENST00000346134,ENST00000381295 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_470226 | chr7:103307595-103307708 | -13.10 | -0.1297 | PMPCB | ENST00000249269 | Frame-shift | | ENSG00000280153 | ENST00000625054 | exon_skip_499791 | chr9:128627924-128627942 | -6.81 | -0.6810 | SPTAN1 | ENST00000372731 | In-frame | | ENSG00000280153 | ENST00000625054 | exon_skip_98384 | chr12:124327408-124327633 | -29.90 | -0.1451 | NCOR2 | ENST00000405201 | In-frame |
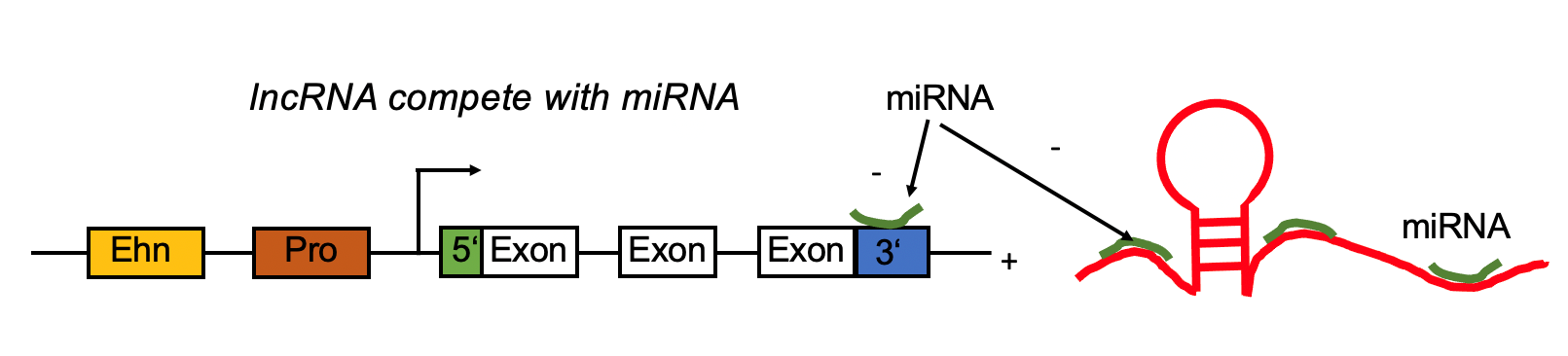
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000280153 | ENSG00000011600 | TYROBP | ACC,CHOL,GBM,KICH,KIRC,KIRP,LGG,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000019582 | CD74 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000023892 | DEF6 | ACC,BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000025708 | TYMP | ACC,BRCA,COAD,GBM,KICH,KIRC,LGG,SARC,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000028137 | TNFRSF1B | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THCA,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000043462 | LCP2 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000076928 | ARHGEF1 | ACC,BLCA,BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000085514 | PILRA | ACC,BRCA,CHOL,ESCA,GBM,KICH,KIRC,LGG,SARC,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000086730 | LAT2 | ACC,BLCA,BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000100450 | GZMH | ACC,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THCA,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000101336 | HCK | ACC,BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000102125 | TAZ | ACC,BRCA,CHOL,KICH,KIRC,KIRP,OV,PAAD,SKCM | KIRC | | | ENSG00000280153 | ENSG00000104894 | CD37 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000106785 | TRIM14 | ACC,KIRC,LGG,PAAD,SARC,SKCM,THCA | KIRC | | | ENSG00000280153 | ENSG00000107551 | RASSF4 | ACC,BLCA,BRCA,COAD,ESCA,KIRC,PAAD,SARC,SKCM,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000108798 | ABI3 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000110031 | LPXN | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000110057 | UNC93B1 | ACC,CHOL,GBM,KICH,LGG,OV,SARC,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000110446 | SLC15A3 | ACC,BRCA,CHOL,COAD,GBM,KICH,KIRC,KIRP,LGG,SARC,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000111348 | ARHGDIB | ACC,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000126264 | HCST | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000128284 | APOL3 | ACC,BLCA,BRCA,CESC,COAD,ESCA,GBM,KIRC,KIRP,LAML,LGG,LUSC,OV,PAAD,SKCM,STAD,TGCT,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000128340 | RAC2 | ACC,BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000129667 | RHBDF2 | ACC,BRCA,GBM,KICH,KIRC,LGG,PAAD,SARC,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000130066 | SAT1 | ACC,GBM,KIRC,KIRP,LGG,OV,SARC,TGCT | GBM | | | ENSG00000280153 | ENSG00000132274 | TRIM22 | ACC,BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000134470 | IL15RA | ACC,BRCA,COAD,ESCA,GBM,KIRC,KIRP,OV,SKCM,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000136048 | DRAM1 | ACC,ESCA,GBM,SKCM,TGCT | GBM | | | ENSG00000280153 | ENSG00000137496 | IL18BP | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000137752 | CASP1 | ACC,BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,LUAD,OV,PAAD,SARC,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000137841 | PLCB2 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000140464 | PML | ACC,BRCA,COAD,GBM,KICH,KIRC,LGG,OV,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000141480 | ARRB2 | ACC,BRCA,CESC,ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000142347 | MYO1F | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000142512 | SIGLEC10 | ACC,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000147010 | SH3KBP1 | ACC,BRCA,CHOL,DLBC,ESCA,LUSC,PAAD | | LUSC | | ENSG00000280153 | ENSG00000147168 | IL2RG | ACC,BLCA,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LUSC,MESO,SARC,SKCM,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000148908 | RGS10 | ACC,CHOL,DLBC,KIRC,KIRP,LGG,PAAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000150977 | RILPL2 | ACC,BLCA,CHOL,COAD,GBM,LGG,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | LUSC | | ENSG00000280153 | ENSG00000153563 | CD8A | ACC,BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000158869 | FCER1G | ACC,BRCA,COAD,GBM,KICH,KIRC,KIRP,LGG,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000159403 | C1R | ACC,BRCA,GBM,KICH,LGG,SKCM,THCA | GBM | | | ENSG00000280153 | ENSG00000160255 | ITGB2 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000161921 | CXCL16 | ACC,CESC,ESCA,GBM,KICH,LGG,SARC,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000162511 | LAPTM5 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000162654 | GBP4 | ACC,BLCA,BRCA,COAD,ESCA,KIRC,KIRP,LAML,LGG,LUSC,SARC,SKCM,TGCT,THCA,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000163131 | CTSS | ACC,BLCA,BRCA,CESC,CHOL,ESCA,GBM,KICH,LGG,LUSC,SARC,SKCM,TGCT,THCA,THYM | GBM | LUSC | | ENSG00000280153 | ENSG00000163568 | AIM2 | ACC,BRCA,CHOL,COAD,KIRC,MESO,PAAD,SKCM,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000163823 | CCR1 | ACC,COAD,ESCA,GBM,KICH,LGG,PAAD,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000163840 | DTX3L | ACC,COAD,GBM,LAML,LGG,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000164308 | ERAP2 | ACC,CESC,GBM,LGG,SKCM,TGCT,THCA,UCEC | GBM | | | ENSG00000280153 | ENSG00000166278 | C2 | ACC,BLCA,BRCA,CESC,GBM,KICH,LGG,OV,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000167286 | CD3D | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000167604 | NFKBID | ACC,BLCA,BRCA,CHOL,COAD,GBM,KICH,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000168394 | TAP1 | ACC,BLCA,BRCA,CESC,COAD,ESCA,KIRC,LGG,OV,SARC,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000168404 | MLKL | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LUSC,PAAD,SARC,SKCM,TGCT,THCA,THYM,UCEC | KIRC | | | ENSG00000280153 | ENSG00000169403 | PTAFR | ACC,BRCA,COAD,GBM,KICH,KIRP,LGG,MESO,SARC,SKCM,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000169442 | CD52 | ACC,BLCA,CESC,CHOL,COAD,DLBC,ESCA,KICH,KIRC,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000170542 | SERPINB9 | ACC,BRCA,CHOL,COAD,GBM,KIRC,KIRP,LAML,LGG,PAAD,SARC,THCA | KIRC | | | ENSG00000280153 | ENSG00000177595 | PIDD1 | ACC,CHOL,DLBC,KICH,KIRC,PAAD | KIRC | | | ENSG00000280153 | ENSG00000177989 | ODF3B | ACC,BLCA,BRCA,COAD,GBM,KICH,KIRC,KIRP,LUSC,MESO,OV,SARC,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000178685 | PARP10 | ACC,BLCA,CESC,CHOL,GBM,KICH,KIRC,LGG,OV,PAAD,SARC,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000180096 | SEPTIN1 | ACC,BLCA,BRCA,CESC,CHOL,COAD,DLBC,ESCA,KICH,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000180644 | PRF1 | ACC,BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,PAAD,SARC,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000182179 | UBA7 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM | LUSC | | ENSG00000280153 | ENSG00000184730 | APOBR | ACC,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,PAAD,SKCM,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000184922 | FMNL1 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000185215 | TNFAIP2 | ACC,BRCA,COAD,GBM,KICH,KIRC,LGG,LUSC,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000188641 | DPYD | ACC,BRCA,COAD,GBM,LGG,OV,TGCT | GBM | | | ENSG00000280153 | ENSG00000188820 | CALHM6 | ACC,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,PAAD,SARC,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000196126 | HLA-DRB1 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000196954 | CASP4 | ACC,BRCA,GBM,KICH,KIRC,KIRP,PAAD,SKCM,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000198502 | HLA-DRB5 | ACC,BLCA,BRCA,CESC,COAD,ESCA,GBM,KICH,LAML,LGG,SARC,SKCM,THCA,THYM,UCEC | GBM | | | ENSG00000280153 | ENSG00000203747 | FCGR3A | ACC,COAD,GBM,KICH,KIRC,KIRP,LGG,SARC,SKCM,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000204252 | HLA-DOA | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000204257 | HLA-DMA | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000204264 | PSMB8 | ACC,BLCA,BRCA,CESC,ESCA,KIRC,LGG,OV,SARC,SKCM,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000204267 | TAP2 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | KIRC | | | ENSG00000280153 | ENSG00000204482 | LST1 | ACC,BLCA,BRCA,CESC,CHOL,ESCA,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000205220 | PSMB10 | ACC,BLCA,BRCA,CESC,CHOL,ESCA,GBM,KICH,KIRC,LGG,LUSC,OV,SARC,SKCM,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000214013 | GANC | ACC,KIRC,LUAD,LUSC,PAAD,TGCT | | LUSC | | ENSG00000280153 | ENSG00000231925 | TAPBP | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,LGG,OV,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | KIRC | | | ENSG00000280153 | ENSG00000240065 | PSMB9 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000005844 | ITGAL | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000006062 | MAP3K14 | BLCA,BRCA,CHOL,ESCA,KIRC,LAML,LGG,LUAD,LUSC,OV,PAAD,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000010295 | IFFO1 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000010671 | BTK | BLCA,BRCA,CHOL,COAD,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000069493 | CLEC2D | BLCA,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000073331 | ALPK1 | BLCA,BRCA,COAD,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,OV,PAAD,SARC,SKCM,STAD,THYM,UCEC | GBM | | | ENSG00000280153 | ENSG00000089012 | SIRPG | BLCA,BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000089820 | ARHGAP4 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000096996 | IL12RB1 | BLCA,COAD,KIRC,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000100385 | IL2RB | BLCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000101082 | SLA2 | BLCA,BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000101347 | SAMHD1 | BLCA,BRCA,CHOL,COAD,GBM,KICH,KIRC,LUSC,MESO,PAAD,SARC,SKCM,STAD,THCA,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000104814 | MAP4K1 | BLCA,BRCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000104951 | IL4I1 | BLCA,BRCA,CHOL,ESCA,GBM,KICH,SKCM,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000105122 | RASAL3 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000105246 | EBI3 | BLCA,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,MESO,PAAD,SARC,SKCM,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000105483 | CARD8 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT | GBM | LUSC | | ENSG00000280153 | ENSG00000105639 | JAK3 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000107099 | DOCK8 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000110934 | BIN2 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000111679 | PTPN6 | BLCA,BRCA,CESC,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000113088 | GZMK | BLCA,BRCA,CHOL,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000115165 | CYTIP | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000115956 | PLEK | BLCA,BRCA,CHOL,COAD,GBM,KICH,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA,THYM | GBM | LUSC | | ENSG00000280153 | ENSG00000117091 | CD48 | BLCA,BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000122986 | HVCN1 | BLCA,BRCA,CHOL,ESCA,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000123329 | ARHGAP9 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LAML,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000123338 | NCKAP1L | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000126246 | IGFLR1 | BLCA,CESC,CHOL,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,SARC,SKCM,TGCT | GBM,KIRC | | | ENSG00000280153 | ENSG00000131378 | RFTN1 | BLCA,BRCA,CHOL,COAD,GBM,LGG,LUSC,OV,PAAD,SKCM,TGCT | | LUSC | | ENSG00000280153 | ENSG00000136286 | MYO1G | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000137078 | SIT1 | BLCA,BRCA,CHOL,ESCA,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000137101 | CD72 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LAML,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000138964 | PARVG | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000139193 | CD27 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000139597 | N4BP2L1 | BLCA,BRCA,CESC,ESCA,KIRC,LUAD,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000140379 | BCL2A1 | BLCA,CHOL,KICH,LGG,LUSC,PAAD,TGCT,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000141968 | VAV1 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000142102 | PGGHG | BLCA,CHOL,GBM,KIRC,PAAD,SKCM,TGCT | GBM,KIRC | | | ENSG00000280153 | ENSG00000143851 | PTPN7 | BLCA,BRCA,CESC,CHOL,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000146094 | DOK3 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LAML,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000149781 | FERMT3 | BLCA,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000155307 | SAMSN1 | BLCA,BRCA,CHOL,COAD,ESCA,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000155629 | PIK3AP1 | BLCA,BRCA,COAD,GBM,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000157873 | TNFRSF14 | BLCA,BRCA,CESC,CHOL,ESCA,GBM,KIRC,LGG,LUAD,LUSC,OV,PAAD,SARC,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000158062 | UBXN11 | BLCA,CESC,CHOL,DLBC,GBM,KIRC,LUAD,LUSC,MESO,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000162739 | SLAMF6 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000166710 | B2M | BLCA,BRCA,CESC,ESCA,LGG,LUSC,SARC,SKCM,TGCT,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000167895 | TMC8 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,UCEC | KIRC | | | ENSG00000280153 | ENSG00000172215 | CXCR6 | BLCA,BRCA,COAD,ESCA,KIRC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000175309 | PHYKPL | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,UCEC | KIRC | | | ENSG00000280153 | ENSG00000175463 | TBC1D10C | BLCA,BRCA,CESC,CHOL,COAD,DLBC,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000177000 | MTHFR | BLCA,CHOL,COAD,DLBC,LGG,LUAD,LUSC,MESO,PAAD,SARC | | LUSC | | ENSG00000280153 | ENSG00000179583 | CIITA | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000180353 | HCLS1 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000180448 | ARHGAP45 | BLCA,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000186517 | ARHGAP30 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000186810 | CXCR3 | BLCA,BRCA,CHOL,ESCA,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000188389 | PDCD1 | BLCA,BRCA,CHOL,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000196684 | HSH2D | BLCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SKCM,STAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000198771 | RCSD1 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LAML,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000198821 | CD247 | BLCA,BRCA,CHOL,COAD,DLBC,KIRC,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000204592 | HLA-E | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM,KIRC | | | ENSG00000280153 | ENSG00000213445 | SIPA1 | BLCA,BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000213886 | UBD | BLCA,BRCA,COAD,ESCA,KIRC,SARC,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000214212 | C19orf38 | BLCA,KIRC,LGG,PAAD,SKCM,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000232629 | HLA-DQB2 | BLCA,BRCA,CESC,CHOL,ESCA,GBM,KIRC,KIRP,LAML,LGG,OV,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000234745 | HLA-B | BLCA,BRCA,CESC,CHOL,ESCA,KIRC,LGG,PAAD,SARC,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000000938 | FGR | BRCA,CHOL,COAD,KICH,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THCA,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000023445 | BIRC3 | BRCA,COAD,ESCA,KIRC,LUSC,OV,PAAD,SKCM,STAD,TGCT,THCA,THYM | KIRC | LUSC | | ENSG00000280153 | ENSG00000023902 | PLEKHO1 | BRCA,CHOL,KICH,KIRC,KIRP,LAML,LGG,LUSC,PAAD,SKCM,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000027869 | SH2D2A | BRCA,CHOL,KIRC,KIRP,PAAD,SARC,SKCM,TGCT | KIRC | | | ENSG00000280153 | ENSG00000051523 | CYBA | BRCA,CHOL,ESCA,GBM,LGG,MESO,SARC,SKCM,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000065357 | DGKA | BRCA,CHOL,GBM,KIRC,KIRP,MESO,PAAD,SARC,SKCM,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000065413 | ANKRD44 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | LUSC | | ENSG00000280153 | ENSG00000072786 | STK10 | BRCA,CHOL,COAD,KICH,KIRC,KIRP,PAAD,STAD,THCA | KIRC | | | ENSG00000280153 | ENSG00000076641 | PAG1 | BRCA,CHOL,ESCA,KIRC,LAML,PAAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000077238 | IL4R | BRCA,GBM,KICH,KIRC,LGG,LUSC,PAAD,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000082074 | FYB1 | BRCA,CHOL,COAD,GBM,KICH,KIRC,KIRP,LGG,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000089639 | GMIP | BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000091592 | NLRP1 | BRCA,CHOL,KIRC,LUAD,LUSC,MESO,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000093072 | ADA2 | BRCA,CHOL,COAD,ESCA,GBM,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000099331 | MYO9B | BRCA,CHOL,COAD,KICH,KIRC,KIRP,LGG,LUAD,LUSC,PAAD,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000100060 | MFNG | BRCA,CHOL,KICH,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000100079 | LGALS2 | BRCA,LUSC,MESO,SKCM,THCA | | LUSC | | ENSG00000280153 | ENSG00000100288 | CHKB | BRCA,CHOL,COAD,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000100365 | NCF4 | BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SKCM,TGCT,THCA,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000101265 | RASSF2 | BRCA,CHOL,COAD,KICH,KIRP,LUAD,LUSC,MESO,PAAD,STAD,TGCT,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000104774 | MAN2B1 | BRCA,GBM,LGG,LUSC,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000105643 | ARRDC2 | BRCA,DLBC,ESCA,KICH,KIRP,LAML,LUSC,MESO,SARC,SKCM,STAD,TGCT,UCEC | | LUSC | | ENSG00000280153 | ENSG00000106952 | TNFSF8 | BRCA,GBM,KIRC,PAAD,STAD | GBM,KIRC | | | ENSG00000280153 | ENSG00000108622 | ICAM2 | BRCA,CHOL,LUSC,PAAD,SKCM | | LUSC | | ENSG00000280153 | ENSG00000108950 | FAM20A | BRCA,GBM,KIRP,LUAD,THCA | GBM | | | ENSG00000280153 | ENSG00000110077 | MS4A6A | BRCA,ESCA,GBM,KICH,KIRC,KIRP,LGG,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000110848 | CD69 | BRCA,CHOL,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000111252 | SH2B3 | BRCA,COAD,ESCA,GBM,LGG,LUSC,PAAD,STAD,THYM | | LUSC | | ENSG00000280153 | ENSG00000113240 | CLK4 | BRCA,CHOL,COAD,DLBC,ESCA,KIRC,KIRP,PAAD,SKCM,STAD,TGCT,UCEC | KIRC | | | ENSG00000280153 | ENSG00000117616 | RSRP1 | BRCA,CHOL,ESCA,GBM,KIRC,LGG,LUAD,LUSC,OV,PAAD,SKCM,STAD,TGCT,UCEC | KIRC | | | ENSG00000280153 | ENSG00000118503 | TNFAIP3 | BRCA,CHOL,COAD,KIRC,PAAD,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000120280 | TASL | BRCA,GBM,KIRC,LGG,LUSC,MESO,PAAD,SKCM,STAD,TGCT,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000123609 | NMI | BRCA,CESC,ESCA,KIRC,LGG,SKCM,THCA | KIRC | | | ENSG00000280153 | ENSG00000125730 | C3 | BRCA,CESC,GBM,LGG,LUSC,SKCM,THCA,UCEC | GBM | LUSC | | ENSG00000280153 | ENSG00000130755 | GMFG | BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000131042 | LILRB2 | BRCA,COAD,ESCA,GBM,KICH,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000132514 | CLEC10A | BRCA,CHOL,LUSC,MESO,PAAD,SARC,SKCM,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000132522 | GPS2 | BRCA,CHOL,GBM,KIRC,LUAD,PAAD,SKCM,TGCT | KIRC | | | ENSG00000280153 | ENSG00000132530 | XAF1 | BRCA,COAD,GBM,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000133106 | EPSTI1 | BRCA,ESCA,KIRC,LGG,LUSC,SARC,SKCM,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000133561 | GIMAP6 | BRCA,CHOL,COAD,ESCA,GBM,LGG,PAAD,SARC,SKCM,STAD,TGCT | GBM | | | ENSG00000280153 | ENSG00000135596 | MICAL1 | BRCA,CHOL,DLBC,ESCA,KICH,KIRC,KIRP,PAAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000135905 | DOCK10 | BRCA,CHOL,ESCA,KIRC,KIRP,LUAD,LUSC,PAAD,SARC,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000136167 | LCP1 | BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000136250 | AOAH | BRCA,CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000141497 | ZMYND15 | BRCA,CESC,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,OV,SARC,TGCT,THCA,THYM,UCEC | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000141506 | PIK3R5 | BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000142634 | EFHD2 | BRCA,CHOL,KICH,KIRC,KIRP,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000143185 | XCL2 | BRCA,KIRC,SARC,SKCM,STAD,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000144843 | ADPRH | BRCA,CHOL,COAD,LGG,LUSC,PAAD,SARC,TGCT,THYM | | LUSC | | ENSG00000280153 | ENSG00000146112 | PPP1R18 | BRCA,CHOL,KIRC,KIRP,LGG,PAAD,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000154237 | LRRK1 | BRCA,KIRC,LAML,LUAD,LUSC,PAAD,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000155465 | SLC7A7 | BRCA,ESCA,GBM,LGG,LUSC,SKCM,THCA,THYM | GBM | LUSC | | ENSG00000280153 | ENSG00000155962 | CLIC2 | BRCA,CHOL,COAD,ESCA,KICH,LGG,LUSC,PAAD,SKCM,STAD,TGCT,THYM | | LUSC | | ENSG00000280153 | ENSG00000159189 | C1QC | BRCA,GBM,KICH,LGG,SKCM,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000160883 | HK3 | BRCA,GBM,KIRC,KIRP,PAAD,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000161791 | FMNL3 | BRCA,CHOL,COAD,KIRC,KIRP,LUAD,LUSC,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000161929 | SCIMP | BRCA,CHOL,DLBC,GBM,KIRC,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000167208 | SNX20 | BRCA,CHOL,GBM,KIRC,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000167615 | LENG8 | BRCA,CHOL,ESCA,KICH,KIRC,LUAD,LUSC,MESO,PAAD,SKCM,STAD | KIRC | | | ENSG00000280153 | ENSG00000167984 | NLRC3 | BRCA,CESC,DLBC,ESCA,KIRC,LUAD,LUSC,PAAD,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000168016 | TRANK1 | BRCA,CHOL,ESCA,GBM,KIRC,LUAD,LUSC,PAAD,SKCM,STAD,UCEC | | LUSC | | ENSG00000280153 | ENSG00000173369 | C1QB | BRCA,GBM,KICH,KIRP,LGG,SKCM,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000173372 | C1QA | BRCA,CHOL,COAD,GBM,KICH,KIRC,KIRP,LGG,MESO,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000173442 | EHBP1L1 | BRCA,CHOL,COAD,GBM,KICH,KIRC,KIRP,LGG,OV,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000174125 | TLR1 | BRCA,GBM,LGG,MESO,PAAD,SARC,SKCM,STAD,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000174600 | CMKLR1 | BRCA,CHOL,COAD,ESCA,KICH,LUSC,MESO,TGCT,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000175489 | LRRC25 | BRCA,CHOL,GBM,KICH,KIRP,LGG,LUSC,PAAD,TGCT,THCA,THYM | GBM | LUSC | | ENSG00000280153 | ENSG00000184574 | LPAR5 | BRCA,CHOL,KIRC,KIRP,LGG,SARC,SKCM | KIRC | | | ENSG00000280153 | ENSG00000184584 | STING1 | BRCA,CHOL,GBM,KICH,KIRP,LGG,OV,THCA,UCEC | GBM | | | ENSG00000280153 | ENSG00000185905 | C16orf54 | BRCA,CHOL,KIRC,LUSC,MESO,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000186166 | CENATAC | BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LUAD,LUSC,PAAD,SARC,SKCM,STAD | KIRC | | | ENSG00000280153 | ENSG00000186818 | LILRB4 | BRCA,CHOL,COAD,GBM,KICH,KIRC,KIRP,LGG,SARC,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000187688 | TRPV2 | BRCA,CHOL,KIRP,LGG,LUSC,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000197540 | GZMM | BRCA,CHOL,COAD,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000197948 | FCHSD1 | BRCA,CHOL,DLBC,GBM,KICH,KIRC,LGG,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000204149 | AGAP6 | BRCA,CHOL,ESCA,KIRC,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000204305 | AGER | BRCA,CHOL,KIRC,KIRP,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000215252 | GOLGA8B | BRCA,COAD,ESCA,KIRC,LUAD,LUSC,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000235568 | NFAM1 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SKCM,TGCT,THYM | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000243646 | IL10RB | BRCA,GBM,KIRP,LGG,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000266094 | RASSF5 | BRCA,CHOL,COAD,KICH,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000100298 | APOBEC3H | CESC,KIRC,LUSC,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000104518 | GSDMD | CESC,CHOL,GBM,LGG,OV,SARC,THCA | GBM | | | ENSG00000280153 | ENSG00000162777 | DENND2D | CESC,CHOL,ESCA,GBM,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000167083 | GNGT2 | CESC,KIRC,LUSC,MESO,PAAD,SKCM,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000169592 | INO80E | CESC,CHOL,DLBC,KICH,KIRC,LUAD,LUSC,PAAD | KIRC | | | ENSG00000280153 | ENSG00000013441 | CLK1 | CHOL,ESCA,KIRC,KIRP,LUAD,LUSC,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000059377 | TBXAS1 | CHOL,GBM,KICH,KIRC,LGG,SARC,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000060491 | OGFR | CHOL,ESCA,KICH,KIRC,KIRP,LUSC,TGCT | KIRC | | | ENSG00000280153 | ENSG00000061938 | TNK2 | CHOL,KICH,KIRC,KIRP,LUAD,MESO,SKCM,THCA | KIRC | | | ENSG00000280153 | ENSG00000064012 | CASP8 | CHOL,COAD,ESCA,GBM,KIRC,OV,PAAD,SKCM,THCA | GBM | | | ENSG00000280153 | ENSG00000068028 | RASSF1 | CHOL,KICH,KIRC,KIRP,LGG,LUAD,LUSC,PAAD,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000072609 | CHFR | CHOL,KIRC,KIRP,PAAD,THCA | KIRC | | | ENSG00000280153 | ENSG00000100429 | HDAC10 | CHOL,GBM,KIRC,KIRP,LUAD,MESO,OV,PAAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000104365 | IKBKB | CHOL,GBM,KIRC,LGG,OV,PAAD,SARC,STAD,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000108773 | KAT2A | CHOL,DLBC,KICH,KIRC,PAAD,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000113108 | APBB3 | CHOL,KICH,KIRC,OV,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000114353 | GNAI2 | CHOL,KICH,LGG,LUSC,SARC,TGCT,THCA | | LUSC | | ENSG00000280153 | ENSG00000114779 | ABHD14B | CHOL,DLBC,GBM,LGG,MESO,TGCT | GBM | | | ENSG00000280153 | ENSG00000115325 | DOK1 | CHOL,ESCA,KICH,KIRC,KIRP,LGG,LUSC,PAAD,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000115604 | IL18R1 | CHOL,ESCA,LUSC,PAAD,STAD,THYM | | LUSC | | ENSG00000280153 | ENSG00000123136 | DDX39A | CHOL,KICH,KIRC,PAAD,THCA | KIRC | | | ENSG00000280153 | ENSG00000124126 | PREX1 | CHOL,COAD,ESCA,KICH,KIRP,LGG,LUSC,MESO,PAAD,SARC,STAD,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000129465 | RIPK3 | CHOL,KIRC,LUAD,LUSC,OV,SKCM,TGCT | | LUSC | | ENSG00000280153 | ENSG00000129675 | ARHGEF6 | CHOL,COAD,ESCA,LAML,LUAD,LUSC,MESO,PAAD,SKCM,STAD,THCA | | LUSC | | ENSG00000280153 | ENSG00000131171 | SH3BGRL | CHOL,ESCA,LUSC,SKCM,TGCT | | LUSC | | ENSG00000280153 | ENSG00000135723 | FHOD1 | CHOL,GBM,KICH,KIRC,LGG,LUAD,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000138834 | MAPK8IP3 | CHOL,ESCA,KIRC,LAML,LUAD,PAAD,STAD | KIRC | | | ENSG00000280153 | ENSG00000139636 | LMBR1L | CHOL,KIRC,KIRP,LUSC,PAAD | KIRC | | | ENSG00000280153 | ENSG00000140511 | HAPLN3 | CHOL,COAD,KIRC,KIRP,LUSC,MESO,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000142185 | TRPM2 | CHOL,KIRC,KIRP,LGG,PAAD,SARC,THYM | KIRC | | | ENSG00000280153 | ENSG00000148288 | GBGT1 | CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD,THCA | GBM | | | ENSG00000280153 | ENSG00000160271 | RALGDS | CHOL,KICH,KIRC,KIRP,SKCM,TGCT,THYM | KIRC | | | ENSG00000280153 | ENSG00000161010 | MRNIP | CHOL,ESCA,GBM,KIRC,PAAD,SKCM,STAD,THYM | KIRC | | | ENSG00000280153 | ENSG00000163644 | PPM1K | CHOL,ESCA,LUAD,LUSC,PAAD,STAD,TGCT | | LUSC | | ENSG00000280153 | ENSG00000164430 | CGAS | CHOL,KIRC,SKCM,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000167261 | DPEP2 | CHOL,GBM,KIRC,KIRP,LUSC,MESO,PAAD,SARC,TGCT | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000170571 | EMB | CHOL,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SKCM,STAD,THYM | KIRC | | | ENSG00000280153 | ENSG00000171700 | RGS19 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000174175 | SELP | CHOL,LUSC,MESO,SARC,SKCM,THCA | | LUSC | | ENSG00000280153 | ENSG00000174943 | KCTD13 | CHOL,DLBC,ESCA,KIRC,OV,TGCT | KIRC | | | ENSG00000280153 | ENSG00000175265 | GOLGA8A | CHOL,ESCA,KIRC,LUAD,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000176533 | GNG7 | CHOL,LAML,LUAD,LUSC,MESO,PAAD | | LUSC | | ENSG00000280153 | ENSG00000179715 | PCED1B | CHOL,GBM,KIRC,KIRP,LGG,LUSC,MESO,PAAD,SARC,SKCM,STAD,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000181481 | RNF135 | CHOL,DLBC,GBM,KIRC,LGG,TGCT | GBM | | | ENSG00000280153 | ENSG00000183160 | TMEM119 | CHOL,DLBC,KICH,LGG,LUSC,MESO,THCA,THYM | | LUSC | | ENSG00000280153 | ENSG00000185101 | ANO9 | CHOL,KIRC,KIRP,PAAD,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000185482 | STAC3 | CHOL,COAD,GBM,KIRC,KIRP,LGG,LUSC,OV,PAAD,SKCM,TGCT,THCA | KIRC | LUSC | | ENSG00000280153 | ENSG00000186088 | GSAP | CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THYM,UCEC | KIRC | | | ENSG00000280153 | ENSG00000186891 | TNFRSF18 | CHOL,KIRC,MESO,PAAD,SKCM,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000188404 | SELL | CHOL,COAD,ESCA,GBM,LAML,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT,THCA | GBM | LUSC | | ENSG00000280153 | ENSG00000197774 | EME2 | CHOL,DLBC,KICH,KIRC,LUAD,OV,PAAD,SKCM,STAD,THYM | KIRC | | | ENSG00000280153 | ENSG00000197872 | CYRIA | CHOL,COAD,LUSC,PAAD,SKCM,STAD,TGCT | | LUSC | | ENSG00000280153 | ENSG00000205268 | PDE7A | CHOL,COAD,KIRC,PAAD,SKCM,STAD | KIRC | | | ENSG00000280153 | ENSG00000205436 | EXOC3L4 | CHOL,ESCA,KIRC,STAD,THYM | KIRC | | | ENSG00000280153 | ENSG00000214021 | TTLL3 | CHOL,ESCA,GBM,KIRC,LUAD,LUSC,OV,PAAD,SKCM,STAD,UCEC | KIRC | | | ENSG00000280153 | ENSG00000215375 | MYL5 | CHOL,DLBC,GBM,KICH,KIRC,KIRP,LUAD | KIRC | | | ENSG00000280153 | ENSG00000221978 | CCNL2 | CHOL,KICH,KIRC,LUAD,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000256525 | POLG2 | CHOL,KIRC,LUAD,PAAD,STAD | KIRC | | | ENSG00000280153 | ENSG00000273899 | NOL12 | CHOL,KICH,KIRC,KIRP,LUAD,MESO,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000066294 | CD84 | COAD,ESCA,GBM,KIRC,LGG,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT,THCA | GBM,KIRC | | | ENSG00000280153 | ENSG00000071246 | VASH1 | COAD,ESCA,KICH,KIRP,LUSC | | LUSC | | ENSG00000280153 | ENSG00000088827 | SIGLEC1 | COAD,GBM,KIRC,SKCM,TGCT,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000109046 | WSB1 | COAD,ESCA,KIRC,PAAD,SKCM | KIRC | | | ENSG00000280153 | ENSG00000115525 | ST3GAL5 | COAD,LAML,LUSC,PAAD,STAD,TGCT | | LUSC | | ENSG00000280153 | ENSG00000118855 | MFSD1 | COAD,GBM,LGG,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000139631 | CSAD | COAD,KIRC,LUAD,TGCT,UCEC | KIRC | | | ENSG00000280153 | ENSG00000145685 | LHFPL2 | COAD,DLBC,GBM,KICH,LGG | GBM | | | ENSG00000280153 | ENSG00000161960 | EIF4A1 | COAD,KIRC,LUAD,PAAD,THYM | KIRC | | | ENSG00000280153 | ENSG00000169245 | CXCL10 | COAD,KIRC,KIRP,LGG,SARC,SKCM,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000172243 | CLEC7A | COAD,GBM,KIRC,KIRP,LGG,SARC,SKCM,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000184060 | ADAP2 | COAD,GBM,KICH,KIRC,KIRP,LGG,SARC,SKCM,STAD,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000184988 | TMEM106A | COAD,ESCA,GBM,LAML,LGG,LUSC,PAAD,STAD | GBM | | | ENSG00000280153 | ENSG00000267534 | S1PR2 | COAD,KICH,KIRP,LUSC,PAAD,THCA | | LUSC | | ENSG00000280153 | ENSG00000157514 | TSC22D3 | DLBC,LUSC,PAAD,SKCM,TGCT | | LUSC | | ENSG00000280153 | ENSG00000164877 | MICALL2 | DLBC,ESCA,GBM,KICH,KIRC,KIRP,LGG,SARC,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000175938 | ORAI3 | DLBC,GBM,KICH,LGG,TGCT | GBM | | | ENSG00000280153 | ENSG00000213903 | LTB4R | DLBC,KIRC,KIRP,LUAD,PAAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000099377 | HSD3B7 | ESCA,GBM,KICH,LGG,TGCT,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000117226 | GBP3 | ESCA,GBM,LAML,LGG,OV,SKCM,THYM | GBM | | | ENSG00000280153 | ENSG00000119535 | CSF3R | ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000127415 | IDUA | ESCA,GBM,KICH,KIRC,LUSC | KIRC | | | ENSG00000280153 | ENSG00000139508 | SLC46A3 | ESCA,GBM,LGG,LUSC,SKCM,TGCT | | LUSC | | ENSG00000280153 | ENSG00000145476 | CYP4V2 | ESCA,GBM,LUAD,LUSC,TGCT | | LUSC | | ENSG00000280153 | ENSG00000006534 | ALDH3B1 | GBM,LGG,SARC,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000007129 | CEACAM21 | GBM,KIRC,LAML,LUSC,MESO,PAAD,SARC,SKCM,TGCT | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000026508 | CD44 | GBM,KICH,LGG,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000062716 | VMP1 | GBM,KIRC,LGG,LUAD,SARC,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000092929 | UNC13D | GBM,KIRC,LUSC,SARC,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000125753 | VASP | GBM,KICH,KIRP,LGG,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000129450 | SIGLEC9 | GBM,KIRC,KIRP,LGG,TGCT | GBM,KIRC | | | ENSG00000280153 | ENSG00000133048 | CHI3L1 | GBM,KICH,KIRP,TGCT,THYM | GBM | | | ENSG00000280153 | ENSG00000133246 | PRAM1 | GBM,KIRC,KIRP,LGG,LUSC,TGCT | KIRC | LUSC | | ENSG00000280153 | ENSG00000139178 | C1RL | GBM,LGG,SARC,SKCM,THCA | GBM | | | ENSG00000280153 | ENSG00000152229 | PSTPIP2 | GBM,KIRC,KIRP,PAAD,SKCM,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000160219 | GAB3 | GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,SARC,STAD,TGCT,THCA | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000163162 | RNF149 | GBM,KICH,KIRC,LGG,TGCT,THCA | KIRC | | | ENSG00000280153 | ENSG00000163545 | NUAK2 | GBM,PAAD,SARC,SKCM,THYM | GBM | | | ENSG00000280153 | ENSG00000170909 | OSCAR | GBM,KIRC,KIRP,LGG,TGCT,THCA,THYM | GBM,KIRC | | | ENSG00000280153 | ENSG00000171860 | C3AR1 | GBM,KICH,LGG,SARC,TGCT,THCA | GBM | | | ENSG00000280153 | ENSG00000173221 | GLRX | GBM,LGG,LUSC,SKCM,TGCT,THCA | GBM | LUSC | | ENSG00000280153 | ENSG00000182511 | FES | GBM,KICH,KIRP,LGG,LUSC,OV,PAAD,SARC,TGCT | | LUSC | | ENSG00000280153 | ENSG00000187554 | TLR5 | GBM,KIRP,LGG,LUAD,SARC,THCA,THYM | GBM | | | ENSG00000280153 | ENSG00000196209 | SIRPB2 | GBM,KIRC,LUSC,MESO,SARC | GBM,KIRC | LUSC | | ENSG00000280153 | ENSG00000134285 | FKBP11 | KICH,KIRC,TGCT,THCA,THYM | KIRC | | | ENSG00000280153 | ENSG00000135722 | FBXL8 | KICH,KIRC,LUAD,MESO,OV | KIRC | | | ENSG00000280153 | ENSG00000113263 | ITK | KIRC,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000135439 | AGAP2 | KIRC,LUAD,PAAD,SKCM,STAD,TGCT | KIRC | | | ENSG00000280153 | ENSG00000142303 | ADAMTS10 | KIRC,KIRP,LAML,LUAD,STAD | KIRC | | | ENSG00000280153 | ENSG00000172578 | KLHL6 | KIRC,LAML,LUAD,LUSC,PAAD,SKCM,STAD,TGCT | KIRC | LUSC | | ENSG00000280153 | ENSG00000163600 | ICOS | LUAD,LUSC,SKCM,STAD,TGCT | | LUSC |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000134186 | PRPF38B | 5 | CHOL,KIRC,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000020633 | RUNX3 | 5 | CHOL,KIRC,PAAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000020633 | RUNX3 | 8 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LUSC,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000020633 | RUNX3 | 10 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LUSC,PAAD,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000020633 | RUNX3 | 9 | BLCA,BRCA,CHOL,KIRC,KIRP,PAAD,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000020633 | RUNX3 | 6 | KIRC,LUSC,PAAD,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000065357 | DGKA | 7 | BRCA,CHOL,KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000065357 | DGKA | 6 | BRCA,CHOL,KIRC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000065357 | DGKA | 6 | BRCA,CHOL,KIRC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000065357 | DGKA | 6 | CHOL,KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000099308 | MAST3 | 6 | CHOL,ESCA,LUSC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000204287 | HLA-DRA | 13 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000204287 | HLA-DRA | 7 | BLCA,BRCA,CHOL,ESCA,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000204287 | HLA-DRA | 13 | BLCA,BRCA,CHOL,ESCA,KICH,KIRC,KIRP,LAML,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000204287 | HLA-DRA | 18 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,OV,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000204287 | HLA-DRA | 9 | BRCA,CHOL,ESCA,KIRC,LAML,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000204287 | HLA-DRA | 14 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,LAML,OV,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000168071 | CCDC88B | 7 | CHOL,GBM,LGG,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000168071 | CCDC88B | 10 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000168071 | CCDC88B | 11 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000168071 | CCDC88B | 9 | BRCA,CHOL,KIRC,KIRP,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000168071 | CCDC88B | 6 | CHOL,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000168071 | CCDC88B | 6 | CHOL,KIRC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000177455 | CD19 | 5 | CHOL,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000177455 | CD19 | 5 | BLCA,CHOL,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000177455 | CD19 | 9 | BLCA,CHOL,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000177455 | CD19 | 5 | CHOL,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000101082 | SLA2 | 10 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000101082 | SLA2 | 7 | BLCA,BRCA,CHOL,ESCA,KIRC,LUSC,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000101082 | SLA2 | 11 | BLCA,BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000101082 | SLA2 | 10 | BLCA,BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000101082 | SLA2 | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000101082 | SLA2 | 8 | BRCA,CHOL,KIRC,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000096996 | IL12RB1 | 7 | KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000096996 | IL12RB1 | 5 | BLCA,KIRC,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000096996 | IL12RB1 | 10 | BLCA,COAD,KIRC,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000096996 | IL12RB1 | 9 | BLCA,COAD,KIRC,LUSC,PAAD,SARC,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000096996 | IL12RB1 | 6 | KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000096996 | IL12RB1 | 6 | KIRC,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000121858 | TNFSF10 | 5 | LGG,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000132530 | XAF1 | 10 | BRCA,COAD,GBM,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000132530 | XAF1 | 7 | BRCA,KIRC,LUAD,LUSC,PAAD,SARC,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000132530 | XAF1 | 10 | BRCA,COAD,GBM,KIRC,LUSC,PAAD,SARC,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000132530 | XAF1 | 8 | BRCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000132530 | XAF1 | 7 | BRCA,COAD,KIRC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000138185 | ENTPD1 | 5 | CHOL,COAD,PAAD,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000138185 | ENTPD1 | 5 | CHOL,ESCA,PAAD,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000091073 | DTX2 | 5 | CHOL,KIRC,KIRP,LGG,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000163840 | DTX3L | 5 | COAD,GBM,LGG,TGCT,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000163840 | DTX3L | 7 | ACC,COAD,GBM,LGG,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000129003 | VPS13C | 5 | COAD,ESCA,LUSC,SKCM,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000129003 | VPS13C | 5 | ESCA,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000129003 | VPS13C | 5 | LUAD,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000023445 | BIRC3 | 9 | BRCA,COAD,KIRC,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000023445 | BIRC3 | 6 | BRCA,ESCA,KIRC,LUSC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000023445 | BIRC3 | 7 | BRCA,ESCA,KIRC,LUSC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000023445 | BIRC3 | 11 | BRCA,COAD,ESCA,KIRC,LUSC,OV,PAAD,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000023445 | BIRC3 | 5 | KIRC,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000023445 | BIRC3 | 8 | BRCA,KIRC,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000162739 | SLAMF6 | 11 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000162739 | SLAMF6 | 7 | BLCA,BRCA,CHOL,KIRC,KIRP,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000162739 | SLAMF6 | 14 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000162739 | SLAMF6 | 13 | BLCA,BRCA,CHOL,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000162739 | SLAMF6 | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000162739 | SLAMF6 | 7 | KIRC,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000150977 | RILPL2 | 8 | CHOL,GBM,LGG,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000150977 | RILPL2 | 7 | BLCA,CHOL,GBM,LGG,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000150977 | RILPL2 | 9 | BLCA,CHOL,GBM,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000150977 | RILPL2 | 9 | ACC,BLCA,CHOL,COAD,LGG,LUSC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000150977 | RILPL2 | 5 | CHOL,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000000938 | FGR | 7 | CHOL,COAD,LUSC,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000000938 | FGR | 5 | BRCA,CHOL,KIRC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000000938 | FGR | 10 | BRCA,CHOL,KICH,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000000938 | FGR | 10 | BRCA,CHOL,COAD,KIRC,LUSC,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000000938 | FGR | 6 | CHOL,KIRC,LUSC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000141480 | ARRB2 | 9 | BRCA,ESCA,GBM,KIRC,KIRP,LUSC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000141480 | ARRB2 | 8 | ACC,BRCA,KIRC,LUSC,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000141480 | ARRB2 | 5 | KIRC,LUSC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000117215 | PLA2G2D | 6 | BRCA,CHOL,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000117215 | PLA2G2D | 5 | BLCA,BRCA,CHOL,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000117215 | PLA2G2D | 9 | BLCA,BRCA,CHOL,COAD,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000117215 | PLA2G2D | 8 | BRCA,CHOL,COAD,PAAD,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000117215 | PLA2G2D | 6 | BRCA,CHOL,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000117215 | PLA2G2D | 6 | CHOL,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000073331 | ALPK1 | 10 | BRCA,COAD,ESCA,GBM,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000073331 | ALPK1 | 5 | GBM,KIRC,KIRP,LUSC,PAAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000073331 | ALPK1 | 7 | ESCA,GBM,KIRC,LUAD,LUSC,SARC,SKCM | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000073331 | ALPK1 | 5 | ESCA,KIRC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000073331 | ALPK1 | 8 | ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000073331 | ALPK1 | 7 | KIRC,LUAD,LUSC,OV,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000171791 | BCL2 | 6 | CHOL,COAD,MESO,PAAD,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000136250 | AOAH | 10 | BRCA,CHOL,GBM,KIRC,LGG,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000136250 | AOAH | 6 | BRCA,CHOL,GBM,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000136250 | AOAH | 11 | BRCA,CHOL,GBM,KICH,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000136250 | AOAH | 10 | BRCA,CHOL,KIRC,KIRP,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000136250 | AOAH | 7 | BRCA,CHOL,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000136250 | AOAH | 6 | KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000134516 | DOCK2 | 13 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000134516 | DOCK2 | 12 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000134516 | DOCK2 | 12 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,LGG,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000134516 | DOCK2 | 8 | BRCA,CHOL,ESCA,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000134516 | DOCK2 | 8 | CHOL,COAD,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000153563 | CD8A | 12 | BRCA,CHOL,COAD,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000153563 | CD8A | 13 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000153563 | CD8A | 14 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000153563 | CD8A | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000153563 | CD8A | 7 | COAD,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000178971 | CTC1 | 10 | CHOL,COAD,KIRC,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000178971 | CTC1 | 6 | CHOL,KIRC,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000178971 | CTC1 | 5 | CHOL,KIRC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000196684 | HSH2D | 9 | CHOL,COAD,ESCA,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000196684 | HSH2D | 8 | CHOL,ESCA,KIRC,KIRP,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000196684 | HSH2D | 11 | BLCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000196684 | HSH2D | 11 | CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000196684 | HSH2D | 7 | CHOL,ESCA,KIRC,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000196684 | HSH2D | 7 | CHOL,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000005059 | MCUB | 7 | CHOL,COAD,LGG,PAAD,SKCM,STAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000005059 | MCUB | 6 | BLCA,CHOL,KICH,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000005059 | MCUB | 7 | BLCA,CHOL,COAD,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000164308 | ERAP2 | 5 | GBM,LGG,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000164308 | ERAP2 | 7 | ACC,CESC,GBM,LGG,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000175463 | TBC1D10C | 8 | BLCA,BRCA,CHOL,KIRC,KIRP,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000175463 | TBC1D10C | 11 | BLCA,BRCA,CHOL,COAD,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000175463 | TBC1D10C | 12 | BLCA,BRCA,CESC,CHOL,COAD,KIRC,KIRP,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000175463 | TBC1D10C | 7 | BRCA,CHOL,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000175463 | TBC1D10C | 5 | KIRC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000126246 | IGFLR1 | 9 | BLCA,CHOL,KIRC,KIRP,LUAD,LUSC,MESO,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000126246 | IGFLR1 | 8 | BLCA,CESC,KIRC,KIRP,LGG,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000126246 | IGFLR1 | 5 | CHOL,KIRC,LUSC,SARC,SKCM | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000167604 | NFKBID | 8 | BLCA,BRCA,CHOL,KIRC,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000167604 | NFKBID | 11 | BLCA,BRCA,CHOL,COAD,KICH,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000167604 | NFKBID | 8 | ACC,BRCA,CHOL,KIRC,LUSC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000167604 | NFKBID | 5 | CHOL,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000196735 | HLA-DQA1 | 13 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000196735 | HLA-DQA1 | 8 | BLCA,BRCA,CHOL,ESCA,GBM,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000196735 | HLA-DQA1 | 15 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LAML,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000196735 | HLA-DQA1 | 18 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,OV,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000196735 | HLA-DQA1 | 8 | CHOL,ESCA,KIRC,LAML,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000196735 | HLA-DQA1 | 13 | BLCA,BRCA,CESC,COAD,ESCA,KIRC,LAML,OV,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000160271 | RALGDS | 6 | CHOL,KICH,KIRC,KIRP,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000160271 | RALGDS | 5 | CHOL,KIRC,KIRP,TGCT,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000174125 | TLR1 | 8 | BRCA,GBM,LGG,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000174125 | TLR1 | 8 | BRCA,GBM,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000174125 | TLR1 | 7 | BRCA,LGG,PAAD,SARC,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000133321 | PLAAT4 | 5 | CHOL,KIRP,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000133321 | PLAAT4 | 11 | ACC,CESC,CHOL,COAD,GBM,KIRP,LGG,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000108819 | PPP1R9B | 5 | CHOL,KICH,LUSC,PAAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000163154 | TNFAIP8L2 | 8 | BLCA,BRCA,CHOL,KIRC,KIRP,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000163154 | TNFAIP8L2 | 16 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000163154 | TNFAIP8L2 | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000182179 | UBA7 | 13 | BRCA,CHOL,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000182179 | UBA7 | 11 | BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000182179 | UBA7 | 16 | BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,LAML,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000182179 | UBA7 | 16 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUSC,OV,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000182179 | UBA7 | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000182179 | UBA7 | 12 | CESC,CHOL,ESCA,KIRC,LAML,LUSC,OV,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000099326 | MZF1 | ENSG00000003756 | RBM5 | 5 | CHOL,DLBC,ESCA,KIRC,LUAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000003756 | RBM5 | 7 | CHOL,COAD,ESCA,KIRC,LUAD,LUSC,PAAD | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000003756 | RBM5 | 5 | CHOL,KIRC,LUAD,LUSC,PAAD | | ENSG00000280153 | ENSG00000166432 | ZMAT1 | ENSG00000003756 | RBM5 | 5 | CHOL,DLBC,KIRC,LUAD,PAAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000128604 | IRF5 | 7 | CHOL,GBM,LGG,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000128604 | IRF5 | 8 | BRCA,CHOL,GBM,KIRC,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000128604 | IRF5 | 8 | BRCA,CHOL,GBM,KIRC,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000128604 | IRF5 | 10 | ACC,BRCA,CHOL,KIRC,LGG,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000128604 | IRF5 | 5 | CHOL,KIRC,PAAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000152601 | MBNL1 | 10 | BRCA,CHOL,COAD,GBM,LGG,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000152601 | MBNL1 | 6 | BRCA,CHOL,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000152601 | MBNL1 | 5 | CHOL,LUAD,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000152601 | MBNL1 | 5 | CHOL,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000185291 | IL3RA | 9 | BRCA,CHOL,ESCA,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000185291 | IL3RA | 5 | CHOL,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000170581 | STAT2 | 9 | BRCA,COAD,GBM,KIRC,LGG,PAAD,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000170581 | STAT2 | 8 | BRCA,COAD,KICH,KIRC,KIRP,PAAD,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000170581 | STAT2 | 8 | BRCA,COAD,GBM,KIRC,LGG,PAAD,TGCT,THCA | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000170581 | STAT2 | 6 | COAD,KIRC,PAAD,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000243811 | APOBEC3D | 9 | BRCA,CHOL,COAD,KIRC,PAAD,SARC,SKCM,STAD,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000243811 | APOBEC3D | 8 | BRCA,CHOL,KIRC,KIRP,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000243811 | APOBEC3D | 12 | BRCA,CHOL,COAD,KIRC,KIRP,LAML,LUSC,MESO,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000243811 | APOBEC3D | 11 | BRCA,CHOL,COAD,KIRC,KIRP,OV,PAAD,SARC,SKCM,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000243811 | APOBEC3D | 8 | BRCA,CHOL,KIRC,LAML,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000243811 | APOBEC3D | 11 | BRCA,CHOL,COAD,KIRC,LAML,LUSC,OV,PAAD,SKCM,STAD,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000072818 | ACAP1 | 9 | BLCA,BRCA,CHOL,KIRC,KIRP,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000072818 | ACAP1 | 15 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000072818 | ACAP1 | 13 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000072818 | ACAP1 | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000072818 | ACAP1 | 6 | KIRC,LUAD,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000070190 | DAPP1 | 8 | BRCA,CHOL,COAD,KIRC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000070190 | DAPP1 | 9 | BRCA,CHOL,KIRC,LUAD,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000070190 | DAPP1 | 7 | BRCA,CHOL,KIRC,PAAD,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000070190 | DAPP1 | 7 | CHOL,KIRC,LAML,LUAD,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000070190 | DAPP1 | 5 | BRCA,KIRC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000123329 | ARHGAP9 | 14 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000123329 | ARHGAP9 | 12 | BLCA,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000123329 | ARHGAP9 | 17 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LAML,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000123329 | ARHGAP9 | 17 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUSC,OV,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000123329 | ARHGAP9 | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000123329 | ARHGAP9 | 9 | COAD,KIRC,LUAD,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000135077 | HAVCR2 | 8 | BRCA,COAD,ESCA,LGG,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000135077 | HAVCR2 | 5 | BRCA,ESCA,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000135077 | HAVCR2 | 5 | BRCA,ESCA,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000135077 | HAVCR2 | 10 | ACC,BRCA,COAD,ESCA,LGG,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000135077 | HAVCR2 | 5 | BRCA,ESCA,LAML,SARC,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000163219 | ARHGAP25 | 13 | BRCA,CHOL,COAD,ESCA,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000163219 | ARHGAP25 | 8 | BLCA,BRCA,CHOL,KIRC,KIRP,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000163219 | ARHGAP25 | 15 | BLCA,BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LUAD,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000163219 | ARHGAP25 | 14 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000163219 | ARHGAP25 | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000163219 | ARHGAP25 | 8 | CHOL,COAD,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000140030 | GPR65 | 9 | BRCA,GBM,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000140030 | GPR65 | 8 | BRCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000140030 | GPR65 | 8 | BRCA,KIRC,LUSC,PAAD,SARC,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000140030 | GPR65 | 7 | BRCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000140030 | GPR65 | 5 | KIRC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000130592 | LSP1 | 6 | CHOL,LGG,PAAD,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000130592 | LSP1 | 7 | BRCA,CHOL,KIRC,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000130592 | LSP1 | 10 | BRCA,CHOL,COAD,ESCA,KICH,KIRC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000130592 | LSP1 | 10 | BRCA,CHOL,COAD,ESCA,KIRC,LGG,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000130592 | LSP1 | 6 | CHOL,ESCA,KIRC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000130592 | LSP1 | 5 | CHOL,KIRC,PAAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000105369 | CD79A | 6 | CHOL,ESCA,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000105369 | CD79A | 6 | BLCA,BRCA,CHOL,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000105369 | CD79A | 11 | BLCA,BRCA,CHOL,ESCA,LAML,LUSC,MESO,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000105369 | CD79A | 6 | CHOL,ESCA,LUSC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000105369 | CD79A | 5 | LAML,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000173372 | C1QA | 7 | CHOL,COAD,GBM,KIRC,LGG,SKCM,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000173372 | C1QA | 5 | BRCA,GBM,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000173372 | C1QA | 9 | BRCA,CHOL,COAD,KICH,KIRC,KIRP,MESO,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000173372 | C1QA | 10 | BRCA,CHOL,COAD,KIRC,KIRP,LGG,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000174718 | RESF1 | 7 | CHOL,COAD,KIRC,PAAD,SKCM,STAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000174718 | RESF1 | 6 | CHOL,KIRC,KIRP,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000174718 | RESF1 | 6 | ACC,KIRC,KIRP,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000174718 | RESF1 | 5 | CHOL,KIRC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000196405 | EVL | 12 | BLCA,CHOL,COAD,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000196405 | EVL | 6 | CHOL,COAD,KIRC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000196405 | EVL | 7 | CHOL,KIRC,LUAD,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000159110 | IFNAR2 | 5 | BRCA,ESCA,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000185905 | C16orf54 | 9 | BRCA,CHOL,KIRC,LUSC,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000185905 | C16orf54 | 5 | BRCA,CHOL,KIRC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000185905 | C16orf54 | 9 | BRCA,CHOL,KIRC,LUSC,MESO,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000185905 | C16orf54 | 8 | BRCA,CHOL,KIRC,LUSC,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000185905 | C16orf54 | 7 | BRCA,CHOL,KIRC,LUSC,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000184730 | APOBR | 6 | CHOL,GBM,LGG,LUSC,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000184730 | APOBR | 8 | CHOL,GBM,KIRC,KIRP,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000184730 | APOBR | 11 | CHOL,ESCA,GBM,KICH,KIRC,KIRP,LUAD,LUSC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000184730 | APOBR | 10 | ACC,CHOL,ESCA,KIRC,LGG,LUSC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000185482 | STAC3 | 8 | CHOL,GBM,KIRC,KIRP,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000185482 | STAC3 | 9 | CHOL,COAD,GBM,KIRC,KIRP,LUSC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000185482 | STAC3 | 9 | CHOL,KIRC,LGG,LUSC,OV,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000185482 | STAC3 | 5 | CHOL,KIRC,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000105122 | RASAL3 | 13 | BRCA,CHOL,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000105122 | RASAL3 | 12 | BLCA,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000105122 | RASAL3 | 16 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000105122 | RASAL3 | 16 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,OV,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000105122 | RASAL3 | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000101017 | CD40 | 8 | CHOL,GBM,LGG,LUSC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000101017 | CD40 | 10 | BLCA,BRCA,CHOL,GBM,KIRC,LGG,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000101017 | CD40 | 11 | BRCA,CHOL,COAD,GBM,KIRC,LGG,LUSC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000101017 | CD40 | 5 | CHOL,KIRC,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000101017 | CD40 | 7 | CHOL,COAD,LUSC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000111796 | KLRB1 | 9 | BRCA,CHOL,ESCA,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000111796 | KLRB1 | 12 | BLCA,BRCA,CHOL,COAD,ESCA,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000111796 | KLRB1 | 9 | BRCA,CESC,CHOL,ESCA,KIRP,LUSC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000111796 | KLRB1 | 7 | CHOL,ESCA,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000027869 | SH2D2A | 6 | BRCA,CHOL,KIRC,KIRP,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000027869 | SH2D2A | 8 | BRCA,CHOL,KIRC,KIRP,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000027869 | SH2D2A | 7 | BRCA,CHOL,KIRC,KIRP,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000027869 | SH2D2A | 5 | CHOL,KIRC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000106785 | TRIM14 | 6 | KIRC,LGG,PAAD,SARC,SKCM,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000106785 | TRIM14 | 7 | ACC,KIRC,LGG,PAAD,SARC,SKCM,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000120280 | TASL | 9 | BRCA,GBM,KIRC,LGG,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000120280 | TASL | 9 | BRCA,GBM,KIRC,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000120280 | TASL | 8 | BRCA,KIRC,LGG,LUSC,PAAD,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000120280 | TASL | 6 | BRCA,KIRC,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000185880 | TRIM69 | 9 | BRCA,COAD,KIRC,LGG,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000185880 | TRIM69 | 6 | BRCA,KIRC,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000185880 | TRIM69 | 11 | BLCA,BRCA,COAD,KIRC,LGG,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000185880 | TRIM69 | 5 | BRCA,KIRC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000185880 | TRIM69 | 7 | BRCA,COAD,KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000171608 | PIK3CD | 10 | BRCA,CHOL,COAD,KIRC,LGG,PAAD,SARC,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000171608 | PIK3CD | 8 | BLCA,BRCA,CHOL,KIRC,KIRP,LGG,LUSC,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000171608 | PIK3CD | 13 | BLCA,BRCA,CHOL,COAD,KICH,KIRC,KIRP,LUAD,LUSC,PAAD,SARC,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000171608 | PIK3CD | 12 | BLCA,BRCA,CHOL,COAD,KIRC,KIRP,LGG,LUSC,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000171608 | PIK3CD | 7 | BRCA,CHOL,KIRC,LUAD,LUSC,PAAD,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000171608 | PIK3CD | 9 | BRCA,CHOL,COAD,KIRC,LUSC,PAAD,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000186818 | LILRB4 | 10 | BRCA,CHOL,COAD,GBM,KIRC,LGG,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000186818 | LILRB4 | 5 | BRCA,CHOL,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000186818 | LILRB4 | 9 | BRCA,CHOL,GBM,KICH,KIRC,KIRP,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000186818 | LILRB4 | 10 | BRCA,CHOL,COAD,KIRC,LGG,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000167208 | SNX20 | 10 | BRCA,CHOL,GBM,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000167208 | SNX20 | 6 | BRCA,CHOL,GBM,KIRC,LUSC,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000167208 | SNX20 | 12 | BRCA,CHOL,GBM,KIRC,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000167208 | SNX20 | 9 | BRCA,CHOL,GBM,KIRC,LUSC,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000167208 | SNX20 | 9 | BRCA,CHOL,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000167208 | SNX20 | 8 | BRCA,CHOL,KIRC,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000103187 | COTL1 | 5 | CHOL,LGG,LUSC,PAAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000103187 | COTL1 | 5 | BRCA,CHOL,ESCA,PAAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000103187 | COTL1 | 7 | BRCA,CHOL,ESCA,LUSC,PAAD,SARC,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000103187 | COTL1 | 9 | BRCA,CHOL,ESCA,LGG,LUSC,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000126264 | HCST | 9 | BLCA,BRCA,CHOL,KIRC,KIRP,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000126264 | HCST | 16 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000126264 | HCST | 8 | CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000163162 | RNF149 | 5 | GBM,KIRC,LGG,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000113088 | GZMK | 10 | BRCA,CHOL,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000113088 | GZMK | 7 | BLCA,BRCA,CHOL,KIRC,LUSC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000113088 | GZMK | 11 | BLCA,BRCA,CHOL,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000113088 | GZMK | 11 | BLCA,BRCA,CHOL,KIRC,KIRP,LUSC,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000113088 | GZMK | 8 | BRCA,CHOL,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000113088 | GZMK | 6 | KIRC,LUSC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000108622 | ICAM2 | 5 | BRCA,CHOL,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000188315 | C3orf62 | 5 | CHOL,KIRC,LUAD,LUSC,PAAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000139190 | VAMP1 | 5 | CHOL,COAD,KIRC,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000139190 | VAMP1 | 12 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,LUAD,LUSC,MESO,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000139190 | VAMP1 | 5 | CHOL,ESCA,KIRC,PAAD,SKCM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000139190 | VAMP1 | 5 | CHOL,KIRC,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000139190 | VAMP1 | 6 | CHOL,KIRC,LUAD,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000197857 | ZNF44 | ENSG00000139190 | VAMP1 | 5 | CHOL,ESCA,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000107263 | RAPGEF1 | 5 | BRCA,CHOL,COAD,PAAD,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000110324 | IL10RA | 14 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000110324 | IL10RA | 11 | BLCA,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,LUSC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000110324 | IL10RA | 17 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000110324 | IL10RA | 16 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000110324 | IL10RA | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000110324 | IL10RA | 7 | CHOL,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000266173 | STRADA | 5 | CHOL,KIRC,LUAD,PAAD,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000266173 | STRADA | 5 | CHOL,KIRC,LAML,PAAD,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000082512 | TRAF5 | 5 | CHOL,KIRC,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000082512 | TRAF5 | 7 | BLCA,CHOL,KIRC,KIRP,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000082512 | TRAF5 | 6 | CHOL,KIRC,LUAD,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000082512 | TRAF5 | 6 | CHOL,KIRC,LUAD,PAAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000204397 | CARD16 | 9 | CHOL,ESCA,GBM,LGG,LUSC,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000204397 | CARD16 | 6 | BLCA,CHOL,KIRC,KIRP,PAAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000204397 | CARD16 | 9 | BLCA,CHOL,ESCA,KICH,KIRC,KIRP,LUSC,PAAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000204397 | CARD16 | 15 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000204397 | CARD16 | 6 | CHOL,ESCA,KIRC,LUSC,PAAD,SARC | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000133561 | GIMAP6 | 9 | BRCA,CHOL,COAD,GBM,LGG,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000133561 | GIMAP6 | 10 | BRCA,CHOL,COAD,ESCA,GBM,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000133561 | GIMAP6 | 7 | CHOL,COAD,ESCA,LGG,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000133561 | GIMAP6 | 6 | CHOL,ESCA,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000133561 | GIMAP6 | 5 | CHOL,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000117228 | GBP1 | 7 | BRCA,COAD,GBM,KIRC,LGG,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000117228 | GBP1 | 10 | ACC,BRCA,COAD,GBM,KIRC,LGG,OV,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000117228 | GBP1 | 5 | BRCA,COAD,KIRC,SKCM,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000162654 | GBP4 | 10 | BRCA,COAD,ESCA,KIRC,LGG,LUSC,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000162654 | GBP4 | 8 | BRCA,ESCA,KIRC,KIRP,LUSC,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000162654 | GBP4 | 14 | ACC,BLCA,BRCA,COAD,ESCA,KIRC,KIRP,LGG,LUSC,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000162654 | GBP4 | 7 | BRCA,ESCA,KIRC,LAML,LUSC,SARC,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000162654 | GBP4 | 7 | COAD,KIRC,LAML,LUSC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000099326 | MZF1 | ENSG00000167280 | ENGASE | 5 | CHOL,DLBC,KICH,KIRC,LUAD | | ENSG00000280153 | ENSG00000166432 | ZMAT1 | ENSG00000167280 | ENGASE | 5 | CHOL,DLBC,KIRC,LUAD,PAAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000129566 | TEP1 | 7 | COAD,ESCA,KIRC,LGG,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000129566 | TEP1 | 8 | COAD,ESCA,GBM,KIRC,LAML,LUAD,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000129566 | TEP1 | 6 | ESCA,KIRC,LGG,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000129566 | TEP1 | 5 | KIRC,LAML,LUAD,SARC,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000129566 | TEP1 | 5 | KIRC,LAML,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000139436 | GIT2 | 6 | CHOL,COAD,ESCA,LUSC,PAAD,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000139436 | GIT2 | 7 | CHOL,ESCA,LAML,LUAD,LUSC,PAAD,STAD | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000139436 | GIT2 | 7 | CHOL,ESCA,LAML,LUAD,LUSC,PAAD,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000139436 | GIT2 | 5 | CHOL,COAD,LUSC,PAAD,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000159958 | TNFRSF13C | 8 | BLCA,CHOL,LUAD,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000159958 | TNFRSF13C | 6 | CHOL,LUAD,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000138496 | PARP9 | 6 | COAD,KIRC,LGG,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000138496 | PARP9 | 7 | COAD,KIRC,LGG,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000138496 | PARP9 | 5 | COAD,KIRC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000133805 | AMPD3 | 6 | BRCA,CHOL,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000136167 | LCP1 | 14 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000136167 | LCP1 | 9 | BRCA,CHOL,ESCA,GBM,KIRP,LGG,LUSC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000136167 | LCP1 | 15 | BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000136167 | LCP1 | 13 | BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000136167 | LCP1 | 8 | CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000136167 | LCP1 | 9 | CHOL,COAD,KIRC,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000204252 | HLA-DOA | 13 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000204252 | HLA-DOA | 7 | BRCA,CHOL,ESCA,GBM,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000204252 | HLA-DOA | 14 | BLCA,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,LAML,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000204252 | HLA-DOA | 18 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,OV,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000204252 | HLA-DOA | 8 | CHOL,ESCA,KIRC,LAML,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000204252 | HLA-DOA | 14 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,LAML,OV,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000141506 | PIK3R5 | 11 | BRCA,CHOL,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000141506 | PIK3R5 | 6 | BRCA,CHOL,ESCA,GBM,LGG,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000141506 | PIK3R5 | 13 | BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000141506 | PIK3R5 | 8 | BRCA,CHOL,ESCA,KIRC,LGG,LUSC,SARC,SKCM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000141506 | PIK3R5 | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000141506 | PIK3R5 | 5 | CHOL,KIRC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000105643 | ARRDC2 | 10 | BRCA,ESCA,KICH,KIRP,LAML,LUSC,MESO,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000047365 | ARAP2 | 6 | BRCA,COAD,KIRC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000047365 | ARAP2 | 6 | BRCA,KIRC,LAML,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000164691 | TAGAP | 9 | BRCA,CHOL,COAD,ESCA,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000164691 | TAGAP | 5 | BLCA,BRCA,CHOL,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000164691 | TAGAP | 9 | BLCA,BRCA,CHOL,COAD,ESCA,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000164691 | TAGAP | 9 | BLCA,BRCA,CHOL,COAD,ESCA,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000164691 | TAGAP | 6 | BRCA,CHOL,ESCA,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000164691 | TAGAP | 7 | BRCA,CHOL,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000117091 | CD48 | 12 | BRCA,CHOL,COAD,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000117091 | CD48 | 7 | BLCA,BRCA,CHOL,KIRC,KIRP,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000117091 | CD48 | 15 | BLCA,BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000117091 | CD48 | 12 | BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000117091 | CD48 | 5 | KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000141968 | VAV1 | 11 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,LUSC,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000141968 | VAV1 | 16 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000141968 | VAV1 | 12 | BRCA,CHOL,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000141968 | VAV1 | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000141968 | VAV1 | 9 | CHOL,KIRC,LUAD,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000012124 | CD22 | 9 | BLCA,BRCA,CHOL,LUAD,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000012124 | CD22 | 5 | CHOL,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000163660 | CCNL1 | 5 | CHOL,KIRC,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000241106 | HLA-DOB | 11 | BRCA,CHOL,COAD,ESCA,KIRC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000241106 | HLA-DOB | 12 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000241106 | HLA-DOB | 14 | BRCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,OV,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000241106 | HLA-DOB | 7 | CHOL,ESCA,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000241106 | HLA-DOB | 13 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,OV,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000134255 | CEPT1 | 5 | CHOL,LGG,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000134255 | CEPT1 | 6 | CHOL,ESCA,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000134255 | CEPT1 | 6 | CHOL,LUAD,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000197857 | ZNF44 | ENSG00000134255 | CEPT1 | 5 | CHOL,ESCA,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000169413 | RNASE6 | 12 | BRCA,CHOL,ESCA,GBM,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000169413 | RNASE6 | 11 | BRCA,CHOL,ESCA,KICH,KIRP,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000169413 | RNASE6 | 11 | ACC,BRCA,CHOL,ESCA,LGG,LUSC,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000169413 | RNASE6 | 8 | BRCA,CHOL,ESCA,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000169413 | RNASE6 | 5 | CHOL,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000172543 | CTSW | 8 | BRCA,CHOL,COAD,ESCA,PAAD,SARC,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000172543 | CTSW | 6 | BRCA,CHOL,KIRC,KIRP,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000172543 | CTSW | 11 | BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000172543 | CTSW | 12 | BRCA,CHOL,COAD,ESCA,KIRC,KIRP,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000197540 | GZMM | 5 | CHOL,LUSC,PAAD,STAD,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000197540 | GZMM | 7 | BRCA,CHOL,KIRC,KIRP,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000197540 | GZMM | 12 | BRCA,CHOL,COAD,KIRC,KIRP,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000197540 | GZMM | 11 | BRCA,CHOL,COAD,KIRC,KIRP,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000197540 | GZMM | 7 | CHOL,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000003402 | CFLAR | 11 | BRCA,COAD,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000003402 | CFLAR | 7 | ESCA,GBM,KIRC,LGG,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000003402 | CFLAR | 12 | BRCA,COAD,ESCA,GBM,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000003402 | CFLAR | 11 | ESCA,GBM,KIRC,LGG,LUSC,OV,PAAD,SARC,SKCM,TGCT,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000003402 | CFLAR | 8 | ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000003402 | CFLAR | 6 | KIRC,OV,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000005844 | ITGAL | 14 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000005844 | ITGAL | 9 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000005844 | ITGAL | 16 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000005844 | ITGAL | 14 | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000005844 | ITGAL | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000005844 | ITGAL | 8 | BRCA,COAD,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000043462 | LCP2 | 13 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000043462 | LCP2 | 10 | BLCA,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000043462 | LCP2 | 13 | BLCA,BRCA,CHOL,ESCA,GBM,KICH,KIRC,KIRP,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000043462 | LCP2 | 14 | ACC,BLCA,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000043462 | LCP2 | 8 | BRCA,CHOL,ESCA,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000043462 | LCP2 | 8 | BRCA,CHOL,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000132274 | TRIM22 | 12 | BRCA,CHOL,COAD,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000132274 | TRIM22 | 9 | BLCA,BRCA,CHOL,KIRC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000132274 | TRIM22 | 14 | ACC,BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000132274 | TRIM22 | 8 | BRCA,CHOL,KIRC,LAML,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000132274 | TRIM22 | 9 | CHOL,COAD,KIRC,LAML,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000169442 | CD52 | 7 | BLCA,CHOL,KIRC,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000169442 | CD52 | 13 | BLCA,CHOL,COAD,ESCA,KICH,KIRC,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000169442 | CD52 | 8 | CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000145649 | GZMA | 10 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000145649 | GZMA | 7 | BRCA,CHOL,GBM,KIRC,KIRP,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000145649 | GZMA | 9 | BRCA,CHOL,ESCA,KIRC,KIRP,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000145649 | GZMA | 13 | ACC,BRCA,CHOL,COAD,ESCA,KIRC,KIRP,PAAD,SARC,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000145649 | GZMA | 7 | BRCA,CHOL,ESCA,KIRC,PAAD,SARC,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000145649 | GZMA | 5 | KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000244509 | APOBEC3C | 9 | BRCA,CHOL,GBM,KIRC,LGG,PAAD,SARC,STAD,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000244509 | APOBEC3C | 6 | BRCA,CHOL,GBM,KIRC,PAAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000244509 | APOBEC3C | 8 | BRCA,CHOL,KICH,KIRC,MESO,PAAD,SARC,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000244509 | APOBEC3C | 7 | BRCA,GBM,KIRC,LGG,PAAD,SARC,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000244509 | APOBEC3C | 5 | CHOL,KIRC,PAAD,SARC,STAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000188827 | SLX4 | 5 | CHOL,KICH,KIRC,PAAD,STAD | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000108639 | SYNGR2 | 5 | LGG,SARC,SKCM,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000105483 | CARD8 | 11 | BRCA,CHOL,COAD,ESCA,KIRC,LGG,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000099326 | MZF1 | ENSG00000105483 | CARD8 | 5 | CHOL,ESCA,KICH,KIRC,LUAD | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000105483 | CARD8 | 13 | BLCA,BRCA,CHOL,ESCA,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000105483 | CARD8 | 7 | CHOL,ESCA,KIRC,KIRP,LUSC,PAAD,TGCT | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000105483 | CARD8 | 9 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000105483 | CARD8 | 5 | CHOL,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000197857 | ZNF44 | ENSG00000105483 | CARD8 | 5 | CHOL,ESCA,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000004534 | RBM6 | 6 | CHOL,ESCA,KIRC,LUAD,LUSC,PAAD | | ENSG00000280153 | ENSG00000197857 | ZNF44 | ENSG00000004534 | RBM6 | 5 | CHOL,ESCA,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000011600 | TYROBP | 5 | CHOL,GBM,LGG,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000011600 | TYROBP | 5 | CHOL,KIRC,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000011600 | TYROBP | 6 | CHOL,KICH,KIRC,KIRP,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000011600 | TYROBP | 8 | ACC,CHOL,KIRC,LGG,SKCM,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000137841 | PLCB2 | 13 | BRCA,CHOL,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000137841 | PLCB2 | 10 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,PAAD,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000137841 | PLCB2 | 17 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000137841 | PLCB2 | 15 | ACC,BRCA,CESC,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000137841 | PLCB2 | 9 | BRCA,CHOL,ESCA,KIRC,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000137841 | PLCB2 | 9 | CESC,CHOL,COAD,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000186517 | ARHGAP30 | 14 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUSC,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000186517 | ARHGAP30 | 9 | BLCA,BRCA,CHOL,GBM,KIRC,KIRP,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000186517 | ARHGAP30 | 17 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KICH,KIRC,KIRP,LUAD,LUSC,MESO,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000186517 | ARHGAP30 | 14 | BRCA,CHOL,COAD,ESCA,KIRC,KIRP,LGG,LUSC,OV,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000186517 | ARHGAP30 | 10 | BRCA,CHOL,ESCA,KIRC,LUAD,LUSC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000186517 | ARHGAP30 | 9 | BRCA,CHOL,COAD,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000156171 | DRAM2 | 6 | CHOL,LGG,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000156171 | DRAM2 | 5 | CHOL,PAAD,SARC,SKCM,TGCT | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000102871 | TRADD | 5 | CHOL,GBM,LGG,SKCM,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000187764 | SEMA4D | 6 | BRCA,CHOL,PAAD,SARC,SKCM,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000187764 | SEMA4D | 10 | BLCA,BRCA,CHOL,COAD,KIRC,PAAD,SARC,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000187764 | SEMA4D | 8 | BRCA,CHOL,KIRC,PAAD,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000187764 | SEMA4D | 7 | BRCA,CHOL,KIRC,PAAD,SARC,SKCM,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000187764 | SEMA4D | 8 | BRCA,CHOL,KIRC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000124508 | BTN2A2 | 13 | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000124508 | BTN2A2 | 8 | BLCA,BRCA,CHOL,ESCA,KIRC,LGG,PAAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000124508 | BTN2A2 | 12 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,PAAD,SARC,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000124508 | BTN2A2 | 14 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,SARC,TGCT,THCA,THYM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000124508 | BTN2A2 | 8 | BRCA,CHOL,ESCA,KIRC,LAML,PAAD,SARC,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000124508 | BTN2A2 | 14 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,KIRC,LAML,OV,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000134262 | AP4B1 | 6 | CHOL,ESCA,KIRC,KIRP,LUSC,SKCM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000134262 | AP4B1 | 5 | CHOL,KIRC,LUSC,PAAD,SKCM | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000134262 | AP4B1 | 6 | CHOL,KIRC,LUAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000197857 | ZNF44 | ENSG00000134262 | AP4B1 | 5 | CHOL,ESCA,LUAD,PAAD,SKCM | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000198625 | MDM4 | 5 | KIRC,LAML,LUAD,PAAD,STAD | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000198625 | MDM4 | 5 | LAML,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000151702 | FLI1 | 11 | BRCA,CHOL,COAD,ESCA,LGG,LUSC,PAAD,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000151702 | FLI1 | 12 | BLCA,BRCA,CHOL,COAD,ESCA,KICH,KIRP,LUSC,PAAD,SKCM,STAD,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000151702 | FLI1 | 7 | CHOL,ESCA,KIRP,LUSC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000135899 | SP110 | ENSG00000151702 | FLI1 | 6 | CHOL,ESCA,LUSC,PAAD,SKCM,STAD | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000196954 | CASP4 | 7 | BRCA,GBM,KIRC,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000077150 | NFKB2 | ENSG00000196954 | CASP4 | 5 | GBM,KIRC,KIRP,PAAD,THCA | | ENSG00000280153 | ENSG00000106948 | AKNA | ENSG00000196954 | CASP4 | 5 | BRCA,KIRC,KIRP,SKCM,TGCT | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000196954 | CASP4 | 9 | ACC,BRCA,GBM,KIRC,KIRP,PAAD,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000067066 | SP100 | ENSG00000166750 | SLFN5 | 7 | COAD,LGG,SARC,SKCM,STAD,TGCT,THCA | | ENSG00000280153 | ENSG00000125347 | IRF1 | ENSG00000166750 | SLFN5 | 5 | LGG,SARC,SKCM,TGCT,THCA | | ENSG00000280153 | ENSG00000143390 | RFX5 | ENSG00000166750 | SLFN5 | 5 | COAD,SKCM,STAD,TGCT,THCA |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation |
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000280153.1 | ENST00000625054.1 | 104 | 177 | 488 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.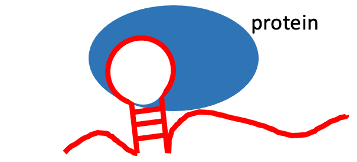
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).