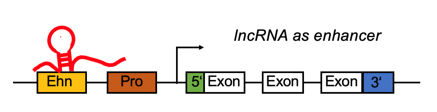
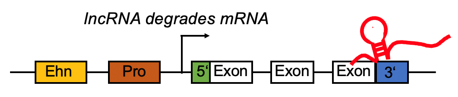
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | ARHGEF17,RPAP2,PMM2,VAMP1,MFSD8,GBF1,DNAJC27,MZF1,TTC31,MAPK8IP3,VAMP4,SLF2,TMEM184C,PIGG,SENP1,ST7L,MSANTD2,CHKA,FAM133B,IBTK,TTC14,CCNK,PPIP5K2,SCML1,C20orf194,NCBP1,FAM219B,UBE3A,RIC8B,THOC2,ZBTB6,OTULIN,OFD1,ATP9A,PEAK1,PURA,XPNPEP3,HGS,ANO8,NEMP1,EML4,MIS18BP1,MDN1,SECISBP2L,ORC2,NFKBIZ,PHC3,ZZEF1,DHX35,INTS2,SUPT20H,DCUN1D2,SMARCAD1,GIGYF1,KHNYN,CDK17,IPO9,ATG2B,HUWE1,AP4E1,TNPO2,DDX42,WDR35,EIF4A1,SEC24C,CRYBG3,PGGT1B,SELENOI,SWT1,BRD3,AMMECR1L,TMEM168,ARHGEF2,FANCL,TTC17,KLF7,PHKA2,UBE2G2,PAWR,MIB1,CEP68,NEMP2,VEZF1,MLLT6,GSK3B,ZNF70,INO80D,ICE2,AGK,SMCR8,CRLF3,PHF7,KPNA1,AP1G2,ZXDA,ZNF25,SCO1,TRNT1,CHD8,EPN2,DENND6A,RALGPS2,UBR4,TMED8,NBEAL2,RELL1,OSBPL8,CBLB,XPO4,PKD1,RIMKLB,METAP1D,SLC25A46,NSD1,UTP14C,CEP83,LIN54,LRIG2,PARP2,IKBKB,SART3,DGKE,PAXBP1,CCDC125,TMOD2,PLEKHA8,RSRP1,EYA3,RNF31,GOLGA4,FLCN,NVL,ATG14,GOLGB1,PWWP2A,ANKLE2,SLC39A10,LINS1,SCAF11,PUS10,NDUFAF7,TOR1AIP2,ZNF862,SLC23A2,PDS5B,SBF1,EDEM3,ATP6V0A2,KIF3A,GGT7,EXOC7,KHDC4,MIDEAS,PIK3CA,LMLN,ATAD2B,NISCH,ALG11,PREPL,PI4KA,ESCO1,MTMR10,PBX2,KLF12,ANGEL2,NAA15,MYSM1,TJAP1,TAF1D,CEP95,NUP153,TTI1,IFT88,TASP1,SUGT1,GTF3C3,EFCAB7,PRR14L,FAM160A1,BARD1,RPS6KB1,PTK2,GABBR1,N6AMT1,KANSL3,DENND4A,MPHOSPH9,HECTD1,EDC4,AGO2,RBM26,PURB,SPATA6,SYNE1,MLH3,KANSL1L,KIF21A,WDFY3,ATP8A1,ENTPD4,SMC5,HEATR5B,VPS53,LCOR,FBXL20,CENPC,NOL9,HERC2,OTUD6B,ABHD13,TUBGCP4,C2orf49,POLK,PLEKHA3,PRKAA1,MICALL2,HSPBAP1,WRN,PRDM2,RBL1,SPATA5,USP12,ZGPAT,AAK1,TIA1,MOSMO,ZNF8,BMPR2,SNRNP200,ERCC3,POLG,FAM13B,GRIPAP1,ARFGEF1,MTMR9,LUC7L,SPOPL,RAPGEF5,PTBP3,WDR6,BRD8,ERCC8,TMCC1,UBE2V1,USP16,MAPKAPK5,BROX,CYTH2,APPL1,RBM33,PAG1,DSTYK,SLC25A27,PLEKHA7,PDZD8,EXOC5,CLEC16A,ERICH1,GXYLT1,MAP3K2,GPATCH2,LAMA5,NHLRC2,RUFY3,TTC28,ZFYVE26,KMT2E,DCAF16,ITPR2,MORC3,PEX1,CHORDC1,S100PBP,TRIM3,KLHL18,KAT6B,FBXW8,INPP4B,TRIM23,TSC2,PCNT,N4BP1,YEATS2,TAF1C,RBM14,IKZF4,PIP4K2B,NF1,ANKRD50,THAP9,THAP6,DHX15,SENP7,AQR,CHFR,MTRF1,TAF2,SEC16A,TRIP11,TRPM7,PXK,TRAPPC8,ERV3-1,MKLN1,PHIP,DOCK9,KLC1,CLMN,CCDC93,LMBR1,SIPA1L3,UBAP2,HP1BP3,ACP6,VPS13C,DIDO1,FKTN,ALG9,PUM2,DHX36,TTLL3,TUBE1,C2CD2,LRP5L,KATNIP,SPG11,FBXO42,PIP5K1A,MAPK1,FRMD4B,ELMO2,PUM1,USP22,RB1CC1,ZRANB1,DCP1A,DMTF1,MKNK1,ZDHHC23,FAM193A,NSD3,LMBR1L,STAT2,VMP1,WWC2,KAT6A,AFF4,ARAP2,RNF38,ZBED5,HUS1,INPP5B,SETD2,SIN3A,MEX3C,CREB1,SLC2A11,SUDS3,DPY19L4,CAPN7,SMAD2,CLCN6,IGIP,PDE7A,CLK2,MYEF2,RNF19A,ASH1L,POLG2,PTAR1,MGRN1,KATNAL1,CPSF7,PARP4,CTC1,GABPB1,VPS36,INTS6,FBXO38,IFT172,HCFC2,RSRC2,SNX27,NAPB,USP46,NAA25,PRMT9,CHD9,RALGAPA2,TOPBP1,BACH1,SBNO1,ABHD16A,NXF1,RNF111,PAN2,STAG2,PDK1,PKN2,NRDE2,MTMR4,ATXN2L,DNASE1,CCNT1,TTC3,STXBP5,ATF7IP,BRD4,MOSPD2,RSPH3,HERC4,MBOAT1,LUZP1,DPY19L3,SMG7,LCORL,CSNK1G1,CEP57,HOOK3,ARIH2,SPICE1,DIP2C,ARMCX4,SAMD8,DDX55,THOC1,WDR59,OTUD7B,UBA6,ACAP2,NUMA1,TNKS2,LANCL1,OXR1,AP5M1,DENND11,LYSMD3,LTO1,RAD54L2,ATP7A,APBB2,MSH3,ALS2CL,NEK9,EP400,UACA,DDX6,SESTD1,NRBP2,SLX4,MECP2,NBPF11,UBAP2L,AASS,ACIN1,TNRC18,GTF2A1,BOD1L1,CHD2,NBPF15,METTL14,ERN1,OTUD4,HEMK1,HERC1,POLR1A,ZSWIM8,PLXNB3,HERC3,GDAP2,RAD50,SBF2,LPP,LRBA,ELP1,SYNGAP1,XAF1,CHD1,HECTD4,ZEB2,YY1AP1,EP400P1,HPS4,NBPF1,USP45,UBE3B,CARD8,GOPC,NCOA2,HECTD2,PLCG1,PRKAB2,RALGAPA1,TMX3,MDM1,MED13L,FCHO2,POLH,FBXO41,SMG6,NEK4,SPIN3,TBRG1,ITSN1,FAT1,PPP1R12A,KLHDC1,RNPC3,MNT,DOP1A,BTBD19,KIAA2026,KMT2C,CCSAP,SEMA4C,SIKE1,FBXO22,SCAF4,CCNT2,RNF123,ZNF91,NBEAL1,TRAPPC11,TUBGCP5,RBM25,UBR5,RESF1,ETNK1,EPC1,ZC3H13,PANK3,ZNF717,HEATR6,MAP3K7,ARGLU1,YJEFN3,NEPRO,LIG4,SRPK2,USP37,SGSM2,SMAD3,VMAC,TMEM161B,KRR1,TBC1D2B,FAM149B1,DENND1B,ADAM17,PHLPP2,HOOK2,PPP4R3A,RFX3,VWA8,USP48,PARP8,HELZ,LRRK1,ELF2,ZNF43,CDC14B,TPP2,HDAC4,ADAT1,SOS1,JMY,AVL9,ZDHHC21,WDR37,GGA3,CPEB2,TRIM66,TNPO1,MSL2,STAG1,ZCCHC8,ITFG2,KAT7,GNA13,AASDH,RABGAP1L,KIAA0895L,TRANK1,MAML2,PHYKPL,SF3B1,POFUT2,ANKIB1,ATP11B,GOLGA1,MINDY2,USP24,SOS2,POGZ,BRWD3,AKAP10,TANC2,MAP3K12,FUBP1,SMPD4,TRIM33,NBAS,KIDINS220,DCAF1,DDX17,FMR1,DHX33,CTDSPL2,SMC6,CDAN1,EXOC6B,SLC24A1,GPALPP1,DLG1,ABCC5,STK36,GOLGA8B,ALPK1,ZFX,CBWD2,GOLGA8A,RABGAP1,SETD4,TP53BP2,TYW5,USP3,DST,CEP162,TRIP12,ZFP28,RAD18,ZNF17,SRCAP,CRBN,RNF169,ZNF84,BAZ2B,CCP110,BICRAL,WDR26,PRPF40B,PCMTD1,JMJD1C,ARHGAP21,REPS1,GSDMB,CPLANE1,ERI2,OXNAD1,NBPF9,BRD1,SPATA13,CHM,SUGP2,PHF10,UBE4A,ZBED4,ZFP14,TUBGCP6,CCDC14,SMARCC2,MYO9A,KLHL42,KNTC1,SAMD12,EXOC8,NIPA1,STRN3,SCYL3,OGT,TNIK,BLZF1,CLOCK,TUT4,DNAJC16,RBM15,BBIP1,OSGEP,CLK1,ATXN7,IGF1R,RFX7,MAP2K4,VTI1A,MBD5,GON4L,CDC27,CERS6,CHIC1,KLHL11,ZNF45,YTHDC1,PASK,CCDC191,HIF1AN,TNKS,NR2C2,TAF1A,BICD1,PIK3C2B,GLS,MBTD1,SEPSECS,MAN1A2,CEP192,ARHGEF11,VPS13A,HSPH1,ZFP3,SH3PXD2A,RNF214,THADA,ILF3,LENG8,SMURF2,PPWD1,RC3H2,SLC30A7,CLASP1,NHLRC3,SH3YL1,CASP2,KRIT1,COG5,PROSER1,MAML3,WAPL,ZZZ3,UGGT2,ZMYM5,C2CD3,TAMM41,CSNK1D,PDCD6IP,PCID2,SETX,TOP3A,TRIM13,PHC1,RAI1,SACS,PUS7L,GGNBP2,ERCC6L2,PPP1R13B,RTTN,REV3L,AFDN,PEX26,DOCK7,ENOSF1,MAU2,DZIP3,GANC,WDR7,COA1,FNBP4,MMS19,TMF1,SSH2,MLLT10,DENND5B,TSTD2,MED23,TSNARE1,NSUN2,SPTAN1,ZNF720,MIER3,BTBD7,RAB11FIP3,GPBP1,ARHGAP32,EHMT1,TTBK2,SPRED2,TP53BP1,CDK19,ZFHX3,ARID4A,SMAD4,TTC37,VCPIP1,TGOLN2,WDR27,EME2,PLXNB1,PNISR,GEN1,SFPQ,TMEM63A,AMT,ZNF10,FAM126B,CEP120,CDC42BPA,B3GALNT2,TGFBRAP1,ROCK1,FAM214A,N4BP2,ARID2,STAM2,MALT1,PLEKHH1,HEATR1,KLHDC4,PBRM1,CCNL2,ANKRD10,TENT4A,PPP4R3B,DDTL,PHF8,SSH1,OSBPL7,EEA1,LPIN1,CHD6,ANKS6,PCSK7,FBXW7,ZFP30,SPRTN,POLI,PRPF39,ITSN2,SRRM2,BTN2A1,MTHFR,SLC4A7,CMTR2,GORAB,UBR3,CEP44,PROSER3,PTPN23,KRAS,ANKRA2,CELF1,CDC40,EPG5,PMS1,QRICH1,FAM160A2,ZNF83,FAM161B,EDA2R,DPH1,UHRF1BP1L,RUSF1,SZT2,USP32,MYO18A,STK4,CRAMP1,CCAR1,SACM1L,CENPJ,TMEM131,AREL1,SLC10A7,PCBD2,APBB3,DYNC1LI2,RUFY2,WBP1,TEPSIN,HCFC1,LRCH4,AGAP6,CDC42BPB,RBM39,GPATCH8,PGS1,MDC1,FAM120B,INTS3,PDS5A,CSAD,PHLDB1,ATRX,RBM28,IKZF5,C2CD5,FRA10AC1,ZBTB5,PTEN,BCLAF1,PIK3CB,OGA,RASAL2,PAN3,MED28,RNF168,MAP3K9,NUMB,SCLT1,ANKRD52,DPP8,MTG1,APAF1,TOGARAM1,FAM76B,GTPBP3,ADNP,MBD1,LMO7,JARID2,ARID1B,ERC1,TCF20,MRNIP,SS18L1,ABL2,RSBN1L,RAD52,CEP43,URB1,DYRK1A,STRN,MARF1,INO80,CEP290,NEK1,WSB1,RSPRY1,ALS2,NSD2,TRIM52,NCBP3,DEPDC5,ZC3HAV1L,TTC13,BAHCC1,BRPF3,CUL4A,MTX3,SAP130,YOD1,MCM3AP,TUT7,STRADA,NAF1,TJP1,LMBRD2,ARNT,CREBBP,HTT,DNAJC11,SFMBT1,C19orf71,DOT1L,SDAD1,LDB1,LONP2,FRYL,USP31,DENND4B,NSUN3,TEP1,RPRD2,NOTCH2,MAN2C1,TAB3,TBC