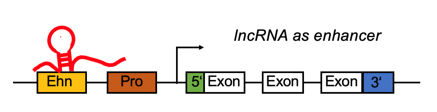
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | TRIR,NDUFB10,RPS17,MRPL52,BLOC1S1,C11orf1,DUT,RTRAF,UXT,ATP5F1E,ZBTB8OS,CFDP1,RPS29,MRPL55,EMG1,C12orf57,NDUFB8,PSMA2,SEM1,PET100,IMP3,SERGEF,NDUFA13,ARF5,ATP5MG,CDPF1,MRPL47,GADD45GIP1,SRP19,SNRPD2,TMUB1,EIF3F,HINT2,RPS27A,RSL24D1,RPL23,XAB2,C19orf53,BUD23,TTC1,BCL7C,MRPL28,NDUFA11,NPRL2,HSCB,RPL27,METTL5,HPF1,MIX23,RPL13A,DTYMK,RPL37A,TRAPPC2L,LSM3,ZNHIT1,UQCRB,PHKG2,BUD31,TIMM9,POLR2J,SS18L2,SWSAP1,LSM5,RFXANK,NDUFA3,COPS9,SVBP,STYXL1,STX10,ECI1,NAA38,SNU13,LRRC23,GTF2H5,TPT1,RPL36AL,CWC15,LYRM4,MPLKIP,ATP5MJ,DPY30,UBL5,RPS8,MAP2K2,RPS11,SF3B6,NELFE,METTL23,ALKBH7,TMEM258,MRPL22,STX8,BTF3,GEMIN6,NDUFB7,MVB12A,UBXN1,MRPS33,THYN1,PGLS,ATP5MK,SMDT1,YJU2,SEC61G,SUMO1,RPL35,NDUFA2,TAF10,COX4I1,C18orf21,SGF29,TIGD1,RPL36A,COA8,DYRK4,CENPS,RPS6,SERF2,RPS10,PSMB3,MRPS9,COMMD6,MRPL57,RPS24,CIAO2B,PYURF,TMA7,C2orf76,PFDN5,CCS,MRPL27,PSMG3,PSMG2,HMGN3,TPRKB,PSMB1,NDUFB1,DUSP28,SREK1IP1,POP5,NACA,CZIB,SYF2,RPS19BP1,HDDC2,COA5,RPA3,MICOS10,NSMCE1,COA6,SDHAF2,SCAND1,RPL23A,ATP5MC2,SSBP1,ATP5F1D,RPS13,CCDC12,PTRHD1,TOMM7,REX1BD,THOC7,COX7A2,ERCC5,SIRT6,RP9,NOP53,SMIM19,MRPL54,SRA1,ATP5IF1 | | Exist in public source | | NA |
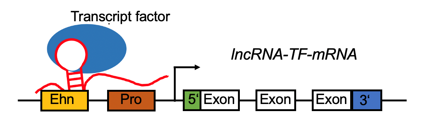
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | ECI1,LAMTOR4,PSMD4,ATPIF1,RPL35,RPS23,RP9,HINT2,C6orf203,EIF3F,GTF2H5,RPSA,DYRK4,MAP2K2,PSMB3,NDUFA3,C14orf2,TIMM9,HSCB,POP5,C4orf27,PSMA2,GADD45GIP1,SGF29,ARIH2OS,PGLS,USMG5,NDUFB1,SHFM1,NDUFS5,MRPL52,UBXN1,SRP14 | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
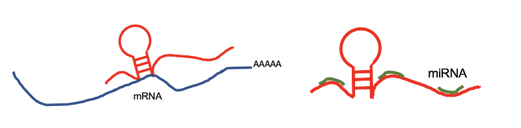
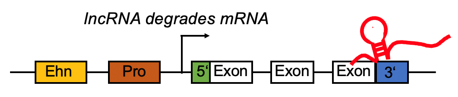
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | TFIP11,ATP2A2,KMT2D,KIF2A,BAZ2A,VPS26C,EIF4G2,FLNA,CSNK1A1,AP3D1,MYH9,PLEKHM1,RNF123,ACTN4,SNX6,USP32,CARM1,LRP10,PI4KA,SSH1,RNF31,SMG6,ITGB1,EIF4G1,AMBRA1,SEC23A,KLHL5,MAN2B2,CLTC | | Exist in public source | | NA |
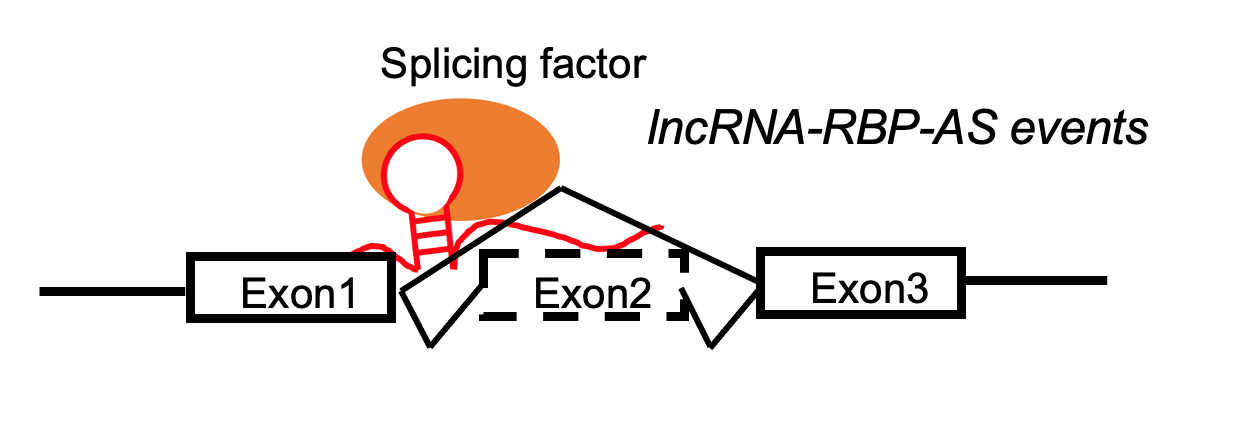
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation |
| -lncRNA and exonskipping events are negatively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000279865 | ENSG00000099385 | BCL7C | DLBC,GBM,KIRP,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000100209 | HSCB | DLBC,GBM,KIRP,LAML,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000106028 | SSBP1 | DLBC,GBM,KIRP,LAML,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000106245 | BUD31 | DLBC,GBM,KIRP,LAML,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000106400 | ZNHIT1 | DLBC,GBM,KIRP,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000110700 | RPS13 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000123349 | PFDN5 | DLBC,GBM,KIRP,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000125691 | RPL23 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000126756 | UXT | DLBC,GBM,KIRP,LAML,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000131469 | RPL27 | DLBC,GBM,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000133112 | TPT1 | DLBC,GBM,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000136942 | RPL35 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000137154 | RPS6 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000140264 | SERF2 | DLBC,GBM,KIRP,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000142534 | RPS11 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000142541 | RPL13A | DLBC,GBM,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000142937 | RPS8 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000145337 | PYURF | DLBC,GBM,KIRP,LAML,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000167272 | POP5 | DLBC,GBM,KIRP,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000188243 | COMMD6 | DLBC,GBM,KIRP,LAML,LGG,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000198242 | RPL23A | DLBC,GBM,LAML,LGG,THYM | GBM | | | ENSG00000279865 | ENSG00000198258 | UBL5 | DLBC,GBM,KIRP,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000213523 | SRA1 | DLBC,GBM,KIRP,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000213741 | RPS29 | DLBC,GBM,LAML,LGG,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000241343 | RPL36A | DLBC,GBM,KIRP,LAML,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000106399 | RPA3 | GBM,KIRP,LAML,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000125743 | SNRPD2 | GBM,KIRP,LAML,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000127922 | SEM1 | GBM,KIRP,LAML,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000134825 | TMEM258 | GBM,KIRP,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000143947 | RPS27A | GBM,LAML,LGG,THCA,THYM,UCS | GBM | | | ENSG00000279865 | ENSG00000144034 | TPRKB | GBM,KIRP,LAML,THCA,THYM | GBM | | | ENSG00000279865 | ENSG00000160124 | MIX23 | GBM,KIRP,LAML,THCA,THYM,UCS | GBM | |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000279865 | ENST00000624929 | AMBRA1 | ENSG00000110497 | ENST00000526545 | -0.6406 | 129 | - | GBM | | ENSG00000279865 | ENST00000624929 | RNF123 | ENSG00000164068 | ENST00000432042 | -0.1143 | 342 | - | GBM | | ENSG00000279865 | ENST00000624929 | RNF123 | ENSG00000164068 | ENST00000443204 | -0.1034 | 167 | - | GBM | | ENSG00000279865 | ENST00000624929 | RNF123 | ENSG00000164068 | ENST00000444689 | -0.1055 | 119 | - | GBM | | ENSG00000279865 | ENST00000624929 | KMT2D | ENSG00000167548 | ENST00000526209 | -0.1305 | 113 | - | GBM | | ENSG00000279865 | ENST00000624929 | USP32 | ENSG00000170832 | ENST00000592339 | -0.2543 | 1 | - | GBM | | ENSG00000279865 | ENST00000624929 | USP32 | ENSG00000170832 | ENST00000593071 | -0.1317 | 608 | - | GBM | | ENSG00000279865 | ENST00000624929 | USP32 | ENSG00000170832 | ENST00000590297 | -0.1422 | 671 | - | GBM | | ENSG00000279865 | ENST00000624929 | ATP2A2 | ENSG00000174437 | ENST00000553144 | -0.1131 | 1203 | - | GBM | | ENSG00000279865 | ENST00000624929 | PI4KA | ENSG00000241973 | ENST00000255882 | -0.1074 | 1 | - | GBM | | ENSG00000279865 | ENST00000624929 | PI4KA | ENSG00000241973 | ENST00000399213 | -0.1074 | 1 | - | GBM |
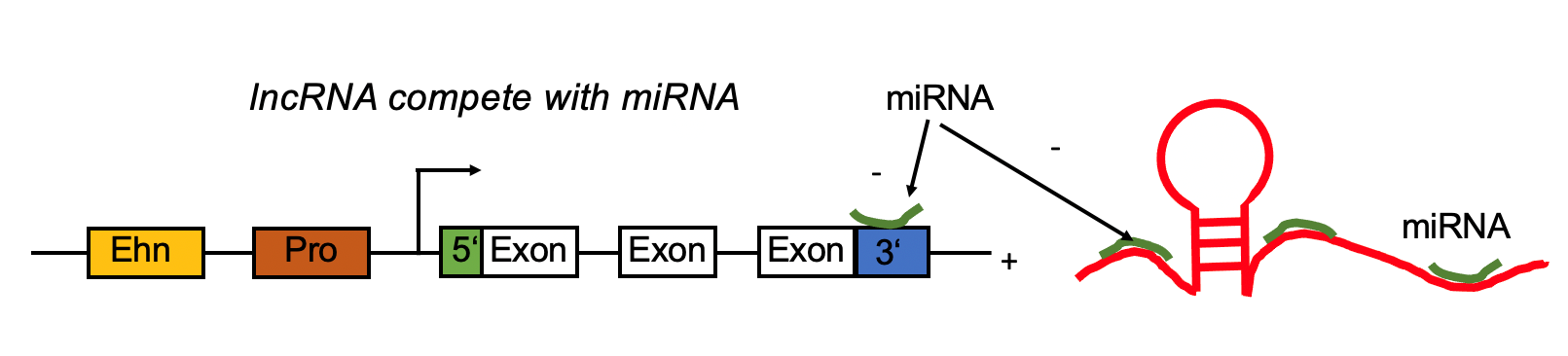
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000279865.1 | ENST00000624929.1 | 65 | 165 | 362 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.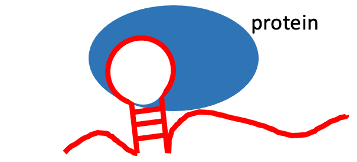
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).