lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | EEF1AKMT3,ARHGEF17,FERMT2,VAMP1,MTMR6,BICD2,MFSD8,DNAJC27,WDFY1,LIFR,MID1,KLF2,VAMP4,SLF2,GNAQ,RAB11FIP5,SLC38A2,PDK4,RNF6,HMGCS1,EDIL3,PEA15,MSANTD2,TERF2IP,IBTK,TTC14,AFAP1,C20orf194,ZFP1,PPIL4,CA11,PSIP1,TMEM242,RIC8B,UBE3A,TMEM30A,ITGA7,ZBTB6,C3orf18,ERG,SERTAD4,SGCA,ATP9A,UBE2H,PURA,EPDR1,CD164,CTDSP2,CBX7,HEG1,KLF10,TSPAN2,PTPRB,COPZ2,DIPK1A,RAB5A,SEC23A,KCTD2,SECISBP2L,SSBP2,PHC3,TES,AIDA,ZZEF1,FYN,INTS2,SUPT20H,DCUN1D2,CBX6,SLU7,NCALD,DZIP1,CDK17,ATG2B,AP4E1,DDX42,WDR35,FAM204A,PGGT1B,MAF,CYFIP2,MECOM,TMEM64,PLPP1,ARHGEF9,SLC26A11,USP27X,FANCL,TTC17,KLF7,WBP4,PAWR,MIB1,CEP68,MLLT6,ZNF70,ICE2,BIVM,CRLF3,MAPK9,DNAJB4,ZNF25,NXN,PPP2CB,EPN2,PIP5K1C,KIAA1143,TMED8,IFT74,OSBPL8,NPR2,MZT1,CBLB,XPO4,S1PR1,UFL1,PKD1,SLC25A46,UTP14C,BBS2,TBC1D9,APOD,PTPRN2,SEC23IP,LMOD1,CHRNB1,REEP5,MCAM,TMOD2,KCTD18,ACACB,RNF20,ERO1B,ACADSB,SMIM14,ATG14,GOLGB1,DNAJC18,UCHL5,SERP2,SLC39A10,HDAC2,CACNA1H,ZFR,SCAF11,TMEM167A,DAB2,KCNMA1,PDS5B,ALDH2,SFRP1,CRMP1,OSBPL5,DCTN4,LDB2,KIF3A,SCAMP1,GSTA4,MAN1A1,BMERB1,CASQ2,MTRF1L,EPB41L2,ABLIM1,FAM107A,PIK3CA,KCNJ8,SAR1B,PREPL,MTMR10,PBX2,KLF12,ANGEL2,MYSM1,IGFBP7,ITPRID2,ELF1,C12orf29,TNS1,TMTC1,KLHL13,IFT88,CYP1B1,COX11,CRY1,MEF2C,AFTPH,DHFR2,DMPK,TOM1L2,RPS6KB1,GABBR1,SH3BGRL,KANSL3,ITGA1,RGL1,SH2B3,INIP,RAB40B,TGFBR3,ARL13B,RBM26,SVIL,CAMK2G,PURB,CEBPZOS,SYNE1,CADM1,CLSTN3,KANSL1L,ANKRD46,WDFY3,HEATR5B,GHITM,LCOR,PHF1,TMEM59L,FBXL20,CENPC,HERC2,ABHD13,RCAN2,AOC3,C2orf49,PGAP4,POLK,PLEKHA3,PRKAA1,PRDM2,R3HDM2,LCA5,VEZT,USP12,AAK1,MOSMO,BAG4,SASH1,BMPR2,CERKL,NAP1L3,CSPG4,PLSCR4,FAM13B,DTX3,SPOPL,RAPGEF5,UBE2V1,CDK6,TMEM47,REEP3,BROX,CFL2,APPL1,MYLK,DSTYK,SLC25A27,EXOC5,CSDE1,GXYLT1,ADGRF5,MYL9,MAP3K2,RPRD1A,TOR1AIP1,NHLRC2,TTC28,B4GAT1,ANKRD42,KMT2E,CWC22,DCAF16,MORC3,DLG4,TRIM3,KAT6B,PDSS2,TRIM23,TECPR2,DTWD1,SLC25A30,IKZF4,PIP4K2B,NF1,ANKRD50,CDC42EP3,THAP6,LSAMP,SEC62,GGPS1,SENP7,PTPN21,TRPM7,TRAPPC8,PHIP,DOCK9,LRRC58,PPP1R21,CRIM1,RBM18,JAK1,SNX1,TM2D1,RASSF8,ATAD1,HP1BP3,DNM1,VPS13C,TAPT1,ITPR1,DIDO1,LTBP1,ALG9,GAS6,EMCN,CLIP3,PUM2,FOXC1,TUBE1,YAP1,RCBTB2,WASL,TPM2,ITFG1,MAPK1,FRMD4B,TSPYL4,PUM1,USP22,RB1CC1,IL6ST,ZRANB1,DYNC2LI1,PRX,PDE5A,CITED2,DMTF1,NSD3,MAP4,VPS39,RFTN1,GNAZ,AFF4,LZTFL1,RNF38,PDE2A,HBS1L,TSPAN3,IK,SIN3A,NIBAN1,UFSP2,MEX3C,CREB1,CLEC14A,TACR2,DPY19L4,CAPN7,ADCY5,ADAM10,SMAD2,CLCN6,IGIP,RFX2,RNF19A,FIG4,ASH1L,NFIA,PTAR1,NEBL,KATNAL1,FLI1,TRIM37,GABPB1,VPS36,FBXO38,SYPL1,HCFC2,PRKACB,DUSP3,RSRC2,AK3,NAPB,PNMA8A,ARL15,GTDC1,C9orf72,CHD9,AGO3,SBNO1,C1orf52,PNMA1,WASF1,MOAP1,STAG2,SLC12A4,HDDC2,N4BP2L1,SDE2,ANKRD13C,CCNT1,TTC3,PRDM8,INSR,STXBP5,ATF7IP,TMEM248,SHOC2,MEF2D,RSPH3,HERC4,CASTOR3,RAB28,DENND5A,SHISAL1,DPY19L3,HOOK3,ENPP4,DIP2C,MORF4L1,NEXN,RTL6,SAMD8,FAM171B,SMAP1,SPART,ATL1,PGRMC2,LARP6,METTL21A,OTUD7B,ACAP2,SWAP70,WSB2,SSC5D,TNKS2,SESN3,LANCL1,PPP3CB,OXR1,POPDC2,PI16,AMIGO1,LYSMD3,FNBP1,NEK9,HMGB1,DDX6,SESTD1,EVC,MECP2,AASS,TNRC18,PPP1R3F,TTC8,USP6NL,ST3GAL3,MBLAC2,IGFBP5,GTF2A1,BOD1L1,FAXDC2,RAB33B,LTBP4,CHD2,LONRF2,MINDY3,DKK3,METTL14,GNAO1,OTUD4,AHNAK,RABL3,HERC1,QKI,RASSF3,TMEM35A,SLC41A3,HERC3,PITPNA,PIGK,SBF2,LPP,ANP32E,SYNGAP1,LRP11,FBXO30,HECTD4,ARHGEF25,USP45,GLOD4,GOPC,EVI5,BCL6,LATS2,TMX3,ZBTB4,UBE2B,MDM1,HES1,RAB4A,MED13L,FCHO2,SBSPON,OLFML1,OSTM1,NDRG4,ITSN1,PPP1R12A,KLHDC1,HEYL,RNPC3,C14orf28,TRPS1,C1orf21,MNT,PCDH18,NTN4,BTBD19,DOP1A,KIAA2026,KMT2C,CCSAP,MARCHF8,CCNT2,SNX14,ORC3,PBXIP1,CLMP,ST6GALNAC6,ZFP91,ARMCX3,KLHL9,RNF150,OPTN,SERINC3,ERMARD,CAVIN2,EPC1,ROBO4,ZC3H13,PANK3,SAP30L,NRP1,HEATR6,MAP3K7,LIG4,GASK1B,PNRC1,FCHSD2,ARHGEF26,KRR1,WBP1L,VAMP3,FAM149B1,NAB2,FBXO32,RASL12,GAS1,UBL3,ABI3BP,TMTC3,RFX3,TMEM245,SNCA,HSPB6,CYLD,REEP1,SRGAP2,HELZ,SEL1L,ELF2,ZNF43,PTPRK,PDE3A,STXBP3,GLT8D2,SAR1A,TNS2,TMX4,TPP2,HDAC4,LTBP3,RAI2,SOS1,RTN4,JMY,ZDHHC21,WDR37,CEP112,CPEB2,MSL2,SLFN5,KLHL2,MAMDC2,GNA13,RAPGEF4,GLI3,RABGAP1L,LHFPL6,MAML2,TUB,SF3B1,ANKIB1,MXRA7,TFE3,SETD7,LDLRAD4,CCN2,ARL3,PEX12,MINDY2,USP24,SOS2,POGZ,AKAP10,RBM4B,MAP3K12,FUBP1,PSD,KLF11,NID1,KIDINS220,DDX17,FMR1,BBS12,CTDSPL2,GNPDA2,EXOC6B,EID1,GPALPP1,SORBS1,NSMF,ANO6,MTFR1L,CRYZL1,LRP12,TM9SF3,ZFX,RMND5A,RABGAP1,PER3,SEPTIN7,SH3BP5,CEP162,DST,RBMS1,ZNF17,CRBN,GNG7,ANTXR2,ZNF84,BAZ2B,CCP110,BICRAL,PCMTD1,JMJD1C,MAPK3,VPS37A,ASB8,FUT11,KBTBD2,IFIT5,ATF2,MRFAP1L1,C6orf89,REPS1,KCNMB1,CSNK1G3,SH3D19,ATP6V1A,SPATA13,TRIQK,CHM,MMRN2,FOXF2,PHF10,RAB6A,ISCU,FBXL17,GDAP1,ANKH,FLT4,ZFP14,SMARCC2,KCTD10,MYO9A,TVP23B,PBX3,SGCB,KLHL42,SMIM15,EXOC8,SARAF,STRN3,HSPA13,CCDC146,GZF1,BLZF1,CNST,CLOCK,SLC22A17,BBIP1,BACE1,CLK1,RFX7,SLC17A5,MAP2K4,MAP7,MBD5,PLK2,RNF11,CCDC28A,CDC27,WWTR1,CHIC1,YTHDC1,MFAP5,FAM229B,MTMR2,CCDC191,TNKS,NR2C2,FBLN5,PPP4R2,SCRN3,CYRIA,PLS1,ARHGAP29,GLS,MBTD1,LNP1,MAN1A2,MANEA,ADGRL4,VPS13A,NACC2,EPB41L3,DACH1,ERCC5,ZFP3,SH3PXD2A,KBTBD6,RARS2,FAM172A,SNX33,PER2,SMURF2,FBXL3,PRR3,PRICKLE2,FAM168B,PPWD1,CLASP1,DNAJB5,NHLRC3,KRIT1,IPMK,DCAF7,LAYN,APBB1,SERINC1,UGGT2,ZMYM5,NOTCH4,PCID2,SETX,TRIM13,FGFR1OP2,OSBPL1A,UBE2E2,SACS,PABIR1,MTRR,PUS7L,MEIS1,CTBS,CD47,GGNBP2,ERCC6L2,RTF1,FBXO3,NEMF,REV3L,WDR47,AFDN,SPRYD3,CD200,DZIP3,FNBP4,PPM1K,TMF1,MLLT10,BHLHB9,DENND5B,MED23,PDLIM3,ACTA2,ZNF720,NCKAP1,MIER3,BTBD7,GPBP1,SCARF1,TTBK2,CASKIN2,CDK19,ZFHX3,ARID4A,RNF115,SMAD4,AASDHPPT,VCPIP1,CCDC80,NELL2,PPP3CA,CLIP4,PLPP3,DSTN,MPDZ,PER1,SFXN3,VEGFC,BBS7,MFAP4,PNISR,SEPTIN8,TMEM59,SMARCA1,CTBP2,PCYOX1,UCHL1,FAM126B,CEP120,CDC42BPA,OSBPL9,ROCK1,IQCE,FAM214A,NLGN2,STAM2,SPARCL1,CCDC6,PBRM1,TCAF1,ANKRD10,PPP4R3B,OBI1,NECAP1,ZER1,PRKACA,CCDC50,KLF6,SSH1,FYCO1,EEA1,UBE2W,CHD6,RBPMS,BMPR1A,CLDND1,FBXW7,AFAP1L2,SPRTN,POLI,SH3BGR,ITSN2,DLC1,PJA2,IDS,ADGRA2,UBR3,COL14A1,CEP44,SLIT3,PROS1,GMCL1,C10orf88,KPNA3,KRAS,ANKRA2,SOGA1,CELF1,CDC40,CCDC121,PEX3,ZCCHC24,FAM161B,UHRF1BP1L,CDS2,GIN1,SLC2A13,ARHGAP31,C1S,POU6F1,CDK14,NCAM1,YPEL5,DYNC1LI2,RUFY2,ERMAP,PDCL,SCARB2,PELI2,RYBP,TPPP,GFOD1,SPOCK1,RBM39,GPATCH8,VSTM4,TEK,C7orf31,FAM120B,FBN1,BLOC1S6,CD99L2,INTS3,PHLDB1,EFCAB14,ATRX,FEZ2,IKZF5,CALHM2,ENPP2,FRA10AC1,PTEN,BCLAF1,PIK3CB,OGA,RASAL2,PAN3,DCHS1,FOXO1,MOXD1,BCAP29,RGS5,TIMP2,TM2D3,CALCOCO1,DPP8,TOGARAM1,FAM217B,FAM76B,LRRFIP1,RNF217,EOGT,RERG,ADNP,CAVIN1,ESYT2,LMO7,ARID1B,PTPRD,NDFIP2,GID4,ITIH5,RSBN1L,DYRK1A,PDE4D,STRN,MARF1,CEP290,NEK1,GNG2,WSB1,TRAK2,GATA6,KCND3,DPYSL3,NPTN,RANBP9,CYBRD1,TJP1,LMBRD2,KANK2,ARNT,MEGF9,CHURC1,FSTL1,JDP2,COLEC12,MICU2,ARHGEF12,FRYL,NGRN,NSUN3,ASB2,TAB3,TBC1D5,LIN7C,RAB23,TSPYL2,SPAG9,TAB2,N4BP2L2,FBXL4,L3HYPDH,ZNF24,ASCC3,ZFAND5,EPB41L1,RHOT1,KLHL28,RSF1,THAP2,CLK4,AMN1,ZUP1,USP30,ATXN7L3,PPP1R3D,GPRASP2,SDHC,FBXO34,LIX1L,ARIH1,DIPK2B,BRSK1,ARID4B,MEIS2,TOP2B,HSPA12A,HS2ST1,FRZB,INMT,FOXJ3,BMI1,SNX3,BNIP3L,S1PR3,MIER1,GET1,NBR1,CYTH3,FGFR1,SETD5,FOXF1,BRAF,MAP1A,PTPN14,LAMA4,DPYD,SSR1,CHRDL1,YWHAZ,USP33,DCN,R3HDM1,KCTD12,NT5DC3,PPP1R9B,BRD2,LSM11,DPYSL2,BTBD9,ZFC3H1,NFYB,SP3,DNAJC3,HSPB7,HIPK1,PIP4P2,SIK3,PAFAH1B1,TXNIP,MARCHF6,LTN1,A2M,ATM,IFT57,ARMCX2,PRUNE2,BBOF1,YWHAE,PAPSS1,RSBN1,NDRG3,RBM27,ATG12,NR1D2,PDZRN3,PIBF1,MICAL2,AHI1,CAMK2N1,RORA,CNOT6L,ATP8B2,GPC6,PGAP1,USP53,ACYP2,PARP11,JPH2,ASF1A,UBR1,PHF12,CLN5,SNRNP48,RC3H1,DACT3,UBE2J1,AP2B1,SLC12A6,SMG1,MTURN,XIAP,BCLAF3,CD34,TMEM43,LPAR1,DNAJB14,PPM1B,KIFBP,MCM9,IFT80,HAND2,MRGPRF,ZCCHC14,NFIC,FILIP1L,CREBRF,FOXN3,KLF3,LATS1,AKT3,ATE1,RAB2B,PGBD1,EPS15,CAMK1,ENC1,SGTB,TADA2A,DCAF6,SYNRG,GNB1,PDGFD,MCFD2,TUBGCP3,ROCK2,SMAD7,MYH9,C18orf32,WAC,GK5,NOL4L,AKTIP,SREK1IP1,C1R,ANXA6,FXR2,TEAD3,ARPP19,FAM171A2,LYRM7,FLT1,CBX5,MYH11,EPM2AIP1,PGM3,DENND2A,FAM168A,ANKFY1,DIXDC1,NUCKS1,MAPK10,CUL4B,TBC1D15,BCL2,SESN1,CEP63,STARD13,FKBP5,EXOC2,SEC24B,KATNBL1,WDTC1,TBP,SLX4IP,DCAKD,BNIP2,PDPK1,UBXN7,LRCH3,CNOT4,SON,RSU1,RLIM,ZMIZ1,CNRIP1,EFEMP1,BRWD1,CSTF3,CHD3,BORCS7,OLFML2A,RBMS3,NAPEPLD,PDK2,DCP1B,MLLT11,RO60,TBCK,ACBD3,BBS10,SP4,RBM43,BCL6B,LNPEP,ZNF34,HDHD2,KIAA0753,DCAF17,VPS41,KIFAP3,ZHX3,MAP4K5,SBDS,RNGTT,FBXW11,KDR,ZDHHC20,SLC9B2,NCOR1,MEAF6,SCMH1,GNB4,GPR176,NFIB,SPOP,WDR36,MED4,GAS7,RPS6KC1,KANSL1,CD302,SYNJ1,JCAD,FRY,DDX5,CWF19L2,TMEM130,DENND2B,CCND2,PDGFC,LUC7L2,IRF2BP2,GAPVD1,ARHGAP5,IQSEC1,KMT2A,ADH1B,REV1,TAGLN,NRIP2,RAB5B,FAM135A,DPT,ETV3,RWDD2A,CAMKK2,JUN,HACE1,GABPB2,KLF9,TMEM167B,ODF2L,TAF9B,JAZF1,RHOQ,TBL1XR1,NRF1,MAPK14,WASHC4,CCDC92,PTGIS,NSRP1,SPTLC1,FNDC3A,CCNC,PMP22,ZC3H7A,RILPL1,RPS6KA2,SSPN,DCAF8,SPRYD7,ABCA5,TSPAN18,NKIRAS1,MAP3K4,CEP350,CLASP2,ATP2B1,EXTL2,RICTOR,SNX16,CTSO,ARL6,RAB11FIP2,VAMP2,SYT11,ANKRD40,COPA,TMEM50A,TENT4B,PTTG1IP,RAB22A,SCAPER,EZH1,FBXW2,DYNC2I1,RRAS,C12orf76,FAM228B,HEXIM1,ELK4,USF3,LMBRD1,FBXO11,CWC25,ZMAT1,IRAG1,CSGALNACT2,RERE,APPBP2,CNOT2,EDNRB,RBM48,GEM,PIKFYVE,RAB3GAP1,ULK2,LEPR,ETS1,ID2,PTPRA,PID1,KLHL7,TSC1,USPL1,TTC33,LACC1,PRPF4B | | Exist in public source | | NA |
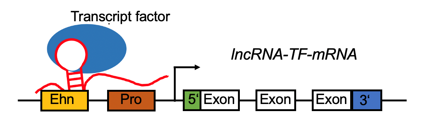
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
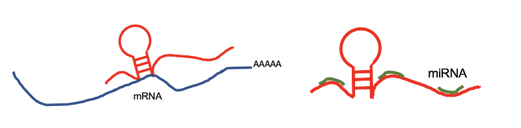
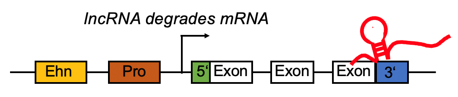
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | PSMB5,TMEM147,FIBP,RUVBL1,REX1BD,SMIM12,TOMM40,ENDOG,NUDT8,NDUFA13,NUDT22,MLST8,LAMTOR2,LRRC45,CENPX,AURKAIP1,NADSYN1,ALDOA,CDK16,NUDT16L1,GSTP1,ANAPC11,CSK,GALE,EIF3B,MAD2L2,SMAGP,CAD,TBRG4,BABAM1,CTU2,NME1,SLC25A39,TUBA1C,BCL2L12,GYS1,CLPP,RUVBL2,PAFAH1B3,REEP4,AUP1,POLR2H,LONP1,MCM5,POLRMT,DMAC2,NOP2,TIMM17B,DUS1L,MYO19,COX4I1,CCNQ,TIMM44,EIF2B3,YIF1A,MAZ,GPX4,ANTKMT,SIL1,QARS1,PNKP,POLR2J,FAM174C,GMPPB,STXBP2,ALG3,STX10,TSR3,SCAND1,ZNRD2,POP5,KRTCAP3,APBA3,FLAD1,LSR,WDR83OS,UQCRC1,NCAPH2,AAAS,TONSL,BOP1,DDX39A,JPT1,TIMM10,THOP1,NDUFB10,TK1,PPP1CA,POLD2,PCCB,DKC1,PKMYT1,APRT,ECE2,FEN1,RAN,PKP3,ECSIT,MRPL41,ATG101,GAPDH,RBM10,SDF2L1,TEDC1,RECQL4,TEAD4,CYBC1,TKT,GPX1,MED16,E2F4,SLC25A3,NOP56,NDUFV1,RBM42,SEC61A1,RPS8,CPSF4,WDR18,RNH1,YDJC,MRPL37,RABGGTA,IRAK1,WDR74,DNPEP,ETHE1,BIRC5,MPND,C19orf25,PSME2,NMRAL1,ZNHIT2,RPL36,SNRPB,ADRM1,EIF3G,TACO1,RPN1,MRPL54,PPP1R14B,CINP,DTYMK,ATG4D,CIAPIN1,SURF2,AP2S1,ECI1,PES1,MRPS11,TSEN54,STOML2,DUS3L,RPSA,DDX56,NDUFB11,QPCTL,CLN6,AURKB,FARSA,NDUFS3,COA4,TSPO,MCM7,SSR4,NAXE,LMAN2,CNOT9,NAA10,CENPM,ASF1B,UBE2J2,GFER,CCNF,NCLN,U2AF2,NOP16,TBL3,TRPT1,ORMDL2,OXLD1,MAPKAPK3,WDR4,RCC1,CHTF18,SRM,CCNB2,GCDH,FAM207A,PSMG3,PSMD8,PYCR3,NDUFS8,TMEM258,NOC4L,NDUFS7,RPL10,FASTK,NSUN5,SNRPA,PSMC4,DCPS,MCAT,RPS6KB2,CENPN,UBE2S,OTUB1,POLR2E,TRIM28,NTMT1,COX5A,MCRS1,UBA52,YIF1B,EIF3K,TARBP2,TAF10,AP1S1,CDC45,TELO2,TEDC2,ATP5MF,ENTR1,ZMYND19,LAMTOR1,MRPL34,MRPL11,INTS11,CRB3,CCNB1,SUGP1,KLF16,NR2C2AP,DPH2,COMMD4,CHEK1,PLK1,NUBP2,TRAF7,PRMT1,APEX1,ILVBL | | Exist in public source | | NA |
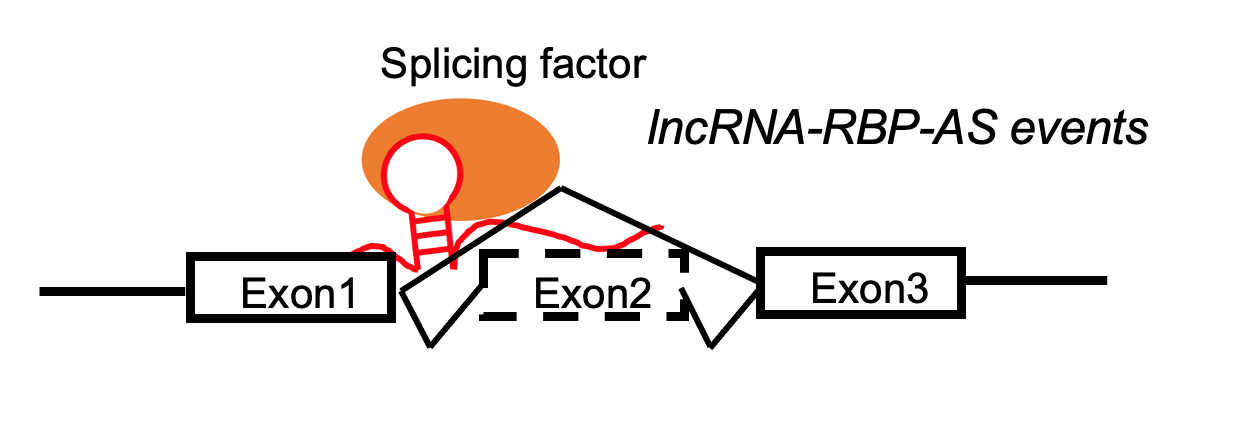
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000279453 | ENST00000624649 | exon_skip_1717 | chr1:15721328-15721388 | -8.72 | -0.2642 | PLEKHM2 | ENST00000375799 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_111672 | chr14:23089981-23090101 | -13.56 | -0.1384 | ACIN1 | ENST00000262710 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_302353 | chr19:11033766-11033865 | -16.52 | -0.2294 | SMARCA4 | ENST00000344626,ENST00000429416 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_313472 | chr19:5216719-5216767 | -8.19 | -0.1998 | PTPRS | ENST00000587303 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_21610 | chr1:9737497-9737554 | -8.89 | -0.2020 | CLSTN1 | ENST00000377298 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_21628 | chr1:9756480-9756510 | -8.25 | -0.4853 | CLSTN1 | ENST00000377298 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_33516 | chr1:156938417-156938513 | -14.48 | -0.1810 | ARHGEF11 | ENST00000361409 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_43507 | chr10:84499874-84499959 | -11.54 | -0.2098 | CCSER2 | ENST00000224756 | Frame-shift | | ENSG00000279453 | ENST00000624649 | exon_skip_48327 | chr10:26771074-26771089 | -4.80 | -0.4364 | ABI1 | ENST00000376142 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_59439 | chr11:57737912-57738026 | -10.53 | -0.1003 | TMX2 | ENST00000278422 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_96112 | chr12:109945259-109945349 | -13.34 | -0.1933 | GIT2 | ENST00000355312 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_97739 | chr12:121244572-121244615 | -8.36 | -0.3215 | CAMKK2 | ENST00000324774,ENST00000402834,ENST00000404169 | Frame-shift | | ENSG00000279453 | ENST00000624649 | exon_skip_287874 | chr17:29085603-29085648 | -6.54 | -0.1817 | MYO18A | ENST00000527372 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_307316 | chr19:40622949-40623021 | -13.04 | -0.2460 | LTBP4 | ENST00000308370 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_335378 | chr2:237751199-237751271 | -10.45 | -0.2133 | LRRFIP1 | ENST00000392000 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_353172 | chr20:58898940-58898985 | -9.21 | -0.3542 | GNAS | ENST00000371085,ENST00000371100 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_382356 | chr3:37091466-37091538 | -8.13 | -0.1626 | LRRFIP2 | ENST00000336686 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_426970 | chr4:159344035-159344059 | -6.65 | -0.6045 | RAPGEF2 | ENST00000264431 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_436488 | chr5:96726793-96726859 | -10.30 | -0.2784 | CAST | ENST00000341926,ENST00000395813 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_451515 | chr6:41080810-41080897 | -16.67 | -0.1938 | NFYA | ENST00000341376 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_515330 | chrX:65524563-65524614 | -9.92 | -0.2255 | LAS1L | ENST00000374811 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_353171 | chr20:58898940-58898985 | -9.21 | -0.3542 | GNAS | ENST00000371085,ENST00000371100 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_390418 | chr3:183842297-183842447 | -12.71 | -0.1105 | PARL | ENST00000317096 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_11827 | chr1:155729041-155729122 | -12.58 | -0.2330 | DAP3 | ENST00000343043,ENST00000368336 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000279453 | ENST00000624649 | exon_skip_10577 | chr1:153642245-153642429 | -20.31 | -0.1104 | CHTOP | ENST00000368694 | Frame-shift | | ENSG00000279453 | ENST00000624649 | exon_skip_37475 | chr1:225504990-225505053 | -8.72 | -0.2907 | ENAH | ENST00000366844 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_42844 | chr10:79310923-79311184 | -24.42 | -0.1115 | ZMIZ1 | ENST00000334512 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_44853 | chr10:104010815-104010908 | -10.67 | -0.1147 | SLK | ENST00000369755 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_44998 | chr10:110132304-110132400 | -9.01 | -0.1086 | ADD3 | ENST00000356080 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_48713 | chr10:34336198-34336243 | -6.40 | -0.1561 | PARD3 | ENST00000374789 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_98384 | chr12:124327408-124327633 | -21.76 | -0.1003 | NCOR2 | ENST00000405201 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_111863 | chr14:23565815-23565892 | -13.65 | -0.2238 | AP1G2 | ENST00000308724,ENST00000397120 | Frame-shift | | ENSG00000279453 | ENST00000624649 | exon_skip_114200 | chr14:73279280-73279424 | -21.77 | -0.1861 | NUMB | ENST00000355058,ENST00000555238 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_148110 | chr17:4892401-4892512 | -12.97 | -0.1441 | MINK1 | ENST00000355280 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_374375 | chr3:49675182-49675309 | -16.67 | -0.1313 | APEH | ENST00000296456 | Frame-shift | | ENSG00000279453 | ENST00000624649 | exon_skip_423795 | chr4:55888887-55888932 | -9.26 | -0.4209 | EXOC1 | ENST00000346134,ENST00000381295 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_456460 | chr6:17771113-17771218 | -13.30 | -0.1547 | KIF13A | ENST00000259711 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_497023 | chr9:89339863-89339953 | -10.65 | -0.1210 | SECISBP2 | ENST00000375807 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_499771 | chr9:128592982-128593042 | -11.18 | -0.1928 | SPTAN1 | ENST00000372731 | In-frame | | ENSG00000279453 | ENST00000624649 | exon_skip_325891 | chr2:62987930-62988038 | -9.38 | -0.3752 | EHBP1 | ENST00000263991 | In-frame |
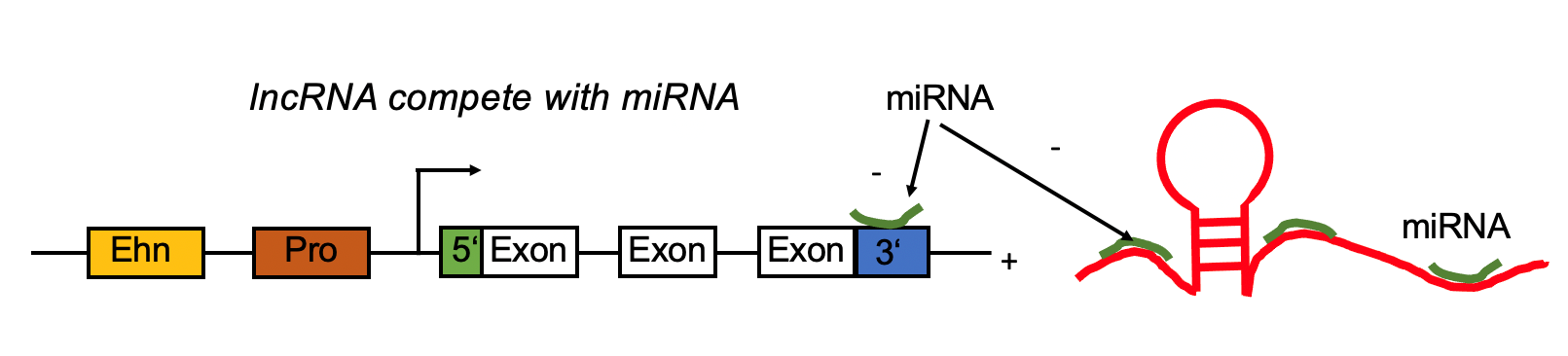
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
 |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122436931-122437013 | 186.00 | -77.86 | 85 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122438210-122438293 | 181.00 | -64.09 | 79 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122438030-122438109 | 174.00 | -66.08 | 81 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122436838-122436920 | 163.00 | -73.54 | 71 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122437715-122437797 | 162.00 | -76.93 | 80 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122437613-122437694 | 161.00 | -71.41 | 83 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122437287-122437359 | 143.00 | -68.90 | 80 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122437507-122437587 | 141.00 | -62.95 | 80 | NA | | ENSG00000279453 | hsa-mir-106b | ENST00000624649 | chr6:122438621-122438709 | 140.00 | -70.86 | 86 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437965-122438047 | 206.00 | -79.93 | 76 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122436949-122437046 | 174.00 | -77.90 | 97 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122438333-122438412 | 170.00 | -60.80 | 81 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437154-122437242 | 160.00 | -61.25 | 89 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437061-122437161 | 159.00 | -69.15 | 95 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437565-122437638 | 158.00 | -67.75 | 81 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437755-122437841 | 158.00 | -69.49 | 85 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122436839-122436918 | 152.00 | -65.91 | 84 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122438057-122438140 | 152.00 | -61.70 | 73 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437631-122437711 | 151.00 | -60.94 | 81 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122438888-122438986 | 147.00 | -62.92 | 89 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122436790-122436846 | 146.00 | -65.77 | 57 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437276-122437357 | 146.00 | -60.80 | 78 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122438197-122438276 | 145.00 | -51.45 | 78 | NA | | ENSG00000279453 | hsa-mir-25 | ENST00000624649 | chr6:122437360-122437438 | 143.00 | -54.66 | 82 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122437709-122437829 | 175.00 | -128.00 | 106 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122437594-122437696 | 173.00 | -116.84 | 100 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122437960-122438052 | 169.00 | -108.73 | 92 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122436997-122437087 | 157.00 | -104.63 | 91 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122438853-122438955 | 153.00 | -103.38 | 86 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122438104-122438196 | 152.00 | -107.89 | 82 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122438599-122438687 | 151.00 | -94.47 | 79 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122436790-122436870 | 146.00 | -107.07 | 83 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122437092-122437186 | 146.00 | -117.53 | 83 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122437291-122437384 | 146.00 | -104.83 | 74 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122437400-122437497 | 145.00 | -98.90 | 91 | NA | | ENSG00000279453 | hsa-mir-629 | ENST00000624649 | chr6:122436852-122436949 | 141.00 | -110.68 | 85 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437596-122437674 | 185.00 | -93.00 | 83 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437841-122437921 | 183.00 | -80.80 | 83 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438957-122439048 | 181.00 | -86.48 | 88 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438096-122438187 | 176.00 | -85.63 | 84 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438725-122438798 | 171.00 | -67.81 | 76 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437427-122437515 | 170.00 | -77.51 | 78 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437130-122437218 | 169.00 | -88.82 | 85 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437933-122438024 | 166.00 | -88.79 | 93 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438607-122438704 | 166.00 | -81.92 | 90 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438350-122438444 | 163.00 | -74.58 | 87 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438528-122438615 | 161.00 | -70.84 | 80 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437047-122437135 | 157.00 | -82.22 | 86 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122438211-122438308 | 156.00 | -78.54 | 93 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122436797-122436895 | 154.00 | -99.32 | 94 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437732-122437812 | 154.00 | -88.66 | 82 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122439038-122439126 | 151.00 | -82.40 | 83 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437269-122437356 | 150.00 | -84.48 | 77 | NA | | ENSG00000279453 | hsa-mir-942 | ENST00000624649 | chr6:122437346-122437432 | 147.00 | -79.26 | 83 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
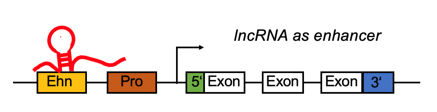
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000279453 | ENSG00000003096 | KLHL13 | ACC,KICH,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000008869 | HEATR5B | ACC,COAD,GBM,KICH,LIHC,READ,THYM | | GBM | | ENSG00000279453 | ENSG00000011021 | CLCN6 | ACC,BRCA,COAD,GBM,HNSC,KICH,KIRP,LIHC,PAAD,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000085832 | EPS15 | ACC,BRCA,COAD,GBM,KICH,KIRC,LAML,LIHC,PAAD,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000088367 | EPB41L1 | ACC,GBM,KICH,LGG,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000100030 | MAPK1 | ACC,CHOL,COAD,GBM,LGG,LIHC,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000102471 | NDFIP2 | ACC,GBM,LGG,LIHC,SKCM,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000107669 | ATE1 | ACC,GBM,KICH,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000111897 | SERINC1 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,HNSC,KICH,KIRC,LGG,LIHC,OV,PAAD,READ,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC,UVM | | BLCA,CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000112305 | SMAP1 | ACC,CHOL,GBM,KICH,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000112339 | HBS1L | ACC,CHOL,COAD,GBM,KICH,KIRC,LIHC,PAAD,SARC,SKCM,UCS,UVM | | GBM | | ENSG00000279453 | ENSG00000113391 | FAM172A | ACC,BLCA,BRCA,CESC,CHOL,COAD,KICH,LAML,LIHC,PAAD,READ,SKCM,STAD,TGCT,THCA,THYM,UCEC,UVM | | BLCA | | ENSG00000279453 | ENSG00000115137 | DNAJC27 | ACC,BRCA,GBM,LGG,PAAD,STAD,THYM | | GBM | | ENSG00000279453 | ENSG00000115295 | CLIP4 | ACC,GBM,LGG,PAAD,STAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000115839 | RAB3GAP1 | ACC,COAD,GBM,KICH,KIRP,LAML,LGG,LIHC,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000116750 | UCHL5 | ACC,CHOL,GBM,KICH,KIRC,LGG,UVM | | GBM | | ENSG00000279453 | ENSG00000120688 | WBP4 | ACC,BLCA,BRCA,CESC,CHOL,COAD,GBM,KICH,KIRC,KIRP,PAAD,READ,SKCM,THYM,UCEC,UVM | | CESC | | ENSG00000279453 | ENSG00000121964 | GTDC1 | ACC,COAD,GBM,KICH,KIRP,LGG,SKCM,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000122042 | UBL3 | ACC,CESC,COAD,LAML,PAAD,READ,THCA,THYM,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000123091 | RNF11 | ACC,BRCA,COAD,GBM,KIRC,LGG,LIHC,PAAD,READ,STAD,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000123200 | ZC3H13 | ACC,BLCA,CHOL,COAD,ESCA,GBM,KICH,KIRP,LAML,PAAD,READ,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000124422 | USP22 | ACC,GBM,KICH,KIRP,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000127870 | RNF6 | ACC,CHOL,GBM,LGG,PAAD,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000128989 | ARPP19 | ACC,COAD,GBM,KIRC,KIRP,LGG,PAAD,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000133104 | SPART | ACC,COAD,KICH,READ,STAD,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000134318 | ROCK2 | ACC,COAD,KICH,LIHC,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000136100 | VPS36 | ACC,BLCA,BRCA,CHOL,COAD,GBM,KICH,KIRC,KIRP,LAML,OV,READ,SKCM,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000136160 | EDNRB | ACC,BRCA,KICH,KIRC,KIRP,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000140299 | BNIP2 | ACC,BLCA,BRCA,CHOL,COAD,ESCA,KICH,KIRC,KIRP,LIHC,READ,STAD,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000140386 | SCAPER | ACC,COAD,GBM,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000150457 | LATS2 | ACC,BLCA,COAD,ESCA,KICH,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000152102 | FAM168B | ACC,CHOL,GBM,KICH,PAAD,STAD,TGCT,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000152484 | USP12 | ACC,COAD,ESCA,GBM,KICH,LAML,LGG,PAAD,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000152894 | PTPRK | ACC,GBM,KICH,KIRC,KIRP,LIHC,OV,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000156642 | NPTN | ACC,COAD,GBM,LGG,STAD,TGCT,THCA,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000157625 | TAB3 | ACC,COAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000162869 | PPP1R21 | ACC,GBM,KICH,PAAD,READ,THCA,UCEC | | GBM | | ENSG00000279453 | ENSG00000163539 | CLASP2 | ACC,COAD,GBM,LIHC,PAAD,STAD,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000165572 | KBTBD6 | ACC,GBM,KICH,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000165678 | GHITM | ACC,CHOL,GBM,LGG,UVM | | GBM | | ENSG00000279453 | ENSG00000165802 | NSMF | ACC,CHOL,GBM,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000174405 | LIG4 | ACC,COAD,GBM,KICH,KIRP,OV,PAAD,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000179630 | LACC1 | ACC,CHOL,COAD,GBM,HNSC,READ,THYM | | GBM | | ENSG00000279453 | ENSG00000182768 | NGRN | ACC,CHOL,GBM,KICH,KIRP,LGG,LIHC,PAAD,TGCT,THYM,UCS,UVM | | GBM | | ENSG00000279453 | ENSG00000183741 | CBX6 | ACC,GBM,KICH,LGG,PAAD,STAD,TGCT,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000197183 | NOL4L | ACC,GBM,KIRP,TGCT,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000215712 | TMEM242 | ACC,CHOL,GBM,KICH,KIRC,LGG,LIHC,PAAD,READ,SKCM,TGCT,THYM,UCS,UVM | | GBM | | ENSG00000279453 | ENSG00000220205 | VAMP2 | ACC,CESC,CHOL,ESCA,GBM,KICH,LGG,PAAD,READ,STAD,TGCT,UCEC | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000004776 | HSPB6 | BLCA,CHOL,KICH,READ,STAD,TGCT,UCEC,UCS | | BLCA,UCEC | | ENSG00000279453 | ENSG00000005812 | FBXL3 | BLCA,BRCA,CESC,COAD,KICH,KIRC,LAML,LIHC,PAAD,READ,SARC,SKCM,TGCT,THYM,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000012822 | CALCOCO1 | BLCA,BRCA,CESC,COAD,ESCA,HNSC,KICH,KIRP,LIHC,OV,PAAD,READ,SARC,STAD,TGCT,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000018408 | WWTR1 | BLCA,COAD,ESCA,KICH,KIRP,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000020181 | ADGRA2 | BLCA,KICH,PAAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000058272 | PPP1R12A | BLCA,COAD,ESCA,KIRP,LAML,LIHC,PAAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000059915 | PSD | BLCA,GBM,LGG,PAAD,READ,STAD,TGCT,UCEC,UCS | | BLCA,GBM,UCEC | | ENSG00000279453 | ENSG00000065534 | MYLK | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000069702 | TGFBR3 | BLCA,COAD,ESCA,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000072952 | IRAG1 | BLCA,CESC,COAD,ESCA,GBM,READ,STAD,TGCT,UCEC | | BLCA,CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000073712 | FERMT2 | BLCA,COAD,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000075073 | TACR2 | BLCA,ESCA,READ,STAD,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000076555 | ACACB | BLCA,ESCA,PAAD,READ,STAD,TGCT,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000079308 | TNS1 | BLCA,COAD,ESCA,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000095637 | SORBS1 | BLCA,COAD,ESCA,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000100307 | CBX7 | BLCA,BRCA,CESC,COAD,ESCA,GBM,KICH,LGG,PAAD,READ,STAD,THCA,THYM,UCEC,UVM | | BLCA,CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000100628 | ASB2 | BLCA,ESCA,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000101335 | MYL9 | BLCA,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000103710 | RASL12 | BLCA,CESC,COAD,ESCA,KICH,PAAD,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000104936 | DMPK | BLCA,ESCA,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000106772 | PRUNE2 | BLCA,COAD,ESCA,KICH,READ,STAD,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000107758 | PPP3CB | BLCA,BRCA,COAD,ESCA,GBM,KICH,KIRC,LAML,LGG,LIHC,PAAD,STAD,TGCT,THCA,THYM,UCEC,UVM | | BLCA,GBM | | ENSG00000279453 | ENSG00000107796 | ACTA2 | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000108823 | SGCA | BLCA,ESCA,READ,STAD,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000109686 | SH3D19 | BLCA,BRCA,KICH,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000111077 | TNS2 | BLCA,CESC,ESCA,PAAD,READ,STAD,TGCT,THCA,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000112210 | RAB23 | BLCA,COAD,KICH,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000112624 | BICRAL | BLCA,CESC,COAD,ESCA,GBM,KICH,KIRP,PAAD,READ,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000113448 | PDE4D | BLCA,COAD,KIRP,READ,STAD | | BLCA | | ENSG00000279453 | ENSG00000113595 | TRIM23 | BLCA,BRCA,CESC,COAD,GBM,HNSC,LAML,LGG,LIHC,PAAD,READ,STAD,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000113657 | DPYSL3 | BLCA,COAD,ESCA,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000114698 | PLSCR4 | BLCA,BRCA,COAD,ESCA,KICH,KIRC,KIRP,PAAD,READ,SARC,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000118496 | FBXO30 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,LAML,LIHC,OV,PAAD,READ,SKCM,STAD,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000118729 | CASQ2 | BLCA,ESCA,KICH,READ,STAD | | BLCA | | ENSG00000279453 | ENSG00000121440 | PDZRN3 | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000121577 | POPDC2 | BLCA,COAD,KICH,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000123096 | SSPN | BLCA,COAD,ESCA,HNSC,KICH,READ,STAD,TGCT,THCA,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000124212 | PTGIS | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000124831 | LRRFIP1 | BLCA,COAD,KICH,KIRP,READ,TGCT,UCEC | | BLCA | | ENSG00000279453 | ENSG00000125868 | DSTN | BLCA,ESCA,KICH,KIRC,KIRP,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000131171 | SH3BGRL | BLCA,CESC,COAD,ESCA,KIRC,PAAD,READ,STAD,TGCT,UCEC,UVM | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000131471 | AOC3 | BLCA,COAD,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000131831 | RAI2 | BLCA,CESC,ESCA,GBM,HNSC,KIRP,PAAD,READ,STAD,TGCT,THYM,UVM | | BLCA,CESC,GBM | | ENSG00000279453 | ENSG00000133392 | MYH11 | BLCA,CESC,KICH,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000135842 | NIBAN1 | BLCA,COAD,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000137094 | DNAJB5 | BLCA,COAD,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000137601 | NEK1 | BLCA,COAD,LAML,PAAD,READ,STAD,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000140092 | FBLN5 | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000143337 | TOR1AIP1 | BLCA,CESC,COAD,ESCA,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000143776 | CDC42BPA | BLCA,COAD,GBM,KICH,READ,TGCT,THYM,UVM | | BLCA,GBM | | ENSG00000279453 | ENSG00000145012 | LPP | BLCA,COAD,ESCA,KICH,KIRP,LAML,LIHC,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000145936 | KCNMB1 | BLCA,CESC,COAD,ESCA,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000149591 | TAGLN | BLCA,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000149596 | JPH2 | BLCA,COAD,ESCA,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000150764 | DIXDC1 | BLCA,BRCA,COAD,ESCA,GBM,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,GBM,UCEC | | ENSG00000279453 | ENSG00000152583 | SPARCL1 | BLCA,BRCA,CESC,COAD,ESCA,PAAD,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000153179 | RASSF3 | BLCA,COAD,KICH,THYM,UCEC | | BLCA | | ENSG00000279453 | ENSG00000154175 | ABI3BP | BLCA,ESCA,KICH,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000154553 | PDLIM3 | BLCA,COAD,ESCA,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000156650 | KAT6B | BLCA,COAD,KICH,PAAD,READ,THCA,THYM,UVM | | BLCA | | ENSG00000279453 | ENSG00000156804 | FBXO32 | BLCA,CESC,COAD,ESCA,LAML,READ,STAD,TGCT,UCEC,UVM | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000157570 | TSPAN18 | BLCA,COAD,KICH,READ,STAD,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000162614 | NEXN | BLCA,COAD,ESCA,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000163297 | ANTXR2 | BLCA,COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000163431 | LMOD1 | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000163820 | FYCO1 | BLCA,COAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000164530 | PI16 | BLCA,ESCA,KICH,READ,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000164764 | SBSPON | BLCA,KICH,STAD,TGCT,THYM | | BLCA | | ENSG00000279453 | ENSG00000165410 | CFL2 | BLCA,COAD,PAAD,READ,STAD,TGCT,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000165424 | ZCCHC24 | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000166780 | BMERB1 | BLCA,CESC,ESCA,GBM,KICH,LGG,PAAD,READ,STAD,TGCT,THCA,UCEC,UVM | | BLCA,CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000168386 | FILIP1L | BLCA,COAD,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000168497 | CAVIN2 | BLCA,BRCA,COAD,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000170271 | FAXDC2 | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000170464 | DNAJC18 | BLCA,ESCA,KICH,PAAD,STAD,TGCT,THCA,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000172348 | RCAN2 | BLCA,COAD,GBM,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,GBM,UCEC | | ENSG00000279453 | ENSG00000172935 | MRGPRF | BLCA,CESC,COAD,ESCA,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000173175 | ADCY5 | BLCA,GBM,LGG,STAD,TGCT | | BLCA,GBM | | ENSG00000279453 | ENSG00000173641 | HSPB7 | BLCA,COAD,ESCA,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000182534 | MXRA7 | BLCA,COAD,KICH,PAAD,READ,STAD,TGCT,THCA,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000182700 | IGIP | BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,HNSC,KIRC,KIRP,LAML,LGG,PAAD,READ,SARC,SKCM,STAD,THYM,UCEC,UVM | | BLCA,CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000184347 | SLIT3 | BLCA,BRCA,COAD,HNSC,KICH,READ,STAD,TGCT,THCA,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000185437 | SH3BGR | BLCA,ESCA,READ,STAD,THYM,UVM | | BLCA | | ENSG00000279453 | ENSG00000187239 | FNBP1 | BLCA,COAD,READ,STAD,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000196557 | CACNA1H | BLCA,ESCA,KICH,PAAD,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000196616 | ADH1B | BLCA,KICH,STAD,THYM,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000196663 | TECPR2 | BLCA,GBM,PAAD,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000197256 | KANK2 | BLCA,CESC,COAD,ESCA,HNSC,KICH,READ,STAD,TGCT,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000197321 | SVIL | BLCA,COAD,KICH,KIRP,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000197380 | DACT3 | BLCA,COAD,ESCA,GBM,READ,STAD,TGCT,THYM,UCEC | | BLCA,GBM,UCEC | | ENSG00000279453 | ENSG00000198467 | TPM2 | BLCA,ESCA,KICH,READ,STAD,TGCT,UCEC | | BLCA,UCEC | | ENSG00000279453 | ENSG00000198961 | PJA2 | BLCA,BRCA,CESC,CHOL,COAD,ESCA,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000240771 | ARHGEF25 | BLCA,COAD,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000255302 | EID1 | BLCA,COAD,KIRC,PAAD,READ,STAD,TGCT,THCA,THYM,UCEC,UVM | | BLCA,UCEC | | ENSG00000279453 | ENSG00000265972 | TXNIP | BLCA,BRCA,CESC,COAD,ESCA,KICH,KIRP,READ,STAD,TGCT,THCA,THYM,UCEC | | BLCA,CESC,UCEC | | ENSG00000279453 | ENSG00000008311 | AASS | BRCA,KICH,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000011523 | CEP68 | BRCA,COAD,GBM,KICH,PAAD,READ,STAD,TGCT,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000031003 | FAM13B | BRCA,COAD,ESCA,GBM,PAAD,READ,STAD,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000049246 | PER3 | BRCA,COAD,READ,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000069122 | ADGRF5 | BRCA,COAD,PAAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000071242 | RPS6KA2 | BRCA,GBM,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000073910 | FRY | BRCA,ESCA,GBM,KICH,STAD | | GBM | | ENSG00000279453 | ENSG00000074935 | TUBE1 | BRCA,CESC,COAD,KIRP,OV,PAAD,THYM,UCEC | | CESC | | ENSG00000279453 | ENSG00000080546 | SESN1 | BRCA,PAAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000085382 | HACE1 | BRCA,CESC,GBM,KICH,KIRC,LAML,SKCM,TGCT,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000088387 | DOCK9 | BRCA,COAD,GBM,KICH,READ,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000102781 | KATNAL1 | BRCA,CESC,ESCA,GBM,KICH,KIRP,PAAD,STAD,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000106771 | TMEM245 | BRCA,COAD,GBM,LAML,PAAD,READ,STAD,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000107560 | RAB11FIP2 | BRCA,COAD,GBM,KICH,KIRC,KIRP,LAML,LIHC,READ,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000108799 | EZH1 | BRCA,CESC,COAD,ESCA,GBM,KICH,KIRP,OV,PAAD,READ,SKCM,STAD,THCA,THYM,UCEC,UVM | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000113594 | LIFR | BRCA,KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000116678 | LEPR | BRCA,KICH,KIRP,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000120156 | TEK | BRCA,COAD,KICH,PAAD,UCEC | | UCEC | | ENSG00000279453 | ENSG00000124942 | AHNAK | BRCA,COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000128872 | TMOD2 | BRCA,ESCA,GBM,LGG,PAAD,STAD | | GBM | | ENSG00000279453 | ENSG00000128881 | TTBK2 | BRCA,GBM,KIRP,LGG,PAAD,STAD,THYM,UCS | | GBM | | ENSG00000279453 | ENSG00000135338 | LCA5 | BRCA,KIRC,KIRP,SARC,TGCT,THCA,UCEC | | UCEC | | ENSG00000279453 | ENSG00000138593 | SECISBP2L | BRCA,COAD,GBM,LIHC,PAAD,READ,STAD,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000140199 | SLC12A6 | BRCA,COAD,GBM,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000141447 | OSBPL1A | BRCA,GBM,LGG,STAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000147113 | DIPK2B | BRCA,CESC,KICH,PAAD,TGCT,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000150938 | CRIM1 | BRCA,COAD,KICH,KIRC,LAML,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000151914 | DST | BRCA,COAD,GBM,READ,STAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000152492 | CCDC50 | BRCA,COAD,ESCA,KICH,KIRC,KIRP,LIHC,READ,STAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000154265 | ABCA5 | BRCA,COAD,GBM,PAAD,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000156052 | GNAQ | BRCA,COAD,GBM,KIRC,KIRP,LGG,READ,THCA,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000162407 | PLPP3 | BRCA,READ,SARC,TGCT,THCA,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000162616 | DNAJB4 | BRCA,COAD,KICH,KIRC,READ,STAD,THYM,UCEC,UCS | | UCEC | | ENSG00000279453 | ENSG00000162618 | ADGRL4 | BRCA,COAD,KICH,KIRC,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000164035 | EMCN | BRCA,COAD,KICH,KIRC,PAAD,READ,STAD,TGCT,THYM,UCEC,UCS | | UCEC | | ENSG00000279453 | ENSG00000164463 | CREBRF | BRCA,CESC,COAD,ESCA,KIRP,LAML,LIHC,PAAD,READ,SKCM,STAD,THCA,THYM,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000164576 | SAP30L | BRCA,CHOL,KICH,PAAD,READ,SKCM,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000169744 | LDB2 | BRCA,CESC,COAD,GBM,KICH,LGG,PAAD,READ,TGCT,UCEC,UVM | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000170962 | PDGFD | BRCA,KICH,KIRC,KIRP,READ,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000170989 | S1PR1 | BRCA,CESC,COAD,KICH,TGCT,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000174059 | CD34 | BRCA,CESC,KICH,PAAD,TGCT,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000178695 | KCTD12 | BRCA,COAD,PAAD,READ,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000180354 | MTURN | BRCA,COAD,GBM,KICH,LGG,PAAD,READ,STAD,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000183722 | LHFPL6 | BRCA,CESC,COAD,KICH,PAAD,READ,STAD,TGCT,THCA,THYM,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000184500 | PROS1 | BRCA,COAD,ESCA,KICH,READ,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000186642 | PDE2A | BRCA,ESCA,GBM,KICH,LGG,STAD | | GBM | | ENSG00000279453 | ENSG00000187866 | PABIR1 | BRCA,COAD,KIRC,LAML,PAAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000035862 | TIMP2 | CESC,COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000101938 | CHRDL1 | CESC,COAD,ESCA,READ,STAD,TGCT,THYM,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000104047 | DTWD1 | CESC,COAD,ESCA,KIRC,STAD,THYM,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000119865 | CNRIP1 | CESC,COAD,ESCA,GBM,KICH,LGG,PAAD,STAD,TGCT,THYM,UCEC | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000121361 | KCNJ8 | CESC,ESCA,KICH,THYM,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000122545 | SEPTIN7 | CESC,COAD,KICH,KIRC,KIRP,LAML,PAAD,READ,STAD,TGCT,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000128052 | KDR | CESC,KICH,PAAD,TGCT,THYM,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000134198 | TSPAN2 | CESC,COAD,ESCA,KICH,STAD,TGCT,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000134533 | RERG | CESC,COAD,READ,STAD,TGCT,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000136960 | ENPP2 | CESC,GBM,PAAD,TGCT,UCEC | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000138172 | CALHM2 | CESC,KICH,TGCT,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000139597 | N4BP2L1 | CESC,COAD,KICH,KIRP,READ,THYM,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000142599 | RERE | CESC,GBM,KICH,LIHC,READ,THCA,THYM,UCEC | | CESC | | ENSG00000279453 | ENSG00000148484 | RSU1 | CESC,KICH,READ,STAD,THYM,UCEC | | CESC | | ENSG00000279453 | ENSG00000165757 | JCAD | CESC,COAD,GBM,KICH,READ,THCA,THYM,UCEC | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000166313 | APBB1 | CESC,ESCA,GBM,KICH,LGG,PAAD,READ,STAD,TGCT,UCEC | | CESC,GBM,UCEC | | ENSG00000279453 | ENSG00000172007 | RAB33B | CESC,COAD,ESCA,KIRC,KIRP,LAML,PAAD,TGCT,THCA,THYM,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000176435 | CLEC14A | CESC,KICH,PAAD,TGCT,UCEC | | CESC,UCEC | | ENSG00000279453 | ENSG00000179476 | C14orf28 | CESC,COAD,LAML,PAAD,STAD,TGCT,THCA,THYM,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000204130 | RUFY2 | CESC,COAD,GBM,KICH,KIRP,LAML,PAAD,READ,STAD,TGCT,THCA,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000271601 | LIX1L | CESC,COAD,ESCA,KICH,PAAD,READ,STAD,TGCT,UCEC,UVM | | CESC,UCEC | | ENSG00000279453 | ENSG00000001561 | ENPP4 | CHOL,COAD,GBM,KIRC,PAAD | | GBM | | ENSG00000279453 | ENSG00000007168 | PAFAH1B1 | CHOL,COAD,GBM,KICH,KIRP,LGG,PAAD,READ,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000023287 | RB1CC1 | CHOL,COAD,GBM,LAML,LGG,THYM | | GBM | | ENSG00000279453 | ENSG00000050748 | MAPK9 | CHOL,GBM,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000051382 | PIK3CB | CHOL,COAD,GBM,KICH,LGG | | GBM | | ENSG00000279453 | ENSG00000078114 | NEBL | CHOL,GBM,KICH,KIRP,THYM | | GBM | | ENSG00000279453 | ENSG00000083642 | PDS5B | CHOL,COAD,GBM,KICH,KIRC,LAML,PAAD,READ,THCA,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000087152 | ATXN7L3 | CHOL,GBM,KICH,KIRP,LGG,LIHC,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000089818 | NECAP1 | CHOL,GBM,LGG,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000102181 | CD99L2 | CHOL,GBM,LGG,PAAD,READ,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000107957 | SH3PXD2A | CHOL,COAD,GBM,READ,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000111696 | NT5DC3 | CHOL,GBM,STAD,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000113638 | TTC33 | CHOL,LAML,PAAD,READ,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000114573 | ATP6V1A | CHOL,COAD,GBM,LGG,THYM | | GBM | | ENSG00000279453 | ENSG00000114790 | ARHGEF26 | CHOL,COAD,PAAD,READ,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000115365 | LANCL1 | CHOL,COAD,GBM,KICH,LGG,READ,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000116903 | EXOC8 | CHOL,COAD,GBM,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000120256 | LRP11 | CHOL,GBM,LGG,LIHC,TGCT,THCA,UVM | | GBM | | ENSG00000279453 | ENSG00000123178 | SPRYD7 | CHOL,GBM,KICH,LAML,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000126091 | ST3GAL3 | CHOL,PAAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000126524 | SBDS | CHOL,KICH,KIRP,READ,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000128731 | HERC2 | CHOL,GBM,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000129636 | ITFG1 | CHOL,ESCA,GBM,LGG,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000139697 | SBNO1 | CHOL,COAD,GBM,KIRP,LIHC,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000151743 | AMN1 | CHOL,GBM,LAML,LGG,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000151835 | SACS | CHOL,GBM,KIRP,PAAD,STAD | | GBM | | ENSG00000279453 | ENSG00000152409 | JMY | CHOL,COAD,GBM,PAAD,READ,STAD,THCA,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000154122 | ANKH | CHOL,GBM,PAAD,TGCT,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000154277 | UCHL1 | CHOL,GBM,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000154429 | CCSAP | CHOL,COAD,GBM,LAML,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000156735 | BAG4 | CHOL,COAD,GBM,KIRC,LGG,LIHC,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000162852 | CNST | CHOL,COAD,GBM,KICH,LGG,LIHC,PAAD,READ | | GBM | | ENSG00000279453 | ENSG00000164327 | RICTOR | CHOL,COAD,GBM,KIRP,LAML,LIHC,READ,THCA,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000164830 | OXR1 | CHOL,COAD,GBM,KIRP,LAML,LGG,TGCT,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000164924 | YWHAZ | CHOL,GBM,KIRP,LGG,UVM | | GBM | | ENSG00000279453 | ENSG00000168438 | CDC40 | CHOL,GBM,KICH,KIRC,LIHC,PAAD,READ,SKCM,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000171368 | TPPP | CHOL,GBM,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000171617 | ENC1 | CHOL,GBM,LGG,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000175073 | VCPIP1 | CHOL,COAD,GBM,LAML,READ,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000175582 | RAB6A | CHOL,GBM,KIRC,LGG,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000176871 | WSB2 | CHOL,GBM,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000213190 | MLLT11 | CHOL,COAD,GBM,PAAD,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000005243 | COPZ2 | COAD,KICH,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000007237 | GAS7 | COAD,GBM,LGG,PAAD,READ,STAD,THYM | | GBM | | ENSG00000279453 | ENSG00000010404 | IDS | COAD,ESCA,GBM,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000010810 | FYN | COAD,HNSC,KICH,PAAD,SARC,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000019995 | ZRANB1 | COAD,GBM,KICH,LGG,PAAD,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000031081 | ARHGAP31 | COAD,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000038219 | BOD1L1 | COAD,ESCA,GBM,KIRP,LIHC,OV,READ,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000053702 | NRIP2 | COAD,ESCA,GBM,KICH,PAAD,STAD,TGCT,THYM,UCEC,UCS | | GBM,UCEC | | ENSG00000279453 | ENSG00000058091 | CDK14 | COAD,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000059758 | CDK17 | COAD,GBM,KIRC,KIRP,LGG,LIHC,PAAD,READ,TGCT,THCA,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000066739 | ATG2B | COAD,GBM,LIHC,PAAD,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000066933 | MYO9A | COAD,ESCA,GBM,KICH,KIRP,PAAD,STAD,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000068024 | HDAC4 | COAD,GBM,STAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000070778 | PTPN21 | COAD,KIRP,PAAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000071967 | CYBRD1 | COAD,ESCA,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000075391 | RASAL2 | COAD,GBM,KICH,KIRP,UVM | | GBM | | ENSG00000279453 | ENSG00000075790 | BCAP29 | COAD,KIRC,PAAD,READ,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000083290 | ULK2 | COAD,GBM,KICH,KIRP,PAAD,STAD,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000083799 | CYLD | COAD,GBM,PAAD,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000085224 | ATRX | COAD,GBM,LAML,LIHC,READ,STAD,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000085433 | WDR47 | COAD,GBM,KICH,LGG,PAAD,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000087448 | KLHL42 | COAD,GBM,LAML,PAAD,READ,STAD,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000091986 | CCDC80 | COAD,ESCA,READ,STAD,TGCT,THCA,UCEC | | UCEC | | ENSG00000279453 | ENSG00000095787 | WAC | COAD,GBM,KICH,LAML,LIHC,PAAD,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000099250 | NRP1 | COAD,KICH,READ,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000100580 | TMED8 | COAD,GBM,LIHC,PAAD,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000100934 | SEC23A | COAD,KICH,LIHC,READ,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000101290 | CDS2 | COAD,ESCA,GBM,PAAD,READ,STAD,UCEC | | GBM | | ENSG00000279453 | ENSG00000103540 | CCP110 | COAD,GBM,KICH,KIRP,READ | | GBM | | ENSG00000279453 | ENSG00000103657 | HERC1 | COAD,GBM,LGG,PAAD,STAD,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000105270 | CLIP3 | COAD,ESCA,GBM,KICH,PAAD,READ,STAD,TGCT,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000106780 | MEGF9 | COAD,GBM,PAAD,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000108061 | SHOC2 | COAD,GBM,KICH,KIRC,LAML,LGG,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000108091 | CCDC6 | COAD,GBM,KICH,LGG,THYM | | GBM | | ENSG00000279453 | ENSG00000108306 | FBXL20 | COAD,GBM,KIRP,LAML,LIHC,STAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000108861 | DUSP3 | COAD,GBM,KICH,LGG,STAD,UCEC | | GBM | | ENSG00000279453 | ENSG00000109099 | PMP22 | COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000109436 | TBC1D9 | COAD,GBM,PAAD,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000109670 | FBXW7 | COAD,GBM,KIRP,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000111647 | UHRF1BP1L | COAD,GBM,LGG,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000111727 | HCFC2 | COAD,ESCA,LAML,PAAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000111961 | SASH1 | COAD,GBM,KICH,LGG,OV,PAAD,READ,SKCM,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000112769 | LAMA4 | COAD,SARC,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000113441 | LNPEP | COAD,ESCA,GBM,KIRP,LAML,PAAD,READ,STAD,TGCT,THCA,UVM | | GBM | | ENSG00000279453 | ENSG00000115380 | EFEMP1 | COAD,ESCA,KICH,READ,UCEC | | UCEC | | ENSG00000279453 | ENSG00000115419 | GLS | COAD,GBM,KICH,LAML,LGG,PAAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000115461 | IGFBP5 | COAD,READ,STAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000115808 | STRN | COAD,GBM,KICH,KIRP,LIHC,THYM | | GBM | | ENSG00000279453 | ENSG00000115977 | AAK1 | COAD,GBM,KICH,LGG,PAAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000116962 | NID1 | COAD,KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000117020 | AKT3 | COAD,GBM,KICH,PAAD,READ,STAD,TGCT,UCEC,UCS,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000120820 | GLT8D2 | COAD,READ,STAD,TGCT,THYM,UCEC,UCS | | UCEC | | ENSG00000279453 | ENSG00000121274 | TENT4B | COAD,GBM,LAML,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000123066 | MED13L | COAD,KIRP,LIHC,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000123094 | RASSF8 | COAD,ESCA,KICH,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000128923 | MINDY2 | COAD,ESCA,KIRP,READ,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000131263 | RLIM | COAD,GBM,LIHC,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000131378 | RFTN1 | COAD,KICH,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000131437 | KIF3A | COAD,ESCA,GBM,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000132718 | SYT11 | COAD,KICH,READ,STAD,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000133121 | STARD13 | COAD,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000133703 | KRAS | COAD,GBM,KIRP,LAML,UVM | | GBM | | ENSG00000279453 | ENSG00000134352 | IL6ST | COAD,ESCA,KICH,KIRC,PAAD,READ,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000135541 | AHI1 | COAD,GBM,KICH,OV,PAAD,SKCM,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000136161 | RCBTB2 | COAD,KICH,KIRP,PAAD,READ,STAD,UCEC | | UCEC | | ENSG00000279453 | ENSG00000136237 | RAPGEF5 | COAD,GBM,KICH,TGCT,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000137075 | RNF38 | COAD,KICH,KIRC,KIRP,LIHC,READ,STAD,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000137693 | YAP1 | COAD,KICH,READ,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000138061 | CYP1B1 | COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000138078 | PREPL | COAD,GBM,KICH,LAML,LGG,PAAD,SARC,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000138138 | ATAD1 | COAD,GBM,LGG,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000138641 | HERC3 | COAD,GBM,LAML,LGG,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000138735 | PDE5A | COAD,ESCA,KIRC,READ,STAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000138814 | PPP3CA | COAD,GBM,LAML,LGG,READ,TGCT,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000139190 | VAMP1 | COAD,GBM,KIRP,LGG,UVM | | GBM | | ENSG00000279453 | ENSG00000143164 | DCAF6 | COAD,GBM,LGG,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000143344 | RGL1 | COAD,KICH,PAAD,READ,TGCT,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000143515 | ATP8B2 | COAD,ESCA,HNSC,KICH,KIRP,PAAD,READ,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000144036 | EXOC6B | COAD,GBM,LGG,READ,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000144357 | UBR3 | COAD,GBM,LAML,LGG,LIHC,PAAD,READ,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000145391 | SETD7 | COAD,STAD,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000145495 | MARCHF6 | COAD,GBM,PAAD,READ,TGCT,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000145990 | GFOD1 | COAD,GBM,LGG,PAAD,STAD | | GBM | | ENSG00000279453 | ENSG00000146278 | PNRC1 | COAD,KICH,LAML,OV,PAAD,READ,SKCM,STAD,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000147027 | TMEM47 | COAD,ESCA,KICH,KIRC,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000148690 | FRA10AC1 | COAD,GBM,KICH,KIRC,LAML,READ,STAD,THCA,UVM | | GBM | | ENSG00000279453 | ENSG00000150995 | ITPR1 | COAD,ESCA,GBM,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000151229 | SLC2A13 | COAD,GBM,KIRP,LGG,PAAD,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000151240 | DIP2C | COAD,GBM,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000152061 | RABGAP1L | COAD,GBM,LAML,LGG,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000152377 | SPOCK1 | COAD,GBM,LGG,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000152404 | CWF19L2 | COAD,LAML,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000153561 | RMND5A | COAD,GBM,KICH,KIRC,LIHC,READ,THCA | | GBM | | ENSG00000279453 | ENSG00000154511 | DIPK1A | COAD,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000155099 | PIP4P2 | COAD,LGG,PAAD,READ,STAD,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000155111 | CDK19 | COAD,GBM,KICH,KIRC,KIRP,LAML,LIHC,OV,PAAD,READ,SKCM,TGCT,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000155744 | FAM126B | COAD,GBM,KICH,LGG,READ,THYM | | GBM | | ENSG00000279453 | ENSG00000156113 | KCNMA1 | COAD,ESCA,GBM,LGG,PAAD,READ,STAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000156671 | SAMD8 | COAD,GBM,KICH,LIHC,READ,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000157106 | SMG1 | COAD,GBM,KICH,KIRP,LIHC,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000157764 | BRAF | COAD,GBM,KIRP,LAML,PAAD,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000158270 | COLEC12 | COAD,HNSC,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000158301 | GPRASP2 | COAD,GBM,KICH,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000159256 | MORC3 | COAD,LIHC,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000159899 | NPR2 | COAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000160584 | SIK3 | COAD,GBM,PAAD,READ,STAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000163171 | CDC42EP3 | COAD,ESCA,KICH,PAAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000163378 | EOGT | COAD,ESCA,KICH,PAAD,STAD,UCEC | | UCEC | | ENSG00000279453 | ENSG00000163602 | RYBP | COAD,GBM,KIRP,LAML,LIHC,READ,THCA,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000163625 | WDFY3 | COAD,GBM,KICH,KIRP,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000164031 | DNAJB14 | COAD,GBM,KIRC,LAML,LGG,PAAD,READ,TGCT,THCA,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000164125 | GASK1B | COAD,HNSC,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000164176 | EDIL3 | COAD,GBM,KICH,READ,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000164187 | LMBRD2 | COAD,GBM,LGG,PAAD,READ,TGCT | | GBM | | ENSG00000279453 | ENSG00000164506 | STXBP5 | COAD,GBM,LGG,PAAD,READ,UVM | | GBM | | ENSG00000279453 | ENSG00000164741 | DLC1 | COAD,ESCA,KICH,READ,STAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000164949 | GEM | COAD,READ,STAD,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000165525 | NEMF | COAD,ESCA,GBM,LAML,LIHC,THYM | | GBM | | ENSG00000279453 | ENSG00000166147 | FBN1 | COAD,ESCA,HNSC,READ,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000166250 | CLMP | COAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000166783 | MARF1 | COAD,ESCA,GBM,KICH,LGG,READ,STAD,THCA,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000166963 | MAP1A | COAD,GBM,LGG,PAAD,READ,STAD,UVM | | GBM | | ENSG00000279453 | ENSG00000168675 | LDLRAD4 | COAD,ESCA,GBM,KICH,PAAD,READ,STAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000168944 | CEP120 | COAD,LAML,PAAD,READ,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000170456 | DENND5B | COAD,GBM,LIHC,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000171055 | FEZ2 | COAD,KICH,KIRC,LAML,READ,STAD,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000173064 | HECTD4 | COAD,GBM,KIRP,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000173706 | HEG1 | COAD,KICH,READ,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000174456 | C12orf76 | COAD,GBM,KIRP,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000176055 | MBLAC2 | COAD,GBM,KIRC,LGG,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000176542 | USF3 | COAD,GBM,LAML,LIHC,READ,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000176714 | CCDC121 | COAD,PAAD,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000177119 | ANO6 | COAD,TGCT,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000177200 | CHD9 | COAD,KICH,LAML,READ,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000177469 | CAVIN1 | COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000178700 | DHFR2 | COAD,KICH,LAML,READ,STAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000180447 | GAS1 | COAD,HNSC,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000182670 | TTC3 | COAD,GBM,HNSC,PAAD,READ,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000185565 | LSAMP | COAD,GBM,READ,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000185585 | OLFML2A | COAD,KICH,READ,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000185658 | BRWD1 | COAD,GBM,KICH,KIRP,LGG,LIHC,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000185716 | MOSMO | COAD,GBM,LAML,LGG,LIHC,READ,STAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000186318 | BACE1 | COAD,GBM,TGCT,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000186469 | GNG2 | COAD,GBM,KICH,PAAD,READ,TGCT | | GBM | | ENSG00000279453 | ENSG00000187189 | TSPYL4 | COAD,GBM,KICH,KIRP,LGG,PAAD,READ,SKCM,STAD,TGCT,THCA,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000187955 | COL14A1 | COAD,KICH,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000188641 | DPYD | COAD,ESCA,READ,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000189184 | PCDH18 | COAD,HNSC,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000196712 | NF1 | COAD,GBM,KIRP,LAML,LIHC,OV,READ,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000196950 | SLC39A10 | COAD,GBM,KICH,LAML,LGG,THYM | | GBM | | ENSG00000279453 | ENSG00000197614 | MFAP5 | COAD,ESCA,READ,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000197860 | SGTB | COAD,GBM,KICH,LGG,PAAD,READ,STAD,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000197872 | CYRIA | COAD,GBM,LGG,READ,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000197885 | NKIRAS1 | COAD,GBM,LAML,LGG,PAAD,READ,STAD,THYM,UCEC | | GBM | | ENSG00000279453 | ENSG00000197969 | VPS13A | COAD,GBM,KIRP,LAML,READ,THYM | | GBM | | ENSG00000279453 | ENSG00000198399 | ITSN2 | COAD,GBM,LIHC,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000198408 | OGA | COAD,ESCA,GBM,KIRP,LAML,READ,STAD,THCA,UVM | | GBM | | ENSG00000279453 | ENSG00000198718 | TOGARAM1 | COAD,GBM,PAAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000204116 | CHIC1 | COAD,GBM,LAML,LGG,PAAD,READ,STAD,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000204217 | BMPR2 | COAD,ESCA,GBM,KICH,KIRC,LIHC,READ,STAD,TGCT,THCA,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000204381 | LAYN | COAD,KICH,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000204681 | GABBR1 | COAD,GBM,KICH,KIRP,LGG,STAD,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000205726 | ITSN1 | COAD,GBM,KICH,READ,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000213694 | S1PR3 | COAD,ESCA,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000222009 | BTBD19 | COAD,KIRP,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000256043 | CTSO | COAD,ESCA,HNSC,KICH,LAML,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000258289 | CHURC1 | COAD,ESCA,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000008710 | PKD1 | ESCA,PAAD,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000101665 | SMAD7 | ESCA,GBM,KIRP,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000104490 | NCALD | ESCA,GBM,LGG,PAAD,STAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000107186 | MPDZ | ESCA,KICH,PAAD,STAD,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000107819 | SFXN3 | ESCA,GBM,KICH,KIRP,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000118523 | CCN2 | ESCA,KICH,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000124067 | SLC12A4 | ESCA,KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000128266 | GNAZ | ESCA,GBM,PAAD,STAD,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000129625 | REEP5 | ESCA,GBM,LGG,PAAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000139625 | MAP3K12 | ESCA,PAAD,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000171385 | KCND3 | ESCA,GBM,PAAD,STAD,UVM | | GBM | | ENSG00000279453 | ENSG00000175899 | A2M | ESCA,KICH,PAAD,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000179954 | SSC5D | ESCA,READ,STAD,TGCT,UCEC,UCS | | UCEC | | ENSG00000279453 | ENSG00000184205 | TSPYL2 | ESCA,GBM,LGG,PAAD,STAD,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000184271 | POU6F1 | ESCA,GBM,KIRP,LGG,PAAD,STAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000186111 | PIP5K1C | ESCA,GBM,STAD,TGCT,UCS | | GBM | | ENSG00000279453 | ENSG00000241644 | INMT | ESCA,KICH,STAD,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000005882 | PDK2 | GBM,KICH,LGG,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000047056 | WDR37 | GBM,KICH,LIHC,PAAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000048991 | R3HDM1 | GBM,KICH,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000049769 | PPP1R3F | GBM,LGG,PAAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000050165 | DKK3 | GBM,KICH,LGG,PAAD,STAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000054793 | ATP9A | GBM,LGG,PAAD,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000055163 | CYFIP2 | GBM,KICH,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000063180 | CA11 | GBM,LGG,TGCT,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000065559 | MAP2K4 | GBM,KICH,KIRP,LAML,LGG,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000070961 | ATP2B1 | GBM,LGG,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000074527 | NTN4 | GBM,KICH,STAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000075945 | KIFAP3 | GBM,KICH,LGG,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000082397 | EPB41L3 | GBM,LGG,PAAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000087258 | GNAO1 | GBM,LGG,PAAD,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000088854 | C20orf194 | GBM,KICH,KIRP,PAAD,SARC,STAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000091428 | RAPGEF4 | GBM,LGG,PAAD,THCA,UVM | | GBM | | ENSG00000279453 | ENSG00000091972 | CD200 | GBM,KICH,KIRP,LGG,PAAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000092964 | DPYSL2 | GBM,HNSC,KICH,PAAD,STAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000099204 | ABLIM1 | GBM,KICH,SARC,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000101079 | NDRG3 | GBM,KICH,LGG,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000102882 | MAPK3 | GBM,KICH,LGG,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000103034 | NDRG4 | GBM,LGG,TGCT,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000104381 | GDAP1 | GBM,LGG,PAAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000105227 | PRX | GBM,KICH,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000105696 | TMEM59L | GBM,LGG,TGCT,THYM,UCS | | GBM | | ENSG00000279453 | ENSG00000106976 | DNM1 | GBM,KIRP,LGG,PAAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000108395 | TRIM37 | GBM,KICH,LGG,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000108819 | PPP1R9B | GBM,KICH,LGG,TGCT,THYM,UCS | | GBM | | ENSG00000279453 | ENSG00000109339 | MAPK10 | GBM,KICH,LGG,PAAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000109466 | KLHL2 | GBM,LGG,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000110171 | TRIM3 | GBM,KICH,LGG,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000110931 | CAMKK2 | GBM,KIRP,LGG,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000111275 | ALDH2 | GBM,KICH,LGG,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000112290 | WASF1 | GBM,KICH,LGG,LIHC,PAAD,READ,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000112972 | HMGCS1 | GBM,KIRP,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000115310 | RTN4 | GBM,LGG,TGCT,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000119242 | CCDC92 | GBM,KIRP,LGG,PAAD,STAD,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000119965 | C10orf88 | GBM,KICH,KIRP,LAML,LGG,PAAD,UVM | | GBM | | ENSG00000279453 | ENSG00000123240 | OPTN | GBM,KICH,LGG,READ,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000126950 | TMEM35A | GBM,LGG,READ,STAD,TGCT,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000131018 | SYNE1 | GBM,KICH,KIRP,LGG,PAAD,SKCM,STAD,UVM | | GBM | | ENSG00000279453 | ENSG00000131089 | ARHGEF9 | GBM,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000131370 | SH3BP5 | GBM,KICH,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000132535 | DLG4 | GBM,LGG,PAAD,STAD,TGCT,THCA,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000133687 | TMTC1 | GBM,KIRC,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000133816 | MICAL2 | GBM,LGG,STAD,TGCT,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000135525 | MAP7 | GBM,KICH,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000135631 | RAB11FIP5 | GBM,KICH,LGG,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000138686 | BBS7 | GBM,PAAD,TGCT,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000138944 | SHISAL1 | GBM,LGG,PAAD,STAD,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000139182 | CLSTN3 | GBM,KICH,LGG,PAAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000140992 | PDPK1 | GBM,LGG,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000142875 | PRKACB | GBM,KIRP,LGG,STAD,TGCT,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000144369 | FAM171B | GBM,LGG,PAAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000144711 | IQSEC1 | GBM,HNSC,LGG,TGCT,UCEC | | GBM | | ENSG00000279453 | ENSG00000145335 | SNCA | GBM,LGG,PAAD,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000145632 | PLK2 | GBM,KICH,LGG,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000148660 | CAMK2G | GBM,KICH,LGG,STAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000149294 | NCAM1 | GBM,PAAD,STAD,TGCT,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000151778 | SERP2 | GBM,KICH,LGG,PAAD,THCA,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000153707 | PTPRD | GBM,KIRP,LGG,TGCT,THCA | | GBM | | ENSG00000279453 | ENSG00000155093 | PTPRN2 | GBM,KICH,LGG,PAAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000156050 | FAM161B | GBM,KICH,LGG,PAAD,TGCT,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000160408 | ST6GALNAC6 | GBM,KICH,LGG,STAD,TGCT,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000160445 | ZER1 | GBM,LGG,PAAD,STAD,TGCT,UCS | | GBM | | ENSG00000279453 | ENSG00000160469 | BRSK1 | GBM,KIRP,LGG,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000161682 | FAM171A2 | GBM,TGCT,THCA,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000162545 | CAMK2N1 | GBM,LGG,READ,TGCT,THCA | | GBM | | ENSG00000279453 | ENSG00000163637 | PRICKLE2 | GBM,LGG,PAAD,STAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000163644 | PPM1K | GBM,KICH,LAML,STAD,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000164038 | SLC9B2 | GBM,LGG,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000164402 | SEPTIN8 | GBM,KICH,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000165152 | PGAP4 | GBM,KIRP,LGG,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000165699 | TSC1 | GBM,LIHC,PAAD,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000165868 | HSPA12A | GBM,KICH,LGG,PAAD,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000165943 | MOAP1 | GBM,LGG,PAAD,STAD,TGCT,UVM | | GBM | | ENSG00000279453 | ENSG00000166402 | TUB | GBM,LGG,PAAD,TGCT,THYM,UCEC,UVM | | GBM,UCEC | | ENSG00000279453 | ENSG00000166448 | TMEM130 | GBM,LGG,TGCT,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000166848 | TERF2IP | GBM,KICH,KIRP,LGG,PAAD,READ,STAD,TGCT,UCEC,UCS,UVM | | GBM | | ENSG00000279453 | ENSG00000170004 | CHD3 | GBM,KICH,PAAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000170153 | RNF150 | GBM,PAAD,STAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000170500 | LONRF2 | GBM,LGG,PAAD,THCA,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000174684 | B4GAT1 | GBM,KICH,LGG,PAAD,TGCT,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000175662 | TOM1L2 | GBM,KICH,LGG,PAAD,TGCT,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000176903 | PNMA1 | GBM,KICH,LGG,PAAD,READ,STAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000178974 | FBXO34 | GBM,LGG,TGCT,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000179912 | R3HDM2 | GBM,KIRP,LAML,LIHC,OV,PAAD,SARC,UCEC,UCS,UVM | | GBM | | ENSG00000279453 | ENSG00000180901 | KCTD2 | GBM,KICH,KIRP,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000181754 | AMIGO1 | GBM,KICH,LGG,PAAD,STAD,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000182013 | PNMA8A | GBM,KICH,KIRP,LGG,PAAD,THYM | | GBM | | ENSG00000279453 | ENSG00000183826 | BTBD9 | GBM,LGG,PAAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000184277 | TM2D3 | GBM,LGG,PAAD,THCA,THYM,UCS,UVM | | GBM | | ENSG00000279453 | ENSG00000184613 | NELL2 | GBM,LGG,TGCT,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000186310 | NAP1L3 | GBM,LGG,PAAD,TGCT,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000188026 | RILPL1 | GBM,KIRP,STAD,TGCT,THYM | | GBM | | ENSG00000279453 | ENSG00000196177 | ACADSB | GBM,KICH,LGG,TGCT,THYM,UCEC,UVM | | GBM | | ENSG00000279453 | ENSG00000197043 | ANXA6 | GBM,PAAD,READ,STAD,UCEC | | UCEC | | ENSG00000279453 | ENSG00000197283 | SYNGAP1 | GBM,KICH,KIRP,LGG,PAAD,STAD,TGCT | | GBM | | ENSG00000279453 | ENSG00000198513 | ATL1 | GBM,KICH,LGG,PAAD,READ,STAD,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000198908 | BHLHB9 | GBM,KICH,LGG,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000198954 | KIFBP | GBM,KICH,PAAD,THYM,UVM | | GBM | | ENSG00000279453 | ENSG00000273820 | USP27X | GBM,KICH,LGG,READ,TGCT,THCA,THYM,UCEC | | GBM,UCEC | | ENSG00000279453 | ENSG00000275023 | MLLT6 | GBM,KICH,KIRP,THCA,UVM | | GBM | | ENSG00000279453 | ENSG00000276293 | PIP4K2B | GBM,KICH,KIRP,LGG,LIHC,PAAD,READ,TGCT,THCA,THYM | | GBM | | ENSG00000279453 | ENSG00000011465 | DCN | HNSC,KICH,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000086289 | EPDR1 | HNSC,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000004799 | PDK4 | KICH,PAAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000021762 | OSBPL5 | KICH,STAD,TGCT,THCA,UCEC | | UCEC | | ENSG00000279453 | ENSG00000037280 | FLT4 | KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000076706 | MCAM | KICH,PAAD,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000090006 | LTBP4 | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000104332 | SFRP1 | KICH,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000126458 | RRAS | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000143196 | DPT | KICH,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000145687 | SSBP2 | KICH,LAML,PAAD,STAD,TGCT,UCEC | | UCEC | | ENSG00000279453 | ENSG00000146966 | DENND2A | KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000153071 | DAB2 | KICH,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000153823 | PID1 | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000154133 | ROBO4 | KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000163453 | IGFBP7 | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000164442 | CITED2 | KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000165406 | MARCHF8 | KICH,PAAD,READ,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000166073 | GPR176 | KICH,THYM,UCEC,UCS,UVM | | UCEC | | ENSG00000279453 | ENSG00000166341 | DCHS1 | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000169129 | AFAP1L2 | KICH,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000170876 | TMEM43 | KICH,KIRP,READ,STAD,TGCT,THCA,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000173269 | MMRN2 | KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000173546 | CSPG4 | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000176533 | GNG7 | KICH,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000189058 | APOD | KICH,READ,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000204301 | NOTCH4 | KICH,PAAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000118971 | CCND2 | KIRP,THCA,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000153790 | C7orf31 | KIRP,LAML,READ,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000154240 | CEP112 | KIRP,PAAD,TGCT,THCA,UCEC | | UCEC | | ENSG00000279453 | ENSG00000171791 | BCL2 | KIRP,LAML,READ,STAD,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000180694 | TMEM64 | LAML,PAAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000108733 | PEX12 | PAAD,READ,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000119638 | NEK9 | PAAD,STAD,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000134072 | CAMK1 | PAAD,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000150630 | VEGFC | PAAD,READ,STAD,TGCT,THYM,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000116667 | C1orf21 | READ,STAD,TGCT,UCEC,UVM | | UCEC | | ENSG00000279453 | ENSG00000166444 | DENND2B | READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000279453 | ENSG00000166482 | MFAP4 | READ,STAD,TGCT,THYM,UCEC,UCS | | UCEC | | ENSG00000279453 | ENSG00000171105 | INSR | READ,TGCT,THCA,THYM,UCEC | | UCEC |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000279453 | ENST00000624649 | SLC25A39 | ENSG00000013306 | ENST00000537904 | -0.2136 | 117 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | SLC25A39 | ENSG00000013306 | ENST00000588767 | -1.9350 | 442 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | AP2S1 | ENSG00000042753 | ENST00000601498 | -0.1012 | 50 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | AP2S1 | ENSG00000042753 | ENST00000599990 | -0.1012 | 50 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | AP2S1 | ENSG00000042753 | ENST00000593442 | -0.1012 | 50 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | AP2S1 | ENSG00000042753 | ENST00000263270 | -0.1012 | 59 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | AP2S1 | ENSG00000042753 | ENST00000597020 | -0.1012 | 64 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | PYCR3 | ENSG00000104524 | ENST00000433751 | -0.1026 | 1005 | BLCA,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PYCR3 | ENSG00000104524 | ENST00000377579 | -0.1396 | 173 | BLCA,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | DDX39A | ENSG00000123136 | ENST00000242776 | -0.2011 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | BCL2L12 | ENSG00000126453 | ENST00000441864 | -0.1703 | 172 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | BCL2L12 | ENSG00000126453 | ENST00000246785 | -0.1159 | 1097 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | BCL2L12 | ENSG00000126453 | ENST00000594157 | -0.2316 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | BCL2L12 | ENSG00000126453 | ENST00000598979 | -0.1049 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | BCL2L12 | ENSG00000126453 | ENST00000246784 | -0.1159 | 1097 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | BCL2L12 | ENSG00000126453 | ENST00000598306 | -0.1159 | 1097 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | TOMM40 | ENSG00000130204 | ENST00000592041 | -0.1323 | 69 | CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | SURF2 | ENSG00000148291 | ENST00000371964 | -0.3022 | 1015 | UCEC | - | | ENSG00000279453 | ENST00000624649 | DTYMK | ENSG00000168393 | ENST00000445261 | -2.1650 | 1237 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | TIMM10 | ENSG00000134809 | ENST00000525587 | -0.1178 | 1386 | UCEC | - | | ENSG00000279453 | ENST00000624649 | TIMM10 | ENSG00000134809 | ENST00000525158 | -0.1605 | 1386 | UCEC | - | | ENSG00000279453 | ENST00000624649 | TMEM258 | ENSG00000134825 | ENST00000257262 | -0.1099 | 907 | GBM | - | | ENSG00000279453 | ENST00000624649 | TMEM258 | ENSG00000134825 | ENST00000543510 | -0.1402 | 196 | GBM | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000571570 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000572639 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000579978 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000577425 | -0.1053 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000571024 | -0.1065 | 92 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000574924 | -0.1065 | 92 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000572851 | -0.1065 | 92 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000357385 | -0.1110 | 126 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000571874 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000578550 | -0.1737 | 304 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000577747 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000579133 | -0.1053 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000392376 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000583839 | -0.1572 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANAPC11 | ENSG00000141552 | ENST00000578544 | -0.1121 | 727 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | RPS8 | ENSG00000142937 | ENST00000396651 | -0.1152 | 976 | GBM | PCPG | | ENSG00000279453 | ENST00000624649 | RPS8 | ENSG00000142937 | ENST00000372209 | -0.1152 | 976 | GBM | PCPG | | ENSG00000279453 | ENST00000624649 | RPL10 | ENSG00000147403 | ENST00000369817 | -0.1677 | 944 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPL10 | ENSG00000147403 | ENST00000406022 | -0.3300 | 952 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPL10 | ENSG00000147403 | ENST00000427682 | -0.2255 | 1003 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPL10 | ENSG00000147403 | ENST00000428169 | -0.4217 | 1339 | GBM | - | | ENSG00000279453 | ENST00000624649 | TRPT1 | ENSG00000149743 | ENST00000394547 | -0.1116 | 1601 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TRPT1 | ENSG00000149743 | ENST00000541278 | -0.1658 | 1590 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TRPT1 | ENSG00000149743 | ENST00000394546 | -0.1658 | 1590 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TRPT1 | ENSG00000149743 | ENST00000546133 | -0.1658 | 1590 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TRPT1 | ENSG00000149743 | ENST00000317459 | -0.5409 | 1597 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TRPT1 | ENSG00000149743 | ENST00000546089 | -0.5409 | 1605 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000338110 | -0.1372 | 1005 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000627059 | -0.1221 | 999 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000395248 | -0.1221 | 999 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000566897 | -0.1372 | 1006 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000569545 | -0.3067 | 367 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000563060 | -0.1372 | 1005 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000412304 | -0.1246 | 1008 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000564546 | -0.1221 | 999 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000564595 | -0.1246 | 1008 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000569798 | -0.1023 | 931 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000395240 | -0.1388 | 1039 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ALDOA | ENSG00000149925 | ENST00000565355 | -0.1468 | 115 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | KRTCAP3 | ENSG00000157992 | ENST00000288873 | -0.1256 | 115 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | KRTCAP3 | ENSG00000157992 | ENST00000407293 | -0.1256 | 115 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | WDR4 | ENSG00000160193 | ENST00000330317 | -0.1249 | 900 | BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | FLAD1 | ENSG00000160688 | ENST00000315144 | -0.1774 | 993 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | FLAD1 | ENSG00000160688 | ENST00000368431 | -0.1594 | 115 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | FLAD1 | ENSG00000160688 | ENST00000292180 | -0.1774 | 993 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | FLAD1 | ENSG00000160688 | ENST00000295530 | -0.1045 | 692 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | FLAD1 | ENSG00000160688 | ENST00000368428 | -0.1774 | 993 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | YDJC | ENSG00000161179 | ENST00000292778 | -0.1099 | 1 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | POLR2H | ENSG00000163882 | ENST00000456318 | -0.1001 | 888 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | POLR2H | ENSG00000163882 | ENST00000438240 | -0.1057 | 932 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | POLR2H | ENSG00000163882 | ENST00000430783 | -0.1001 | 889 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | POLR2H | ENSG00000163882 | ENST00000443489 | -0.1001 | 891 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | POLR2H | ENSG00000163882 | ENST00000452961 | -0.1111 | 949 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ENTR1 | ENSG00000165689 | ENST00000417512 | -0.1607 | 1168 | GBM | - | | ENSG00000279453 | ENST00000624649 | ENDOG | ENSG00000167136 | ENST00000372642 | -0.2827 | 180 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | POP5 | ENSG00000167272 | ENST00000357500 | -0.1306 | 1 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | GPX4 | ENSG00000167468 | ENST00000616066 | -0.1315 | 30 | UCEC | - | | ENSG00000279453 | ENST00000624649 | GPX4 | ENSG00000167468 | ENST00000354171 | -0.1315 | 27 | UCEC | - | | ENSG00000279453 | ENST00000624649 | GPX4 | ENSG00000167468 | ENST00000611653 | -0.1315 | 38 | UCEC | - | | ENSG00000279453 | ENST00000624649 | GPX4 | ENSG00000167468 | ENST00000588919 | -0.8491 | 27 | UCEC | - | | ENSG00000279453 | ENST00000624649 | GPX4 | ENSG00000167468 | ENST00000622390 | -0.1315 | 30 | UCEC | - | | ENSG00000279453 | ENST00000624649 | TUBA1C | ENSG00000167553 | ENST00000541364 | -0.1407 | 21 | GBM,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | TUBA1C | ENSG00000167553 | ENST00000549818 | -0.1114 | 811 | GBM,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | TUBA1C | ENSG00000167553 | ENST00000552125 | -0.1174 | 1 | GBM,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | YIF1B | ENSG00000167645 | ENST00000592694 | -0.1124 | 795 | BLCA,UCEC | - | | ENSG00000279453 | ENST00000624649 | YIF1B | ENSG00000167645 | ENST00000337679 | -0.1024 | 1 | BLCA,UCEC | - | | ENSG00000279453 | ENST00000624649 | YIF1B | ENSG00000167645 | ENST00000392124 | -0.1124 | 806 | BLCA,UCEC | - | | ENSG00000279453 | ENST00000624649 | YIF1B | ENSG00000167645 | ENST00000339413 | -0.1124 | 809 | BLCA,UCEC | - | | ENSG00000279453 | ENST00000624649 | YIF1B | ENSG00000167645 | ENST00000592246 | -0.1796 | 795 | BLCA,UCEC | - | | ENSG00000279453 | ENST00000624649 | YIF1B | ENSG00000167645 | ENST00000591755 | -0.1077 | 1008 | BLCA,UCEC | - | | ENSG00000279453 | ENST00000624649 | TK1 | ENSG00000167900 | ENST00000590862 | -0.2248 | 219 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ECI1 | ENSG00000167969 | ENST00000562238 | -0.1999 | 166 | GBM | - | | ENSG00000279453 | ENST00000624649 | ECI1 | ENSG00000167969 | ENST00000570258 | -0.5552 | 874 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPSA | ENSG00000168028 | ENST00000443003 | -0.1949 | 390 | GBM | - | | ENSG00000279453 | ENST00000624649 | FEN1 | ENSG00000168496 | ENST00000535307 | -0.1113 | 1 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | LMAN2 | ENSG00000169223 | ENST00000515209 | -0.1506 | 68 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | LRRC45 | ENSG00000169683 | ENST00000581227 | -0.1060 | 1 | BLCA,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | CENPX | ENSG00000169689 | ENST00000579520 | -0.1037 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | CENPX | ENSG00000169689 | ENST00000584600 | -0.1037 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | CENPX | ENSG00000169689 | ENST00000580435 | -0.1037 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | CENPX | ENSG00000169689 | ENST00000306704 | -0.1037 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | CENPX | ENSG00000169689 | ENST00000392359 | -0.1037 | 1 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | DUS1L | ENSG00000169718 | ENST00000354321 | -0.1232 | 89 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | DUS1L | ENSG00000169718 | ENST00000577574 | -0.5775 | 519 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | DUS1L | ENSG00000169718 | ENST00000542088 | -0.1060 | 729 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | DUS1L | ENSG00000169718 | ENST00000580731 | -0.3758 | 1 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | THOP1 | ENSG00000172009 | ENST00000586677 | -0.2374 | 992 | CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | THOP1 | ENSG00000172009 | ENST00000591363 | -0.4094 | 587 | CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | THOP1 | ENSG00000172009 | ENST00000590970 | -0.1045 | 907 | CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | THOP1 | ENSG00000172009 | ENST00000587401 | -0.1891 | 188 | CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | THOP1 | ENSG00000172009 | ENST00000395212 | -0.1045 | 908 | CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | PPP1CA | ENSG00000172531 | ENST00000358239 | -0.1334 | 731 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | GMPPB | ENSG00000173540 | ENST00000308388 | -0.1085 | 1 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | MRPL11 | ENSG00000174547 | ENST00000528272 | -0.1040 | 1 | GBM,LUSC | - | | ENSG00000279453 | ENST00000624649 | RPS6KB2 | ENSG00000175634 | ENST00000312629 | -0.1515 | 75 | UCEC | - | | ENSG00000279453 | ENST00000624649 | AURKAIP1 | ENSG00000175756 | ENST00000338370 | -0.3796 | 1127 | UCEC | - | | ENSG00000279453 | ENST00000624649 | AURKAIP1 | ENSG00000175756 | ENST00000338338 | -0.3796 | 1127 | UCEC | - | | ENSG00000279453 | ENST00000624649 | AURKAIP1 | ENSG00000175756 | ENST00000321751 | -0.3796 | 1129 | UCEC | - | | ENSG00000279453 | ENST00000624649 | AURKAIP1 | ENSG00000175756 | ENST00000378853 | -0.3796 | 1134 | UCEC | - | | ENSG00000279453 | ENST00000624649 | COX5A | ENSG00000178741 | ENST00000562233 | -0.1126 | 316 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | COX5A | ENSG00000178741 | ENST00000568783 | -0.2593 | 401 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | COX5A | ENSG00000178741 | ENST00000564811 | -0.2415 | 67 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | RCC1 | ENSG00000180198 | ENST00000373831 | -0.1434 | 92 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | SSR4 | ENSG00000180879 | ENST00000320857 | -0.2407 | 895 | GBM,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | SSR4 | ENSG00000180879 | ENST00000370087 | -0.2407 | 895 | GBM,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | SSR4 | ENSG00000180879 | ENST00000370086 | -0.2242 | 888 | GBM,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | SSR4 | ENSG00000180879 | ENST00000370085 | -0.2407 | 895 | GBM,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | COA4 | ENSG00000181924 | ENST00000537289 | -0.1164 | 91 | GBM | PCPG | | ENSG00000279453 | ENST00000624649 | NR2C2AP | ENSG00000184162 | ENST00000420605 | -0.6850 | 262 | GBM,CESC,LUSC | - | | ENSG00000279453 | ENST00000624649 | NR2C2AP | ENSG00000184162 | ENST00000538165 | -0.1050 | 989 | GBM,CESC,LUSC | - | | ENSG00000279453 | ENST00000624649 | PKP3 | ENSG00000184363 | ENST00000331563 | -0.1296 | 83 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PKP3 | ENSG00000184363 | ENST00000525642 | -0.1867 | 201 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | TEDC1 | ENSG00000185347 | ENST00000392527 | -0.1356 | 123 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | TEDC1 | ENSG00000185347 | ENST00000392522 | -0.1304 | 141 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | TEDC1 | ENSG00000185347 | ENST00000392523 | -0.1356 | 127 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | TEDC1 | ENSG00000185347 | ENST00000354560 | -0.1090 | 100 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | TEDC1 | ENSG00000185347 | ENST00000334656 | -0.1194 | 81 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | NDUFA13 | ENSG00000186010 | ENST00000507754 | -0.2533 | 1247 | UCEC | - | | ENSG00000279453 | ENST00000624649 | NDUFA13 | ENSG00000186010 | ENST00000428459 | -0.2108 | 794 | UCEC | - | | ENSG00000279453 | ENST00000624649 | JPT1 | ENSG00000189159 | ENST00000481647 | -0.1192 | 1281 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | JPT1 | ENSG00000189159 | ENST00000581874 | -0.3266 | 1 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | APRT | ENSG00000198931 | ENST00000378364 | -0.1522 | 1 | GBM | - | | ENSG00000279453 | ENST00000624649 | APRT | ENSG00000198931 | ENST00000426324 | -0.1343 | 21 | GBM | - | | ENSG00000279453 | ENST00000624649 | APRT | ENSG00000198931 | ENST00000568319 | -0.1138 | 21 | GBM | - | | ENSG00000279453 | ENST00000624649 | APRT | ENSG00000198931 | ENST00000563655 | -0.1423 | 24 | GBM | - | | ENSG00000279453 | ENST00000624649 | APRT | ENSG00000198931 | ENST00000569616 | -0.1174 | 70 | GBM | - | | ENSG00000279453 | ENST00000624649 | ALG3 | ENSG00000214160 | ENST00000397676 | -0.1182 | 278 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | ALG3 | ENSG00000214160 | ENST00000445626 | -0.2107 | 818 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | ALG3 | ENSG00000214160 | ENST00000455059 | -0.3628 | 818 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | FAM174C | ENSG00000228300 | ENST00000469144 | -0.4967 | 227 | UCEC | - | | ENSG00000279453 | ENST00000624649 | GPX1 | ENSG00000233276 | ENST00000419783 | -0.1269 | 31 | GBM,UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | GPX1 | ENSG00000233276 | ENST00000620890 | -0.1269 | 35 | GBM,UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | REEP4 | ENSG00000168476 | ENST00000306306 | -0.1097 | 1 | GBM,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | REEP4 | ENSG00000168476 | ENST00000523293 | -0.1827 | 170 | GBM,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | POLR2J | ENSG00000005075 | ENST00000393794 | -0.1127 | 1078 | GBM | - | | ENSG00000279453 | ENST00000624649 | SEC61A1 | ENSG00000058262 | ENST00000464451 | -0.1055 | 2040 | GBM | - | | ENSG00000279453 | ENST00000624649 | GSTP1 | ENSG00000084207 | ENST00000398606 | -0.4454 | 1010 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | GSTP1 | ENSG00000084207 | ENST00000398603 | -0.4454 | 1020 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | GSTP1 | ENSG00000084207 | ENST00000495996 | -0.7575 | 1139 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | PSME2 | ENSG00000100911 | ENST00000560410 | -0.1990 | 203 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | PSME2 | ENSG00000100911 | ENST00000216802 | -0.1990 | 203 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | PSME2 | ENSG00000100911 | ENST00000615264 | -0.1990 | 203 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | ETHE1 | ENSG00000105755 | ENST00000292147 | -0.1138 | 1075 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ETHE1 | ENSG00000105755 | ENST00000600651 | -0.1464 | 5 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | DCPS | ENSG00000110063 | ENST00000263579 | -0.1403 | 60 | GBM | - | | ENSG00000279453 | ENST00000624649 | MAD2L2 | ENSG00000116670 | ENST00000235310 | -0.1042 | 78 | GBM,BLCA,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | MAD2L2 | ENSG00000116670 | ENST00000376692 | -0.1009 | 79 | GBM,BLCA,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | MAD2L2 | ENSG00000116670 | ENST00000376672 | -0.1130 | 41 | GBM,BLCA,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | MAD2L2 | ENSG00000116670 | ENST00000376669 | -0.2103 | 986 | GBM,BLCA,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | MAD2L2 | ENSG00000116670 | ENST00000376667 | -0.5184 | 150 | GBM,BLCA,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | ORMDL2 | ENSG00000123353 | ENST00000550836 | -0.1824 | 153 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | ORMDL2 | ENSG00000123353 | ENST00000548974 | -0.1136 | 4 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | SNRPB | ENSG00000125835 | ENST00000438552 | -0.1315 | 117 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | SNRPB | ENSG00000125835 | ENST00000339610 | -0.1315 | 126 | GBM,BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | SDF2L1 | ENSG00000128228 | ENST00000248958 | -0.1767 | 58 | GBM,BLCA,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | CCNB1 | ENSG00000134057 | ENST00000507798 | -0.1106 | 231 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | CHEK1 | ENSG00000149554 | ENST00000524737 | -0.1419 | 942 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | CCNB2 | ENSG00000157456 | ENST00000559622 | -0.1025 | 108 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | SCAND1 | ENSG00000171222 | ENST00000305978 | -0.1899 | 754 | UCEC | - | | ENSG00000279453 | ENST00000624649 | SCAND1 | ENSG00000171222 | ENST00000615116 | -0.1899 | 754 | UCEC | - | | ENSG00000279453 | ENST00000624649 | SCAND1 | ENSG00000171222 | ENST00000373991 | -0.1899 | 766 | UCEC | - | | ENSG00000279453 | ENST00000624649 | OXLD1 | ENSG00000204237 | ENST00000374741 | -0.1673 | 1 | BLCA,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | NME1 | ENSG00000239672 | ENST00000475573 | -0.1532 | 1 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | NME1 | ENSG00000239672 | ENST00000013034 | -0.2009 | 1408 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ATP5MF | ENSG00000241468 | ENST00000394186 | -0.3357 | 29 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | ANTKMT | ENSG00000103254 | ENST00000569529 | -0.2721 | 1003 | UCEC | - | | ENSG00000279453 | ENST00000624649 | ANTKMT | ENSG00000103254 | ENST00000219535 | -0.4090 | 2204 | UCEC | - | | ENSG00000279453 | ENST00000624649 | ANTKMT | ENSG00000103254 | ENST00000568916 | -0.2721 | 1003 | UCEC | - | | ENSG00000279453 | ENST00000624649 | TEDC2 | ENSG00000162062 | ENST00000361837 | -0.1021 | 151 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | TEDC2 | ENSG00000162062 | ENST00000483320 | -0.1021 | 151 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | CLPP | ENSG00000125656 | ENST00000245816 | -0.1199 | 116 | GBM | - | | ENSG00000279453 | ENST00000624649 | CLPP | ENSG00000125656 | ENST00000596149 | -0.3370 | 15 | GBM | - | | ENSG00000279453 | ENST00000624649 | NCLN | ENSG00000125912 | ENST00000590671 | -0.1499 | 71 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | NCLN | ENSG00000125912 | ENST00000587740 | -0.1418 | 1 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | NCLN | ENSG00000125912 | ENST00000592235 | -0.1499 | 77 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | TBRG4 | ENSG00000136270 | ENST00000494076 | -0.1046 | 134 | UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PSMG3 | ENSG00000157778 | ENST00000252329 | -0.2531 | 95 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PPP1R14B | ENSG00000173457 | ENST00000392210 | -0.1073 | 11 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PPP1R14B | ENSG00000173457 | ENST00000309318 | -0.1073 | 15 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PPP1R14B | ENSG00000173457 | ENST00000542235 | -0.1229 | 80 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | RUVBL1 | ENSG00000175792 | ENST00000478892 | -0.1600 | 978 | GBM,BLCA,CESC,LUSC | - | | ENSG00000279453 | ENST00000624649 | RUVBL2 | ENSG00000183207 | ENST00000595090 | -0.1086 | 254 | UCEC | - | | ENSG00000279453 | ENST00000624649 | RUVBL2 | ENSG00000183207 | ENST00000601968 | -0.1858 | 292 | UCEC | - | | ENSG00000279453 | ENST00000624649 | LONP1 | ENSG00000196365 | ENST00000360614 | -0.1119 | 744 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | LONP1 | ENSG00000196365 | ENST00000585374 | -0.1119 | 745 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | LONP1 | ENSG00000196365 | ENST00000593119 | -0.1119 | 745 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | LONP1 | ENSG00000196365 | ENST00000540670 | -0.1119 | 745 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | LONP1 | ENSG00000196365 | ENST00000590729 | -0.2021 | 164 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | LONP1 | ENSG00000196365 | ENST00000586617 | -0.1019 | 1139 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | BOP1 | ENSG00000261236 | ENST00000569669 | -0.2771 | 955 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | ECE2 | ENSG00000145194 | ENST00000357474 | -0.1827 | 158 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | YIF1A | ENSG00000174851 | ENST00000496746 | -1.0671 | 765 | GBM | - | | ENSG00000279453 | ENST00000624649 | YIF1A | ENSG00000174851 | ENST00000471387 | -0.1270 | 951 | GBM | - | | ENSG00000279453 | ENST00000624649 | YIF1A | ENSG00000174851 | ENST00000359461 | -1.0671 | 765 | GBM | - | | ENSG00000279453 | ENST00000624649 | YIF1A | ENSG00000174851 | ENST00000376901 | -1.0671 | 765 | GBM | - | | ENSG00000279453 | ENST00000624649 | YIF1A | ENSG00000174851 | ENST00000484814 | -0.1481 | 925 | GBM | - | | ENSG00000279453 | ENST00000624649 | DPP3 | ENSG00000254986 | ENST00000531863 | -0.1029 | 1 | BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | DPP3 | ENSG00000254986 | ENST00000532677 | -0.1029 | 1 | BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | DPP3 | ENSG00000254986 | ENST00000541961 | -0.1029 | 1 | BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | DPP3 | ENSG00000254986 | ENST00000530165 | -0.1700 | 189 | BLCA,CESC,UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | UQCRC1 | ENSG00000010256 | ENST00000203407 | -0.1621 | 78 | UCEC | - | | ENSG00000279453 | ENST00000624649 | NOP16 | ENSG00000048162 | ENST00000619979 | -2.7100 | 818 | UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | NOP16 | ENSG00000048162 | ENST00000621444 | -0.1811 | 2048 | UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | NOP16 | ENSG00000048162 | ENST00000614830 | -0.1106 | 1 | UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | NOP16 | ENSG00000048162 | ENST00000613081 | -0.1811 | 2049 | UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | NOP16 | ENSG00000048162 | ENST00000502663 | -0.1128 | 1 | UCEC,LUSC | - | | ENSG00000279453 | ENST00000624649 | WDR18 | ENSG00000065268 | ENST00000585809 | -0.1323 | 1 | UCEC | - | | ENSG00000279453 | ENST00000624649 | WDR18 | ENSG00000065268 | ENST00000587001 | -0.1323 | 9 | UCEC | - | | ENSG00000279453 | ENST00000624649 | WDR18 | ENSG00000065268 | ENST00000251289 | -0.1323 | 10 | UCEC | - | | ENSG00000279453 | ENST00000624649 | WDR18 | ENSG00000065268 | ENST00000607440 | -0.1323 | 9 | UCEC | - | | ENSG00000279453 | ENST00000624649 | WDR18 | ENSG00000065268 | ENST00000591997 | -0.1211 | 93 | UCEC | - | | ENSG00000279453 | ENST00000624649 | STXBP2 | ENSG00000076944 | ENST00000622853 | -0.1427 | 1822 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | STXBP2 | ENSG00000076944 | ENST00000441779 | -0.2175 | 189 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | STXBP2 | ENSG00000076944 | ENST00000221283 | -0.2175 | 189 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | STXBP2 | ENSG00000076944 | ENST00000597068 | -0.1515 | 1 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | STXBP2 | ENSG00000076944 | ENST00000414284 | -0.2175 | 189 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | STXBP2 | ENSG00000076944 | ENST00000600702 | -0.1127 | 815 | GBM,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | PAFAH1B3 | ENSG00000079462 | ENST00000262890 | -0.2472 | 205 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PAFAH1B3 | ENSG00000079462 | ENST00000538771 | -0.2472 | 205 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PAFAH1B3 | ENSG00000079462 | ENST00000594989 | -0.2472 | 208 | BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | BIRC5 | ENSG00000089685 | ENST00000592734 | -0.1125 | 269 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | BIRC5 | ENSG00000089685 | ENST00000587746 | -0.1153 | 899 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | CENPM | ENSG00000100162 | ENST00000402420 | -1.1046 | 1220 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | TSPO | ENSG00000100300 | ENST00000337554 | -0.1050 | 1088 | GBM | - | | ENSG00000279453 | ENST00000624649 | TSPO | ENSG00000100300 | ENST00000583777 | -0.1529 | 101 | GBM | - | | ENSG00000279453 | ENST00000624649 | TSPO | ENSG00000100300 | ENST00000329563 | -0.1529 | 132 | GBM | - | | ENSG00000279453 | ENST00000624649 | TSPO | ENSG00000100300 | ENST00000396265 | -0.1529 | 101 | GBM | - | | ENSG00000279453 | ENST00000624649 | NAA10 | ENSG00000102030 | ENST00000370015 | -0.1173 | 1 | UCEC | - | | ENSG00000279453 | ENST00000624649 | NAA10 | ENSG00000102030 | ENST00000393712 | -0.1384 | 1 | UCEC | - | | ENSG00000279453 | ENST00000624649 | NAA10 | ENSG00000102030 | ENST00000370009 | -0.1868 | 135 | UCEC | - | | ENSG00000279453 | ENST00000624649 | STX10 | ENSG00000104915 | ENST00000343587 | -0.1020 | 763 | GBM | - | | ENSG00000279453 | ENST00000624649 | STX10 | ENSG00000104915 | ENST00000587318 | -0.8186 | 223 | GBM | - | | ENSG00000279453 | ENST00000624649 | STX10 | ENSG00000104915 | ENST00000587230 | -0.1061 | 794 | GBM | - | | ENSG00000279453 | ENST00000624649 | STX10 | ENSG00000104915 | ENST00000593126 | -0.1951 | 18 | GBM | - | | ENSG00000279453 | ENST00000624649 | ASF1B | ENSG00000105011 | ENST00000474890 | -0.1050 | 888 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | TMEM147 | ENSG00000105677 | ENST00000222284 | -0.1261 | 1605 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TMEM147 | ENSG00000105677 | ENST00000392204 | -0.1261 | 1605 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | TMEM147 | ENSG00000105677 | ENST00000392205 | -0.1430 | 879 | GBM,UCEC | - | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000602122 | -0.2856 | 806 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000621372 | -0.2856 | 808 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000361790 | -0.2856 | 808 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000360798 | -0.2856 | 806 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000354900 | -0.2856 | 804 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000347609 | -0.1034 | 1118 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000427250 | -0.2856 | 806 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | LSR | ENSG00000105699 | ENST00000605618 | -0.1331 | 1397 | BLCA,CESC,UCEC,LUSC,LUAD | PCPG | | ENSG00000279453 | ENST00000624649 | AP1S1 | ENSG00000106367 | ENST00000443943 | -0.1085 | 7 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | POLD2 | ENSG00000106628 | ENST00000610533 | -0.1492 | 1159 | GBM,LUSC | - | | ENSG00000279453 | ENST00000624649 | POLD2 | ENSG00000106628 | ENST00000406581 | -0.1492 | 1165 | GBM,LUSC | - | | ENSG00000279453 | ENST00000624649 | POLD2 | ENSG00000106628 | ENST00000223361 | -0.1756 | 1128 | GBM,LUSC | - | | ENSG00000279453 | ENST00000624649 | UBE2S | ENSG00000108106 | ENST00000589978 | -0.1752 | 187 | GBM,BLCA,CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | NDUFS8 | ENSG00000110717 | ENST00000313468 | -0.3167 | 781 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | NDUFS8 | ENSG00000110717 | ENST00000528492 | -0.3167 | 786 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | NDUFS8 | ENSG00000110717 | ENST00000531228 | -0.1960 | 1 | UCEC | PCPG | | ENSG00000279453 | ENST00000624649 | GAPDH | ENSG00000111640 | ENST00000396856 | -0.1058 | 773 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | GAPDH | ENSG00000111640 | ENST00000396858 | -0.1101 | 1229 | CESC,UCEC,LUSC,LUAD | - | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000251654 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000490504 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000483687 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000468777 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000462637 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000466072 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | PCCB | ENSG00000114054 | ENST00000482086 | -0.1216 | 943 | CESC,UCEC,LUSC | PCPG | | ENSG00000279453 | ENST00000624649 | MAPKAPK3 | ENSG00000114738 | ENST00000451680 | -0.1004 | 1 | GBM | - | | ENSG00000279453 | ENST00000624649 | LAMTOR2 | ENSG00000116586 | ENST00000368305 | -0.1095 | 8 | UCEC | - | | ENSG00000279453 | ENST00000624649 | LAMTOR2 | ENSG00000116586 | ENST00000368304 | -0.1095 | 12 | UCEC | - | | ENSG00000279453 | ENST00000624649 | LAMTOR2 | ENSG00000116586 | ENST00000368302 | -0.2465 | 159 | UCEC | - | | ENSG00000279453 | ENST00000624649 | GALE | ENSG00000117308 | ENST00000617979 | -0.1004 | 736 | CESC,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | GALE | ENSG00000117308 | ENST00000374497 | -0.1004 | 737 | CESC,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | GALE | ENSG00000117308 | ENST00000456977 | -0.1004 | 741 | CESC,UCEC,LUAD | - | | ENSG00000279453 | ENST00000624649 | CLN6 | ENSG00000128973 | ENST00000566347 | -0.2003 | 936 | BLCA,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | CLN6 | ENSG00000128973 | ENST00000538696 | -0.2594 | 963 | BLCA,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | CLN6 | ENSG00000128973 | ENST00000564752 | -0.1202 | 782 | BLCA,CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | RPL36 | ENSG00000130255 | ENST00000579649 | -0.1695 | 1 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPL36 | ENSG00000130255 | ENST00000577222 | -2.4900 | 1277 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPL36 | ENSG00000130255 | ENST00000579446 | -0.1704 | 933 | GBM | - | | ENSG00000279453 | ENST00000624649 | RPL36 | ENSG00000130255 | ENST00000394580 | -0.1695 | 1 | GBM | - | | ENSG00000279453 | ENST00000624649 | CRB3 | ENSG00000130545 | ENST00000356762 | -0.1256 | 75 | CESC,UCEC | - | | ENSG00000279453 | ENST00000624649 | CRB3 | ENSG00000130545 | ENST00000308243 | -0.1720 | 977 | CESC,UCEC | - |
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000279453 | RP3-425C14.4,hsa-mir-25,CALHM2 | UCEC,TGCT,PRAD,UVM,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-25,ITM2B | UCEC,TGCT,PRAD,UVM,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-25,FBXO32 | UCEC,TGCT,PRAD,UVM,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-942,GPRASP1 | UCEC,KICH,TGCT,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-942,FAXDC2 | UCEC,KICH,TGCT,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-942,MSRB3 | UCEC,KICH,TGCT,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-942,ZBTB4 | UCEC,KICH,TGCT,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-942,SLC22A17 | UCEC,KICH,TGCT,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-106b,SERINC1 | UCEC,TGCT,PRAD,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-106b,MSRB3 | UCEC,TGCT,PRAD,MESO,THYM | | ENSG00000279453 | RP3-425C14.4,hsa-mir-106b,MBNL2 | UCEC,TGCT,PRAD,MESO,THYM |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000279453.1 | ENST00000624649.1 | 127 | 32 | 415 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.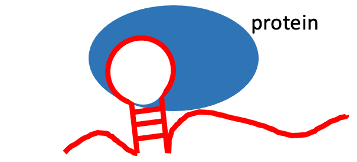
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).