1D5,WHAMM,CCDC57,TET2,RABL2B,CFAP97,SPAG9,TTF2,MYCBP2,GLYR1,ANAPC1,DHX38,LYSMD4,CCDC88A,RPS6KA3,N4BP2L2,L3HYPDH,FBXL4,UNKL,NEDD4,GPCPD1,ZNF24,ASCC3,SIPA1L1,KMT2B,TP53INP1,KLHL28,KDM5B,RSF1,NUP58,PIAS3,CLK4,HMGXB3,KDM5A,SERAC1,NOD1,ARIH1,ARID4B,CAPN8,FAN1,RABL2A,SLC25A29,FOXJ3,KDM7A,MIER1,NCOA6,TTC21B,NPAT,ANKRD11,AP1G1,ARID1A,RAB3GAP2,SETD5,ZFP62,SOCS4,BRAF,OSBPL3,PTPN14,GUF1,CUL9,CAMSAP1,LRRC37B,SKIL,TAOK3,ANKRD28,R3HDM1,SLC25A16,SMURF1,FAM193B,LSM11,CCDC186,BTBD9,ZFC3H1,SP3,ARHGEF7,VCPKMT,HIPK1,SIK3,PIK3C2A,PAFAH1B1,DHX16,EIF2AK2,ZNF69,FMNL3,MARCHF6,RFFL,EIF4ENIF1,CUL3,LTN1,ATM,UBN1,SNX19,MED14,PLEKHG5,CDK13,TRAF6,RELCH,RSBN1,RBM27,ATG12,PIBF1,AHI1,RORA,CNOT6L,PGAP1,FAM111A,USP53,PARP11,ANKMY1,UBR1,PHF12,SNRNP48,RC3H1,ZNF44,SLC12A6,SMG1,XIAP,BCLAF3,MFSD9,DNAJB14,MTOR,MCM9,IFT80,ZCCHC14,MYO5C,ATP11A,UBN2,FMN1,KIAA0586,ABHD18,SLC5A3,SMCHD1,PRR12,POLR2A,CREBRF,ANGEL1,SMYD4,PRKD3,ZNF41,ZC3H8,LATS1,AKT3,TRAPPC6B,RAB2B,SLC38A9,APPL2,EPS15,HEATR5A,KIAA1958,KMT2D,SYNRG,USP40,TUBGCP3,ROCK2,BCL9L,ANKRD26,GSAP,C2orf68,WAC,MRE11,GK5,NOL4L,RBM6,TRMT13,ARHGAP12,NT5C2,VPS54,CCDC142,PAXIP1,SYNE2,CBX5,EPM2AIP1,NAA35,FAM168A,MAML1,ANKFY1,BORCS5,TPR,PIGB,CEPT1,DMXL2,ATAT1,INVS,WDR11,KDM3B,ADPGK,AKAP8,CEP57L1,SEC24B,VPS13B,KATNBL1,TAF4,TBP,GABPA,SLX4IP,GATAD1,TET3,TTC19,BNIP2,PDPK1,PPM1A,CLK3,INCA1,LRCH3,UBXN7,CNOT4,SON,MC1R,RLIM,ZMIZ1,KLHL29,PTCD3,MAGI1,NFRKB,BRWD1,CNOT1,NIN,DDI2,CHD3,NAPEPLD,RO60,TBCK,SP4,SETD6,LNPEP,KIAA0753,DDHD1,MAP4K5,DLG5,RBM41,CCNL1,TLK1,NCOR1,MADD,RBM45,NFXL1,ADCY6,WDR36,RANBP10,RPS6KC1,KANSL1,ZMYM3,DNMT3A,ABCD4,HAUS6,TBC1D8,SYNJ1,NEIL1,DDX5,STX17,PPP2R5E,LYST,PNN,MICAL3,LUC7L2,GAPVD1,ARHGAP5,MROH1,SNX13,TRA2A,KMT2A,REV1,NEURL4,NRIP2,FAM135A,CFAP44,RBM19,HACE1,GABPB2,ATXN7L1,MAPKBP1,ODF2L,TAF9B,SLC26A6,TRIM25,NRF1,WASHC4,ZXDC,FNDC3A,PIGL,ZC3H7A,BAZ2A,DCAF8,ABCA5,SNX29,MAP3K4,CEP350,CLASP2,SUPT7L,PSME4,RICTOR,OSGEPL1,CYTH1,RBM4,RAB11FIP2,EXOG,GABRE,SCAI,MYO19,GAN,DCAF5,GCN1,TENT4B,TIRAP,RAB22A,SCAPER,WDR20,LRP6,EZH1,ELMOD3,FBXW2,POLR3A,SLC7A6,ASXL2,MAP4K3,PFAS,VPS52,ELK4,CNNM3,ASAP1,USF3,FBXO11,LPIN2,ZMAT1,RERE,APPBP2,CNOT2,ABCA1,RBM48,PIKFYVE,AARSD1,RBM5,ULK2,NKTR,MTF1,FARP2,CEP170,SCYL2,ERCC6,PARP6,WDR33,OBSCN,NRIP1,TRA2B,PIAS2,ASXL1,TSC1,USPL1,PRPF4B | | Exist in public source | | NA |
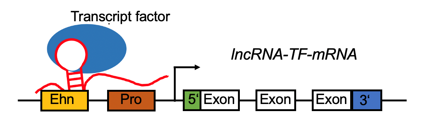
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
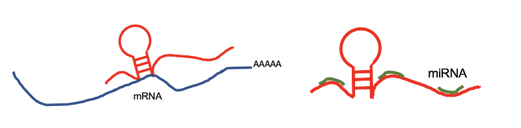
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | DDT,RNF181,ECH1,GET3,LAMTOR2,MRPL58,UXT,MRPL51,NEDD8,AURKAIP1,NDUFB2,ALDOA,MRPL40,BLOC1S1,ATP5PO,SLC25A11,GSTP1,UROD,NAA38,CIB1,BCL7C,CLIC1,ATP6V0B,COX6A1,RPS19BP1,CHMP2A,NDUFS5,MFSD5,ARPC4,EDF1,CLPP,EBP,TOMM7,COMMD9,DYNLRB1,SERF2,ATP5ME,COX4I1,UBL5,C12orf57,VAMP8,OST4,SH3BGRL3,RPL35,GPX4,POLR2J,SNRPD2,FAM174C,ATP5MC2,MYG1,MICOS13,NDUFA11,TSR3,COX6B1,PFDN5,CAPNS1,WDR83OS,UQCRC1,PSMC3,TMEM208,ISOC2,FKBP8,ROMO1,COPE,MRPS12,NDUFB10,BAD,PPP1CA,PFN1,NDUFB9,NELFE,APRT,ATP5PD,PPP4C,COPS9,PSENEN,COMMD1,CCDC124,COX14,PARK7,NDUFB1,NDUFA12,ATP5MK,GPX1,DYNLL1,PFDN1,MRPL28,NDUFC1,RNH1,NDUFB4,ELOB,HSD17B10,EIF6,UQCRQ,TXN2,FAM32A,MRPL47,TEX264,TMEM205,PRDX5,HIGD2A,PSMD4,GSTO1,MRPL54,CUTA,MZT2B,AP2S1,ECI1,ATP5MJ,NDUFB11,NDUFS6,VKORC1,NDUFS3,LAMTOR4,NAXE,CIAO2B,ATP6V1F,BANF1,DGCR6L,MRPL20,PGLS,ATP5MC1,ATP5PF,TRAPPC1,CFL1,HINT2,PSMD8,EMC7,NDUFS8,TMEM258,ELOF1,ARF5,COX8A,MYL6,COX7C,C19orf53,COPS6,COX5A,COX7A2,UBA52,EIF3K,ALKBH7,TRAPPC2L,MRPS15,TMEM219,ARL2,NDUFB7,ATP5MF,PSMB6,PPIA,NDUFC2,MRPL34,NSMCE1,PSMB3,BRI3,CD63,MTLN,HCFC1R1,OCEL1,NDUFB8,NDUFA3,NOP10,NDUFA1,NDUFA6,SWI5,MSRB1 | | Exist in public source | | NA |
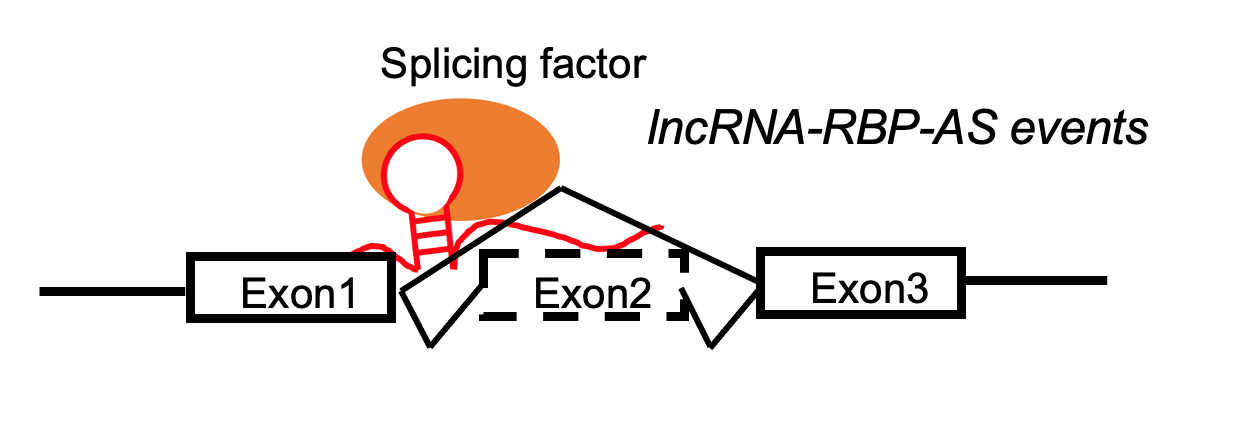
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation |
| -lncRNA and exonskipping events are negatively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000278864 | ENSG00000005810 | MYCBP2 | BLCA,BRCA,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000007168 | PAFAH1B1 | BLCA,CHOL,COAD,GBM,KIRC,LAML,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000007545 | CRAMP1 | BLCA,CHOL,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000008869 | HEATR5B | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000011021 | CLCN6 | BLCA,BRCA,COAD,GBM,KIRC,LAML,LUAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000018189 | RUFY3 | BLCA,BRCA,COAD,GBM,KIRC,LUAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000019995 | ZRANB1 | BLCA,COAD,GBM,KIRC,LAML,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000031003 | FAM13B | BLCA,BRCA,COAD,GBM,KIRC,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000038219 | BOD1L1 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000047056 | WDR37 | BLCA,BRCA,COAD,GBM,KIRC,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000047578 | KATNIP | BLCA,CHOL,COAD,GBM,KIRC,PAAD,PCPG,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000053702 | NRIP2 | BLCA,BRCA,CHOL,COAD,GBM,LAML,LUAD | | GBM | | ENSG00000278864 | ENSG00000054690 | PLEKHH1 | BLCA,BRCA,COAD,ESCA,GBM,KIRC,LGG,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000054793 | ATP9A | BLCA,CHOL,ESCA,GBM,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000059758 | CDK17 | BLCA,COAD,GBM,KIRC,LAML,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000065559 | MAP2K4 | BLCA,COAD,GBM,LUAD,PCPG | | GBM | | ENSG00000278864 | ENSG00000066739 | ATG2B | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000066933 | MYO9A | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000067369 | TP53BP1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000075391 | RASAL2 | BLCA,BRCA,COAD,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000075539 | FRYL | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000075711 | DLG1 | BLCA,CHOL,COAD,GBM,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000082269 | FAM135A | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000085224 | ATRX | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000085382 | HACE1 | BLCA,BRCA,GBM,KIRC,LAML,LGG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000088387 | DOCK9 | BLCA,BRCA,CHOL,COAD,GBM,LUAD,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000088854 | C20orf194 | BLCA,BRCA,GBM,KIRC,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000089057 | SLC23A2 | BLCA,BRCA,COAD,GBM,KIRC,LAML,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000092148 | HECTD1 | BLCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000092978 | GPATCH2 | BLCA,CHOL,COAD,KIRC,LUAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000095787 | WAC | BLCA,CHOL,COAD,GBM,KIRC,TGCT | | GBM | | ENSG00000278864 | ENSG00000100030 | MAPK1 | BLCA,CHOL,COAD,GBM,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000100483 | VCPKMT | BLCA,CHOL,COAD,GBM,KIRC,LUAD,PAAD,THCA | | GBM | | ENSG00000278864 | ENSG00000100523 | DDHD1 | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000100614 | PPM1A | BLCA,CHOL,COAD,GBM,KIRC,LUAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000102606 | ARHGEF7 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PAAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000103404 | USP31 | BLCA,BRCA,COAD,GBM,KIRC,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000103657 | HERC1 | BLCA,BRCA,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000104093 | DMXL2 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000104643 | MTMR9 | BLCA,COAD,GBM,KIRC,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000106144 | CASP2 | BLCA,CHOL,COAD,KIRC,PAAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000108306 | FBXL20 | BLCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000109452 | INPP4B | BLCA,CHOL,COAD,KIRC,LAML,LUAD | CHOL | | | ENSG00000278864 | ENSG00000109670 | FBXW7 | BLCA,BRCA,COAD,GBM,KIRC,LUAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000110844 | PRPF40B | BLCA,BRCA,CHOL,GBM,LGG,PCPG,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000112159 | MDN1 | BLCA,BRCA,COAD,ESCA,GBM,KIRC,LAML,LGG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000112624 | BICRAL | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000113441 | LNPEP | BLCA,BRCA,COAD,GBM,KIRC,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000115808 | STRN | BLCA,BRCA,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000115977 | AAK1 | BLCA,BRCA,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000116580 | GON4L | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000116903 | EXOC8 | BLCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000120694 | HSPH1 | BLCA,CHOL,GBM,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000121274 | TENT4B | BLCA,COAD,GBM,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000125633 | CCDC93 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000127481 | UBR4 | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000128833 | MYO5C | BLCA,CHOL,COAD,ESCA,LAML,LUAD,PAAD,STAD,THCA | CHOL | | | ENSG00000278864 | ENSG00000128872 | TMOD2 | BLCA,GBM,KIRC,LUAD,PCPG | | GBM | | ENSG00000278864 | ENSG00000129534 | MIS18BP1 | BLCA,CHOL,COAD,KIRC,LUAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000131437 | KIF3A | BLCA,CHOL,COAD,GBM,KIRC,LAML,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000132680 | KHDC4 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000132694 | ARHGEF11 | BLCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000133056 | PIK3C2B | BLCA,CHOL,ESCA,KIRC,LAML,PAAD,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000133460 | SLC2A11 | BLCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000134318 | ROCK2 | BLCA,CHOL,COAD,ESCA,GBM,KIRC,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000134444 | RELCH | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000134909 | ARHGAP32 | BLCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,PAAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000135541 | AHI1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000136100 | VPS36 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000138018 | SELENOI | BLCA,COAD,GBM,KIRC,PCPG | | GBM | | ENSG00000278864 | ENSG00000138593 | SECISBP2L | BLCA,COAD,ESCA,GBM,KIRC,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000139697 | SBNO1 | BLCA,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000140386 | SCAPER | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000140443 | IGF1R | BLCA,CHOL,COAD,LAML,PCPG,TGCT | CHOL | | | ENSG00000278864 | ENSG00000140992 | PDPK1 | BLCA,CHOL,ESCA,GBM,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000141252 | VPS53 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000144036 | EXOC6B | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000144357 | UBR3 | BLCA,BRCA,COAD,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000144445 | KANSL1L | BLCA,BRCA,CHOL,COAD,ESCA,KIRC,LAML,LUAD,PAAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000146858 | ZC3HAV1L | BLCA,BRCA,CHOL,COAD,KIRC,LAML,PAAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000148690 | FRA10AC1 | BLCA,COAD,GBM,KIRC,LAML | | GBM | | ENSG00000278864 | ENSG00000151208 | DLG5 | BLCA,CHOL,GBM,LAML,LGG,PAAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000151276 | MAGI1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,STAD | | GBM | | ENSG00000278864 | ENSG00000151914 | DST | BLCA,BRCA,COAD,GBM,KIRC,LGG,TGCT | | GBM | | ENSG00000278864 | ENSG00000152061 | RABGAP1L | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LUAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000152409 | JMY | BLCA,BRCA,COAD,GBM,LGG,LUAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000155111 | CDK19 | BLCA,CHOL,COAD,GBM,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000155744 | FAM126B | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LUAD,PAAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000156671 | SAMD8 | BLCA,CHOL,COAD,GBM,KIRC,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000157106 | SMG1 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000157764 | BRAF | BLCA,BRCA,CHOL,COAD,GBM,KIRC,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000160299 | PCNT | BLCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,THCA | | GBM | | ENSG00000278864 | ENSG00000162885 | B3GALNT2 | BLCA,BRCA,CHOL,COAD,KIRC,LUAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000163257 | DCAF16 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LUAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000163539 | CLASP2 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000163625 | WDFY3 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000164187 | LMBRD2 | BLCA,COAD,ESCA,GBM,KIRC,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000164327 | RICTOR | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000164506 | STXBP5 | BLCA,BRCA,COAD,GBM,KIRC,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000165338 | HECTD2 | BLCA,BRCA,GBM,KIRC,LGG,THCA | | GBM | | ENSG00000278864 | ENSG00000165650 | PDZD8 | BLCA,BRCA,COAD,ESCA,GBM,TGCT | | GBM | | ENSG00000278864 | ENSG00000165699 | TSC1 | BLCA,BRCA,COAD,ESCA,GBM,KIRC,LAML,LGG,PAAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000166783 | MARF1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000167548 | KMT2D | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000167615 | LENG8 | BLCA,BRCA,CHOL,ESCA,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000168438 | CDC40 | BLCA,COAD,GBM,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000169118 | CSNK1G1 | BLCA,CHOL,COAD,KIRC,LAML,LUAD,PCPG,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000170921 | TANC2 | BLCA,CHOL,GBM,KIRC,LGG,LUAD,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000173064 | HECTD4 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000173681 | BCLAF3 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000174405 | LIG4 | BLCA,BRCA,COAD,ESCA,GBM,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000175073 | VCPIP1 | BLCA,CHOL,COAD,GBM,KIRC,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000176542 | USF3 | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000176994 | SMCR8 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000178761 | FAM219B | BLCA,BRCA,CHOL,COAD,GBM,LGG,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000179832 | MROH1 | BLCA,CHOL,ESCA,GBM,KIRC,PAAD,STAD | CHOL | GBM | | ENSG00000278864 | ENSG00000182400 | TRAPPC6B | BLCA,CHOL,COAD,GBM,KIRC,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000182670 | TTC3 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000182700 | IGIP | BLCA,COAD,GBM,KIRC,TGCT | | GBM | | ENSG00000278864 | ENSG00000183530 | PRR14L | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000183826 | BTBD9 | BLCA,BRCA,COAD,GBM,LAML,TGCT | | GBM | | ENSG00000278864 | ENSG00000184384 | MAML2 | BLCA,CHOL,COAD,LUAD,PCPG,THCA | CHOL | | | ENSG00000278864 | ENSG00000184402 | SS18L1 | BLCA,COAD,GBM,KIRC,LGG,LUAD,PAAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000185658 | BRWD1 | BLCA,BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000185716 | MOSMO | BLCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000196187 | TMEM63A | BLCA,CHOL,ESCA,GBM,LAML,LUAD,PAAD,STAD,THCA | | GBM | | ENSG00000278864 | ENSG00000196712 | NF1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000197217 | ENTPD4 | BLCA,COAD,ESCA,GBM,KIRC,LAML,LGG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000197283 | SYNGAP1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LGG,STAD,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000197312 | DDI2 | BLCA,COAD,GBM,KIRC,LUAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000197386 | HTT | BLCA,COAD,GBM,KIRC,LAML,LUAD,PCPG,STAD | | GBM | | ENSG00000278864 | ENSG00000197969 | VPS13A | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000198408 | OGA | BLCA,BRCA,COAD,ESCA,GBM,KIRC,LAML,PAAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000198707 | CEP290 | BLCA,BRCA,CHOL,COAD,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000198718 | TOGARAM1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000198920 | KIAA0753 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000204116 | CHIC1 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LUAD,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000204130 | RUFY2 | BLCA,BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,PAAD,STAD,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000204217 | BMPR2 | BLCA,BRCA,CHOL,COAD,GBM,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000213462 | ERV3-1 | BLCA,BRCA,CHOL,COAD,KIRC,LAML,LGG,PAAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000214021 | TTLL3 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000257093 | DENND11 | BLCA,BRCA,CHOL,COAD,ESCA,GBM,KIRC,LAML,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000258839 | MC1R | BLCA,CHOL,LAML,LGG,PAAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000040199 | PHLPP2 | BRCA,ESCA,GBM,KIRC,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000068024 | HDAC4 | BRCA,CHOL,COAD,GBM,KIRC,LGG,PCPG,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000083290 | ULK2 | BRCA,COAD,GBM,KIRC,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000083642 | PDS5B | BRCA,CHOL,COAD,GBM,KIRC,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000083857 | FAT1 | BRCA,CHOL,COAD,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000087448 | KLHL42 | BRCA,COAD,GBM,KIRC,PCPG,THCA | | GBM | | ENSG00000278864 | ENSG00000091157 | WDR7 | BRCA,GBM,KIRC,LUAD,PCPG | | GBM | | ENSG00000278864 | ENSG00000100150 | DEPDC5 | BRCA,COAD,GBM,KIRC,LUAD,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000107560 | RAB11FIP2 | BRCA,CHOL,COAD,GBM,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000111731 | C2CD5 | BRCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,PCPG,STAD,THCA | | GBM | | ENSG00000278864 | ENSG00000111752 | PHC1 | BRCA,CHOL,GBM,KIRC,LAML,LGG,PAAD,PCPG,THCA | CHOL | | | ENSG00000278864 | ENSG00000115137 | DNAJC27 | BRCA,GBM,KIRC,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000124422 | USP22 | BRCA,CHOL,GBM,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000128731 | HERC2 | BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000128881 | TTBK2 | BRCA,GBM,KIRC,LAML,LUAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000131018 | SYNE1 | BRCA,GBM,KIRC,LAML,LGG | | GBM | | ENSG00000278864 | ENSG00000137343 | ATAT1 | BRCA,CHOL,COAD,KIRC,LAML,PAAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000137802 | MAPKBP1 | BRCA,COAD,GBM,KIRC,LAML,LGG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000138002 | IFT172 | BRCA,CHOL,ESCA,GBM,KIRC,LAML,LGG,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000140199 | SLC12A6 | BRCA,COAD,GBM,KIRC,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000143322 | ABL2 | BRCA,CHOL,COAD,GBM,KIRC,LAML,LUAD,PCPG,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000143776 | CDC42BPA | BRCA,CHOL,COAD,ESCA,GBM,KIRC,LUAD,PAAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000151240 | DIP2C | BRCA,CHOL,GBM,LGG,TGCT | | GBM | | ENSG00000278864 | ENSG00000151835 | SACS | BRCA,GBM,KIRC,LAML,LGG,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000151849 | CENPJ | BRCA,CHOL,COAD,GBM,KIRC,LGG,LUAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000153317 | ASAP1 | BRCA,CHOL,COAD,KIRC,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000167522 | ANKRD11 | BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000169641 | LUZP1 | BRCA,COAD,GBM,LAML,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000170456 | DENND5B | BRCA,COAD,ESCA,GBM,KIRC,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000173611 | SCAI | BRCA,GBM,LAML,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000174373 | RALGAPA1 | BRCA,CHOL,COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000178502 | KLHL11 | BRCA,COAD,GBM,KIRC,LAML,LGG,LUAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000185158 | LRRC37B | BRCA,CHOL,KIRC,LAML,LUAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000185917 | SETD4 | BRCA,CHOL,COAD,KIRC,LAML,LGG,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000196950 | SLC39A10 | BRCA,CHOL,COAD,LAML,LUAD,PCPG,THCA | CHOL | | | ENSG00000278864 | ENSG00000197555 | SIPA1L1 | BRCA,COAD,GBM,LUAD,PCPG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000197776 | KLHDC1 | BRCA,GBM,KIRC,LGG,LUAD,PCPG,THCA | | GBM | | ENSG00000278864 | ENSG00000198399 | ITSN2 | BRCA,COAD,ESCA,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000204681 | GABBR1 | BRCA,CHOL,COAD,GBM,KIRC,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000008710 | PKD1 | CHOL,GBM,KIRC,LAML,LGG,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000010322 | NISCH | CHOL,GBM,KIRC,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000051382 | PIK3CB | CHOL,COAD,GBM,LUAD,STAD | | GBM | | ENSG00000278864 | ENSG00000068650 | ATP11A | CHOL,COAD,ESCA,GBM,LGG,PAAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000070882 | OSBPL3 | CHOL,COAD,ESCA,KIRC,PAAD,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000080839 | RBL1 | CHOL,COAD,LUAD,PCPG,TGCT | CHOL | | | ENSG00000278864 | ENSG00000088808 | PPP1R13B | CHOL,GBM,PAAD,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000103540 | CCP110 | CHOL,COAD,GBM,KIRC,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000105443 | CYTH2 | CHOL,ESCA,GBM,LAML,PAAD | CHOL | | | ENSG00000278864 | ENSG00000105738 | SIPA1L3 | CHOL,GBM,LAML,STAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000107862 | GBF1 | CHOL,GBM,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000107863 | ARHGAP21 | CHOL,COAD,GBM,KIRC,LGG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000108175 | ZMIZ1 | CHOL,GBM,LAML,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000108799 | EZH1 | CHOL,COAD,GBM,KIRC,LGG,LUAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000110171 | TRIM3 | CHOL,GBM,KIRC,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000115365 | LANCL1 | CHOL,GBM,LAML,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000115419 | GLS | CHOL,COAD,GBM,LAML,LUAD,PCPG | CHOL | GBM | | ENSG00000278864 | ENSG00000116584 | ARHGEF2 | CHOL,ESCA,GBM,PAAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000118965 | WDR35 | CHOL,COAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000119285 | HEATR1 | CHOL,COAD,KIRC,LAML,LGG,PAAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000119771 | KLHL29 | CHOL,GBM,KIRC,PCPG,TGCT | CHOL | | | ENSG00000278864 | ENSG00000120370 | GORAB | CHOL,KIRC,LAML,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000123200 | ZC3H13 | CHOL,COAD,GBM,KIRC,LGG,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000130559 | CAMSAP1 | CHOL,COAD,GBM,KIRC,LGG,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000130702 | LAMA5 | CHOL,GBM,LGG,PAAD,STAD,THCA | CHOL | | | ENSG00000278864 | ENSG00000131080 | EDA2R | CHOL,KIRC,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000131788 | PIAS3 | CHOL,KIRC,LAML,LGG,PAAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000133059 | DSTYK | CHOL,COAD,LUAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000136237 | RAPGEF5 | CHOL,COAD,GBM,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000138078 | PREPL | CHOL,COAD,GBM,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000138834 | MAPK8IP3 | CHOL,ESCA,GBM,KIRC,LAML,LUAD,PAAD,PCPG,STAD,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000139116 | KIF21A | CHOL,COAD,GBM,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000143376 | SNX27 | CHOL,COAD,GBM,LUAD,PAAD,PCPG | CHOL | | | ENSG00000278864 | ENSG00000143498 | TAF1A | CHOL,KIRC,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000143569 | UBAP2L | CHOL,LAML,PAAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000143643 | TTC13 | CHOL,COAD,ESCA,KIRC,LUAD,PCPG | CHOL | | | ENSG00000278864 | ENSG00000143748 | NVL | CHOL,COAD,KIRC,LAML,PCPG | CHOL | | | ENSG00000278864 | ENSG00000148606 | POLR3A | CHOL,GBM,KIRC,LAML,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000152104 | PTPN14 | CHOL,COAD,KIRC,LUAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000156050 | FAM161B | CHOL,GBM,LAML,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000160796 | NBEAL2 | CHOL,ESCA,GBM,KIRC,PAAD,STAD,THCA | CHOL | | | ENSG00000278864 | ENSG00000163013 | FBXO41 | CHOL,GBM,LUAD,PCPG,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000163482 | STK36 | CHOL,COAD,ESCA,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000163617 | CCDC191 | CHOL,COAD,KIRC,LUAD,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000163872 | YEATS2 | CHOL,COAD,GBM,KIRC,LAML,PAAD,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000164877 | MICALL2 | CHOL,ESCA,GBM,KIRC,PAAD,STAD | CHOL | | | ENSG00000278864 | ENSG00000165138 | ANKS6 | CHOL,ESCA,GBM,LAML,PAAD,TGCT | CHOL | | | ENSG00000278864 | ENSG00000167595 | PROSER3 | CHOL,GBM,KIRC,PAAD,PCPG,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000169398 | PTK2 | CHOL,GBM,KIRC,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000170004 | CHD3 | CHOL,GBM,PAAD,PCPG,TGCT | CHOL | GBM | | ENSG00000278864 | ENSG00000170113 | NIPA1 | CHOL,COAD,GBM,LUAD,PCPG | CHOL | GBM | | ENSG00000278864 | ENSG00000171045 | TSNARE1 | CHOL,GBM,LGG,PAAD,THCA | | GBM | | ENSG00000278864 | ENSG00000171680 | PLEKHG5 | CHOL,GBM,KIRC,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000172534 | HCFC1 | CHOL,GBM,LGG,PCPG,STAD,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000186174 | BCL9L | CHOL,GBM,LAML,LGG,PAAD,PCPG,THCA | CHOL | | | ENSG00000278864 | ENSG00000196535 | MYO18A | CHOL,GBM,LAML,LGG,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000197119 | SLC25A29 | CHOL,COAD,ESCA,GBM,KIRC,LAML,LGG,STAD | CHOL | | | ENSG00000278864 | ENSG00000197183 | NOL4L | CHOL,ESCA,GBM,KIRC,LAML,PAAD,PCPG,TGCT,THCA | CHOL | GBM | | ENSG00000278864 | ENSG00000198728 | LDB1 | CHOL,GBM,KIRC,TGCT,THCA | CHOL | | | ENSG00000278864 | ENSG00000198752 | CDC42BPB | CHOL,ESCA,GBM,LGG,PAAD,PCPG,STAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000241973 | PI4KA | CHOL,COAD,GBM,KIRC,LAML,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000243156 | MICAL3 | CHOL,COAD,GBM,KIRC,LAML,LGG,TGCT | | GBM | | ENSG00000278864 | ENSG00000278259 | MYO19 | CHOL,GBM,KIRC,LAML,LGG,PAAD,PCPG | CHOL | | | ENSG00000278864 | ENSG00000011523 | CEP68 | COAD,GBM,KIRC,LAML,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000019144 | PHLDB1 | COAD,GBM,KIRC,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000023287 | RB1CC1 | COAD,GBM,KIRC,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000048991 | R3HDM1 | COAD,GBM,KIRC,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000100580 | TMED8 | COAD,GBM,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000108669 | CYTH1 | COAD,GBM,KIRC,LGG,LUAD,PAAD,PCPG,STAD,THCA | | GBM | | ENSG00000278864 | ENSG00000117020 | AKT3 | COAD,GBM,KIRC,LGG,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000124406 | ATP8A1 | COAD,ESCA,GBM,KIRC,LUAD,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000138641 | HERC3 | COAD,GBM,KIRC,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000139190 | VAMP1 | COAD,ESCA,GBM,KIRC,LAML,LUAD,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000145495 | MARCHF6 | COAD,GBM,KIRC,LAML,LGG,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000152484 | USP12 | COAD,ESCA,GBM,KIRC,LUAD,PCPG,TGCT | | GBM | | ENSG00000278864 | ENSG00000160584 | SIK3 | COAD,GBM,KIRC,LAML,LUAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000170832 | USP32 | COAD,GBM,LUAD,PCPG,STAD | | GBM | | ENSG00000278864 | ENSG00000183060 | LYSMD4 | COAD,GBM,KIRC,LAML,LUAD,THCA | | GBM | | ENSG00000278864 | ENSG00000205726 | ITSN1 | COAD,GBM,KIRC,LUAD,THCA | | GBM | | ENSG00000278864 | ENSG00000197694 | SPTAN1 | ESCA,GBM,LAML,STAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000250067 | YJEFN3 | ESCA,GBM,KIRC,LGG,PAAD | | GBM | | ENSG00000278864 | ENSG00000068400 | GRIPAP1 | GBM,KIRC,LGG,PCPG,THCA | | GBM | | ENSG00000278864 | ENSG00000100241 | SBF1 | GBM,KIRC,LAML,LGG,PAAD,TGCT | | GBM | | ENSG00000278864 | ENSG00000168758 | SEMA4C | GBM,LAML,LGG,PAAD,THCA | | GBM | | ENSG00000278864 | ENSG00000196440 | ARMCX4 | GBM,KIRC,LAML,LGG,PCPG,THCA | | GBM | | ENSG00000278864 | ENSG00000275023 | MLLT6 | GBM,KIRC,LAML,LGG,PAAD,PCPG,TGCT,THCA | | GBM | | ENSG00000278864 | ENSG00000276293 | PIP4K2B | GBM,LAML,PCPG,TGCT,THCA | | GBM |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000548898 | -0.1009 | 1405 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000420846 | -0.1338 | 1504 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000552067 | -0.1278 | 1504 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000548160 | -0.1318 | 388 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000546939 | -0.1318 | 388 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000552692 | -0.1318 | 388 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000549117 | -0.2418 | 948 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000257857 | -0.2418 | 953 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000552754 | -0.5480 | 1295 | GBM | - | | ENSG00000278864 | ENST00000624039 | CD63 | ENSG00000135404 | ENST00000550776 | -0.8244 | 876 | GBM | - | | ENSG00000278864 | ENST00000624039 | GSTO1 | ENSG00000148834 | ENST00000539281 | -0.2929 | 1380 | - | CHOL | | ENSG00000278864 | ENST00000624039 | GSTO1 | ENSG00000148834 | ENST00000369710 | -0.2929 | 1380 | - | CHOL | | ENSG00000278864 | ENST00000624039 | GSTO1 | ENSG00000148834 | ENST00000369713 | -0.2929 | 1382 | - | CHOL | | ENSG00000278864 | ENST00000624039 | CLIC1 | ENSG00000213719 | ENST00000616760 | -0.1002 | 765 | GBM | - | | ENSG00000278864 | ENST00000624039 | ISOC2 | ENSG00000063241 | ENST00000438389 | -0.1084 | 1176 | - | CHOL | | ENSG00000278864 | ENST00000624039 | ISOC2 | ENSG00000063241 | ENST00000425675 | -0.1130 | 1180 | - | CHOL | | ENSG00000278864 | ENST00000624039 | ISOC2 | ENSG00000063241 | ENST00000085068 | -0.1130 | 1180 | - | CHOL | | ENSG00000278864 | ENST00000624039 | HSD17B10 | ENSG00000072506 | ENST00000375304 | -0.2067 | 870 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | HSD17B10 | ENSG00000072506 | ENST00000168216 | -0.2067 | 870 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000215061 | -0.1071 | 952 | - | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000601529 | -0.1069 | 1043 | - | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000600232 | -0.2092 | 1123 | - | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000597836 | -0.1129 | 1093 | - | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000599286 | -0.1272 | 328 | - | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000595573 | -0.2278 | 318 | - | CHOL | | ENSG00000278864 | ENST00000624039 | OCEL1 | ENSG00000099330 | ENST00000600826 | -0.2856 | 1127 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DDT | ENSG00000099977 | ENST00000398344 | -0.1649 | 222 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DDT | ENSG00000099977 | ENST00000350608 | -0.1649 | 222 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DDT | ENSG00000099977 | ENST00000428792 | -0.1649 | 227 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DDT | ENSG00000099977 | ENST00000404092 | -0.1220 | 235 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DDT | ENSG00000099977 | ENST00000403754 | -3.2400 | 1489 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DDT | ENSG00000099977 | ENST00000430101 | -0.1092 | 1255 | - | CHOL | | ENSG00000278864 | ENST00000624039 | ECH1 | ENSG00000104823 | ENST00000221418 | -0.1496 | 1441 | - | CHOL | | ENSG00000278864 | ENST00000624039 | UROD | ENSG00000126088 | ENST00000246337 | -0.1587 | 307 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DGCR6L | ENSG00000128185 | ENST00000248879 | -0.1204 | 824 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DGCR6L | ENSG00000128185 | ENST00000405465 | -0.1204 | 831 | - | CHOL | | ENSG00000278864 | ENST00000624039 | DGCR6L | ENSG00000128185 | ENST00000443409 | -0.1006 | 942 | - | CHOL | | ENSG00000278864 | ENST00000624039 | EBP | ENSG00000147155 | ENST00000495186 | -0.1221 | 217 | - | CHOL | | ENSG00000278864 | ENST00000624039 | VKORC1 | ENSG00000167397 | ENST00000394975 | -0.1096 | 1159 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | VKORC1 | ENSG00000167397 | ENST00000394971 | -0.1705 | 349 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | VKORC1 | ENSG00000167397 | ENST00000498155 | -0.1889 | 89 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | AURKAIP1 | ENSG00000175756 | ENST00000338370 | -0.2272 | 566 | - | CHOL | | ENSG00000278864 | ENST00000624039 | AURKAIP1 | ENSG00000175756 | ENST00000338338 | -0.2272 | 566 | - | CHOL | | ENSG00000278864 | ENST00000624039 | AURKAIP1 | ENSG00000175756 | ENST00000321751 | -0.2272 | 566 | - | CHOL | | ENSG00000278864 | ENST00000624039 | AURKAIP1 | ENSG00000175756 | ENST00000378853 | -0.4045 | 541 | - | CHOL | | ENSG00000278864 | ENST00000624039 | MSRB1 | ENSG00000198736 | ENST00000473663 | -0.1325 | 183 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | MSRB1 | ENSG00000198736 | ENST00000399753 | -0.2068 | 260 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | MSRB1 | ENSG00000198736 | ENST00000564908 | -0.7358 | 1416 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | POLR2J | ENSG00000005075 | ENST00000393794 | -0.1411 | 1231 | GBM | - | | ENSG00000278864 | ENST00000624039 | BCL7C | ENSG00000099385 | ENST00000215115 | -0.2751 | 903 | GBM | - | | ENSG00000278864 | ENST00000624039 | VAMP8 | ENSG00000118640 | ENST00000432071 | -0.1218 | 747 | GBM | - | | ENSG00000278864 | ENST00000624039 | VAMP8 | ENSG00000118640 | ENST00000263864 | -0.1103 | 330 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFDN5 | ENSG00000123349 | ENST00000551018 | -0.1824 | 850 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFDN5 | ENSG00000123349 | ENST00000351500 | -0.1824 | 850 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFDN5 | ENSG00000123349 | ENST00000550846 | -0.1824 | 850 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFDN5 | ENSG00000123349 | ENST00000243040 | -0.1064 | 295 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFDN5 | ENSG00000123349 | ENST00000334478 | -0.1824 | 850 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFDN5 | ENSG00000123349 | ENST00000549759 | -0.1864 | 853 | GBM | - | | ENSG00000278864 | ENST00000624039 | CLPP | ENSG00000125656 | ENST00000245816 | -0.1339 | 236 | GBM | - | | ENSG00000278864 | ENST00000624039 | CLPP | ENSG00000125656 | ENST00000596149 | -0.2629 | 390 | GBM | - | | ENSG00000278864 | ENST00000624039 | CLPP | ENSG00000125656 | ENST00000596605 | -0.1316 | 176 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000342669 | -0.1498 | 974 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000585392 | -0.1498 | 974 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000587367 | -0.1578 | 974 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000588599 | -0.1513 | 1340 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000391932 | -0.1513 | 1340 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000588301 | -0.1513 | 1340 | GBM | - | | ENSG00000278864 | ENST00000624039 | SNRPD2 | ENSG00000125743 | ENST00000590212 | -0.1275 | 266 | GBM | - | | ENSG00000278864 | ENST00000624039 | UXT | ENSG00000126756 | ENST00000333119 | -0.2822 | 907 | GBM | - | | ENSG00000278864 | ENST00000624039 | UXT | ENSG00000126756 | ENST00000485641 | -0.1430 | 857 | GBM | - | | ENSG00000278864 | ENST00000624039 | UXT | ENSG00000126756 | ENST00000335890 | -0.3617 | 306 | GBM | - | | ENSG00000278864 | ENST00000624039 | PGLS | ENSG00000130313 | ENST00000252603 | -0.1691 | 302 | GBM | - | | ENSG00000278864 | ENST00000624039 | PGLS | ENSG00000130313 | ENST00000594761 | -0.1173 | 130 | GBM | - | | ENSG00000278864 | ENST00000624039 | PGLS | ENSG00000130313 | ENST00000595782 | -0.1691 | 302 | GBM | - | | ENSG00000278864 | ENST00000624039 | TMEM258 | ENSG00000134825 | ENST00000537328 | -0.1037 | 929 | GBM | - | | ENSG00000278864 | ENST00000624039 | TMEM258 | ENSG00000134825 | ENST00000541893 | -0.1083 | 774 | GBM | - | | ENSG00000278864 | ENST00000624039 | TMEM258 | ENSG00000134825 | ENST00000545210 | -0.1074 | 774 | GBM | - | | ENSG00000278864 | ENST00000624039 | TMEM258 | ENSG00000134825 | ENST00000257262 | -0.1429 | 236 | GBM | - | | ENSG00000278864 | ENST00000624039 | TMEM258 | ENSG00000134825 | ENST00000535297 | -0.1423 | 293 | GBM | - | | ENSG00000278864 | ENST00000624039 | TMEM258 | ENSG00000134825 | ENST00000543510 | -0.2010 | 294 | GBM | - | | ENSG00000278864 | ENST00000624039 | RPL35 | ENSG00000136942 | ENST00000373570 | -0.1360 | 1190 | GBM | - | | ENSG00000278864 | ENST00000624039 | RPL35 | ENSG00000136942 | ENST00000493018 | -0.1044 | 1193 | GBM | - | | ENSG00000278864 | ENST00000624039 | RPL35 | ENSG00000136942 | ENST00000348462 | -0.5900 | 1418 | GBM | - | | ENSG00000278864 | ENST00000624039 | SERF2 | ENSG00000140264 | ENST00000409646 | -0.1037 | 1227 | GBM | - | | ENSG00000278864 | ENST00000624039 | SERF2 | ENSG00000140264 | ENST00000339624 | -0.1864 | 1199 | GBM | - | | ENSG00000278864 | ENST00000624039 | SERF2 | ENSG00000140264 | ENST00000402131 | -0.1467 | 896 | GBM | - | | ENSG00000278864 | ENST00000624039 | SERF2 | ENSG00000140264 | ENST00000403425 | -0.1240 | 846 | GBM | - | | ENSG00000278864 | ENST00000624039 | SERF2 | ENSG00000140264 | ENST00000409614 | -0.1240 | 846 | GBM | - | | ENSG00000278864 | ENST00000624039 | PPP4C | ENSG00000149923 | ENST00000279387 | -0.1069 | 295 | GBM | - | | ENSG00000278864 | ENST00000624039 | PPP4C | ENSG00000149923 | ENST00000561610 | -0.1069 | 354 | GBM | - | | ENSG00000278864 | ENST00000624039 | BANF1 | ENSG00000175334 | ENST00000527348 | -0.1336 | 305 | GBM | - | | ENSG00000278864 | ENST00000624039 | NOP10 | ENSG00000182117 | ENST00000557912 | -0.1086 | 938 | GBM | - | | ENSG00000278864 | ENST00000624039 | UBL5 | ENSG00000198258 | ENST00000586895 | -0.1210 | 1032 | GBM | - | | ENSG00000278864 | ENST00000624039 | UBL5 | ENSG00000198258 | ENST00000358666 | -0.1330 | 1031 | GBM | - | | ENSG00000278864 | ENST00000624039 | UBL5 | ENSG00000198258 | ENST00000590068 | -0.1330 | 1031 | GBM | - | | ENSG00000278864 | ENST00000624039 | UBL5 | ENSG00000198258 | ENST00000593087 | -0.1387 | 1267 | GBM | - | | ENSG00000278864 | ENST00000624039 | APRT | ENSG00000198931 | ENST00000378364 | -0.1239 | 1173 | GBM | - | | ENSG00000278864 | ENST00000624039 | APRT | ENSG00000198931 | ENST00000426324 | -0.1705 | 1348 | GBM | - | | ENSG00000278864 | ENST00000624039 | APRT | ENSG00000198931 | ENST00000567391 | -0.1360 | 1348 | GBM | - | | ENSG00000278864 | ENST00000624039 | APRT | ENSG00000198931 | ENST00000568319 | -0.1570 | 1348 | GBM | - | | ENSG00000278864 | ENST00000624039 | APRT | ENSG00000198931 | ENST00000563655 | -0.1705 | 1351 | GBM | - | | ENSG00000278864 | ENST00000624039 | APRT | ENSG00000198931 | ENST00000569616 | -0.1343 | 842 | GBM | - | | ENSG00000278864 | ENST00000624039 | GPX1 | ENSG00000233276 | ENST00000419783 | -0.1226 | 569 | GBM | - | | ENSG00000278864 | ENST00000624039 | GPX1 | ENSG00000233276 | ENST00000620890 | -0.1507 | 569 | GBM | - | | ENSG00000278864 | ENST00000624039 | ATP5MF | ENSG00000241468 | ENST00000414062 | -0.1041 | 902 | GBM | - | | ENSG00000278864 | ENST00000624039 | ATP5MF | ENSG00000241468 | ENST00000449683 | -0.1457 | 513 | GBM | - | | ENSG00000278864 | ENST00000624039 | ATP5MF | ENSG00000241468 | ENST00000359832 | -0.2760 | 908 | GBM | - | | ENSG00000278864 | ENST00000624039 | ATP5MF | ENSG00000241468 | ENST00000292475 | -0.2760 | 908 | GBM | - | | ENSG00000278864 | ENST00000624039 | ATP5MF | ENSG00000241468 | ENST00000488775 | -0.2760 | 908 | GBM | - | | ENSG00000278864 | ENST00000624039 | ATP5MF | ENSG00000241468 | ENST00000394186 | -0.5481 | 506 | GBM | - | | ENSG00000278864 | ENST00000624039 | GSTP1 | ENSG00000084207 | ENST00000398606 | -0.1742 | 973 | GBM | - | | ENSG00000278864 | ENST00000624039 | GSTP1 | ENSG00000084207 | ENST00000398603 | -0.1799 | 1147 | GBM | - | | ENSG00000278864 | ENST00000624039 | GSTP1 | ENSG00000084207 | ENST00000495996 | -0.6900 | 660 | GBM | - | | ENSG00000278864 | ENST00000624039 | C12orf57 | ENSG00000111678 | ENST00000537087 | -0.2348 | 1169 | GBM | - | | ENSG00000278864 | ENST00000624039 | C12orf57 | ENSG00000111678 | ENST00000229281 | -0.2348 | 1169 | GBM | - | | ENSG00000278864 | ENST00000624039 | C12orf57 | ENSG00000111678 | ENST00000540506 | -0.3160 | 1192 | GBM | - | | ENSG00000278864 | ENST00000624039 | PPIA | ENSG00000196262 | ENST00000451562 | -0.1516 | 312 | GBM | - | | ENSG00000278864 | ENST00000624039 | PPIA | ENSG00000196262 | ENST00000355968 | -0.1176 | 565 | GBM | - | | ENSG00000278864 | ENST00000624039 | PPIA | ENSG00000196262 | ENST00000620047 | -0.1176 | 565 | GBM | - | | ENSG00000278864 | ENST00000624039 | OST4 | ENSG00000228474 | ENST00000447619 | -0.1119 | 932 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000550697 | -0.1180 | 1025 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000548580 | -0.1096 | 1531 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000293422 | -0.1096 | 1532 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000348108 | -0.1180 | 1012 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000549017 | -0.1096 | 1532 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000549566 | -0.2650 | 1165 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000536128 | -0.1026 | 1083 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000547649 | -0.1096 | 1532 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000548400 | -0.1208 | 1024 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000548293 | -0.1151 | 1139 | GBM | - | | ENSG00000278864 | ENST00000624039 | MYL6 | ENSG00000092841 | ENST00000546845 | -0.1185 | 1227 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFN1 | ENSG00000108518 | ENST00000225655 | -0.1133 | 1287 | GBM | - | | ENSG00000278864 | ENST00000624039 | PFN1 | ENSG00000108518 | ENST00000574872 | -0.1133 | 1302 | GBM | - | | ENSG00000278864 | ENST00000624039 | ROMO1 | ENSG00000125995 | ENST00000374078 | -0.1603 | 1531 | GBM | - | | ENSG00000278864 | ENST00000624039 | ROMO1 | ENSG00000125995 | ENST00000374077 | -0.1603 | 1531 | GBM | - | | ENSG00000278864 | ENST00000624039 | ROMO1 | ENSG00000125995 | ENST00000374072 | -0.1770 | 251 | GBM | - | | ENSG00000278864 | ENST00000624039 | ROMO1 | ENSG00000125995 | ENST00000336695 | -0.1603 | 1531 | GBM | - | | ENSG00000278864 | ENST00000624039 | BRI3 | ENSG00000164713 | ENST00000539286 | -0.1150 | 984 | GBM | - | | ENSG00000278864 | ENST00000624039 | ECI1 | ENSG00000167969 | ENST00000562238 | -0.2069 | 455 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | ECI1 | ENSG00000167969 | ENST00000570258 | -0.3059 | 1411 | GBM | CHOL | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000341942 | -0.1433 | 1255 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000474141 | -0.1433 | 1255 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000460732 | -0.1038 | 1354 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000441173 | -0.1336 | 360 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000468582 | -0.1433 | 1246 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000488241 | -0.1038 | 1354 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000473459 | -0.1433 | 1255 | GBM | - | | ENSG00000278864 | ENST00000624039 | LAMTOR4 | ENSG00000188186 | ENST00000490633 | -0.1042 | 847 | GBM | - |
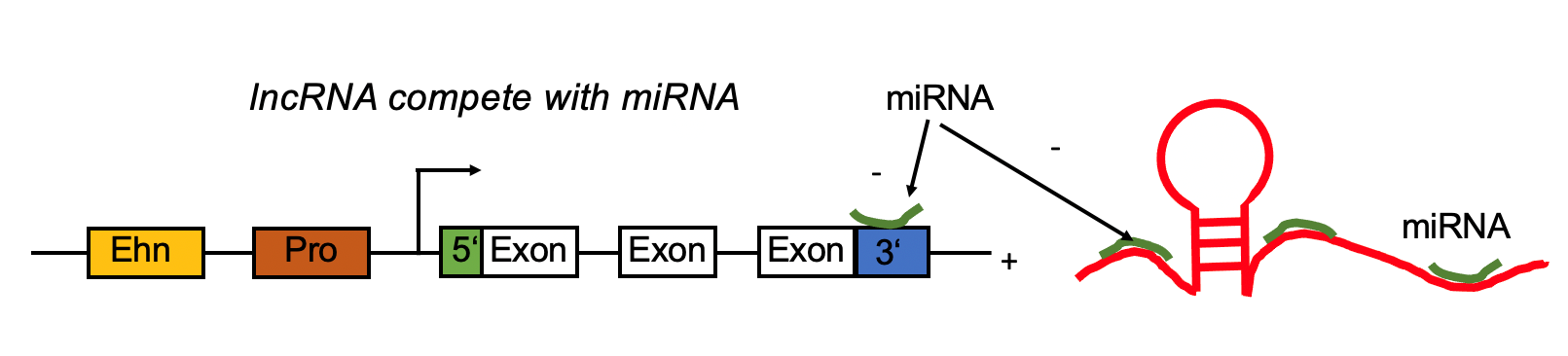
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000278864.1 | ENST00000624039.1 | 209 | 492 | 1121 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).