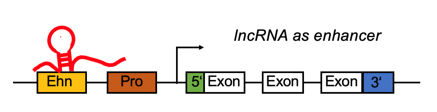
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | RPAP2,CCDC65,PMM2,GSDMB,B3GNTL1,VAMP1,EPOR,MFSD8,TMEM234,NBPF9,BRD1,FASTKD3,CIRBP,PET100,MZF1,TTC31,CNOT9,QPCTL,LYSMD4,SRP19,SUGP2,PPP2R3B,FKRP,DCLRE1C,MKNK1,L3HYPDH,DMTF1,RNF34,ZDHHC23,NUP62,FAM193A,UNKL,OPA3,CXXC5,PIGG,LMBR1L,ZNF24,NUP50,WDCP,ULK3,SERTAD3,KMT2B,IRAK4,STAT2,METTL5,ENY2,VMP1,ZFP14,CENATAC,NPEPL1,WDR5B,TP53INP1,SAFB2,KCTD13,MSANTD2,CCDC14,FAM200A,OSER1,ACAD8,CLK4,C6orf136,POLR2K,MAEA,ZUP1,FAM133B,ZBED5,EIF2B5,PPIH,RABGGTB,UBA2,RECQL4,TTC14,KNTC1,CEP72,PAF1,SLC2A11,HMGXB3,SUDS3,GKAP1,SPRN,SPG7,MIDN,PTPRH,ATR,SIAH1,PPP1R13L,USP49,DNAJC2,PPIP5K1,OGT,RCCD1,IGIP,CBX3,RAB4B,PDE7A,COX19,CLK2,MYEF2,BRSK1,TARBP2,RAPGEF6,MPLKIP,POLG2,DHX34,RBM15,CHKB,CEP152,CEP89,METTL17,UBE3A,ANAPC10,RPAP3,ESRP1,BBC3,CBY1,MORC2,SLC12A9,WDR83,OSGEP,THOC2,CLK1,ZBTB6,ATXN7,MRRF,WDR5,ITGB3BP,GABPB1,RABL2A,NASP,LLGL2,ANKRD39,PRC1,OFD1,MOV10,CARS2,ACYP1,CCDC9B,TOMM40,MTFMT,SLC25A29,TMEM39A,FCHSD1,TMEM243,IQGAP3,HELLS,TIAM2,CFAP298,GAR1,XPNPEP3,B3GALT6,SKP2,RSRC2,TTF1,C19orf48,PFDN6,HSH2D,FANCF,NAPB,C3orf62,NAA25,RDH14,LPCAT4,UTP23,ZFP82,SETD5,ZFP62,ALKBH2,ANAPC5,ZBED8,ORMDL1,ZNF45,BRAF,RALGAPA2,LPAR2,COQ7,C18orf21,COQ6,BAZ1A,C12orf73,DDX51,DONSON,AGER,BRIX1,MBD4,AGO3,C19orf12,SPOUT1,GUF1,SBNO1,SRSF2,C1orf52,PHC3,CUL9,CDK11A,TFCP2,MRPL32,PASK,TSEN54,LRRC37B,DHX35,ATG4B,SKIL,UBE2O,LMNTD2,TAOK3,DCUN1D2,NXF1,DAGLB,EIF2B1,FIGNL1,NR2C2,PAN2,TEAD2,BORCS8,BMF,FAM229A,SLFN13,SAE1,GIGYF1,ZNF14,NOP2,FAM193B,CENPH,LSM11,ABCA7,TMCO6,NOL3,XPO1,COG7,SEC31B,ZFC3H1,PDRG1,KLHL17,NFYB,ARHGAP33,MBTD1,DMAC2L,DNASE1,TMEM184A,HEXD,TNPO2,HDAC8,VCPKMT,TMEM50B,TTC3,CEP192,SCNN1D,EIF4A1,CCDC24,CSNK2A2,MRPS25,CDK7,CCDC12,BFAR,VPS13A,POLM,PALB2,DECR2,SHROOM1,TRIM27,CCDC34,INSR,ATF7IP,TMEM248,CCDC154,ERCC5,MARCHF6,RNF214,RSL24D1,IQCC,ZNF30,TMEM168,SCNM1,ATM,FLVCR1,TEDC2,FANCL,ILF3,NUP54,GLI4,TTC17,CAPN12,BRAP,CDK13,EMG1,DPY19L3,SMG7,LENG8,MED7,RBBP8NL,SH2B1,LCORL,PHKA2,C1orf159,FAM98C,UBE2G2,DNMBP,IP6K2,ARIH2,SPICE1,SH3D21,WDR53,PIF1,PPWD1,RELCH,MLLT6,SASS6,RBM27,ATG12,ZNF70,FASTKD1,INO80D,ICE2,AGK,ASTE1,TRIM65,EIF2B4,TMED4,TIMM44,C20orf96,DDX55,MIX23,HPF1,CASP2,KRIT1,AP1G2,RPL13A,GLMN,FAM111A,BANP,MICA,THOC1,CCDC40,ANKMY1,WDR59,CCDC59,UBA6,AIMP1,NUTF2,PHF12,BEND3,ACBD6,EZH2,TAMM41,MMP15,SLC4A3,GDPD3,DRC3,RC3H1,CBLB,TOP3A,TMEM138,SMG1,ARHGEF38,FAM71E1,LYSMD3,FAAP24,RIMKLB,LTO1,GRPEL2,KANSL2,ZSWIM3,PIDD1,METAP1D,BCLAF3,CWC15,PUS7L,CCDC106,CCDC137,GGCT,LAS1L,CEP83,RFC4,NAA10,UBN2,ANKDD1A,EXD3,DDX6,NFKBID,RTF1,LRIG2,PARP2,ABHD18,PCED1A,ABTB1,NRBP2,MAPK15,SLX4,C11orf98,TBCB,ANGEL1,SMYD4,MTA3,IKBKB,NBPF11,DYNLT2,BRF1,ACIN1,ZNF41,ZC3H8,GNL2,ARHGAP17,TNRC18,PRR7,RAB17,ENOSF1,MAU2,DZIP3,RAB2B,UBOX5,ANKRD49,DNAJC19,KNOP1,SLC38A9,PGBD1,LENG1,MEX3A,FNTA,FNBP4,ZNF28,ZFYVE19,LRRC56,INPP5E,KMT2D,RSRP1,PTRH2,LRR1,NEK8,CHD2,SERF2,NVL,ATG14,GOLGB1,CLEC2D,PWWP2A,ZSWIM1,NBPF15,TRPT1,TATDN2,MYNN,METTL14,KIFC2,TSNARE1,ANKLE2,NSUN2,ZNF720,MED19,HEMK1,MIER3,UPF3B,LINS1,RABEP2,PCSK4,ANKRD26,SCAF11,PYURF,HAUS1,CLHC1,DDX11,PUS10,FRS3,NDUFAF5,GPBP1,C2orf68,FBXL8,MRE11,GNRH1,AP4M1,PSMG2,GK5,ZNRF1,RBM6,FRG1,TOR1AIP2,EHMT1,TRMT13,KRTCAP2,ZNF862,SREK1IP1,ARHGAP12,PRMT1,POP5,ORAI3,TMEM91,PAXIP1,MYL5,VCPIP1,NOM1,EPM2AIP1,TYW1,ABCB6,SYNGAP1,MLLT1,CHD1,PSME3IP1,NAP1L4,MAML1,WDR27,HECTD4,PTCD1,PPCDC,HARS2,KHDC4,CROCC,CERS5,PDCD2L,EP400P1,HPS4,SLC25A19,PNISR,RP9,SFPQ,PLPP5,GEN1,TMEM63A,TRIOBP,FKBP11,CARD8,NCBP2,ATAD2B,DUSP18,ZNF10,LRRC14,KCNC3,TBRG4,LRIF1,DUT,RNF149,PYM1,PLCG1,ALG2,ESCO1,ZBTB8OS,RASSF1,TRMU,TMX3,NAGK,MYPOP,CTBP1,BICRA,NFKBIB,ATXN7L2,PSMA2,SEM1,POC5,IQCE,CHTF18,ADPGK,YTHDF1,AKAP8,CDPF1,CEP250,ARID2,MSTO1,UIMC1,DPY19L2,UBE2J2,C12orf29,RNF44,TAF1D,KATNBL1,MKS1,TAF4,PLEKHH1,C1orf35,CCDC174,TTI1,CDK5RAP1,PIP4P1,SPIN3,NDUFA7,BUD23,TBRG1,PSMG1,SLX4IP,NPRL2,ZCWPW1,RDH13,KLHDC4,EFNA4,MIS18A,CCNL2,TET3,EFCAB7,SMIM22,FBXO46,DMPK,PRR14L,RNPC3,CLK3,C14orf28,ANKRD10,TENT4A,INCA1,UQCRB,LRCH3,UBXN7,NUSAP1,CNOT4,CCDC77,TFAP4,N6AMT1,MSL3,LSM5,SLC66A1,DDTL,BCAT2,TP53I13,SCAF4,MC1R,COQ4,VILL,TDP1,CCNT2,LLPH,KAT2A,PHTF2,NFRKB,ZNF91,CHD6,RAD1,AGO2,SLC17A9,U2AF2,CSTF3,DDI2,XRCC2,PLA2G6,CSNK1E,TMEM86B,RNPS1,NR2F6,ZBTB3,TUBGCP5,RBM25,ENTR1,CYBC1,DCP1B,FBXW7,SLC27A3,CEBPZOS,RHPN1,SPATA2,ENDOV,VN1R1,ZFP30,FLAD1,BLCAP,MRPL50,PPIE,CLDN15,SETD6,POLI,PRPF39,ETNK1,ENTPD4,TRIM45,PRR36,TMEM41B,GEMIN6,RNF114,ZNF22,PHF1,PABIR2,ABHD11,TMC4,HDAC10,FBXL20,TGIF2,CEP44,CENPC,RBM41,HSF4,PROSER3,NOL9,YJEFN3,NEPRO,NOXA1,POLR2D,CCNL1,TLK1,RGL3,ANKRD54,MEX3D,ANKRA2,VPS33A,VMAC,KRI1,CELF1,BCL2L12,TMEM60,TMEM161B,KRR1,GPR35,RBM45,MICALL2,HSPBAP1,CRYGS,NFXL1,YAE1,DYRK4,R3HDM2,LBHD1,RPS6,FAM160A2,NSMAF,EGFL8,NR2C2AP,PSMG4,DDX31,WDR36,PBX4,TIA1,ALDH16A1,HOOK2,ZNF83,RANBP10,PPP4R3A,MOSMO,SAAL1,POLR1G,RETREG3,KANSL1,ZNF3,MCRIP2,ZNF8,LETM1,ABCD4,PTPN2,WDR77,CDK2,BTG3,VSIG10,TMEM68,SAYSD1,C19orf25,NEIL1,SZT2,POLE,EIF5B,ABHD10,DTX3,KPTN,MTG2,E2F1,RNF215,ZCCHC10,NAT9,ELF2,CRAMP1,LUC7L,ZNF43,PTBP3,PNN,DSN1,LUC7L2,BRD8,UBE2V1,PPAN,CHAF1A,MROH1,CENPJ,C14orf93,DPH7,C4orf47,TRA2A,MAPKAPK5,ZMYND8,POLR3F,NEURL4,HINFP,PCBD2,NR2C1,BRD9,AMY2B,CYTH2,C8orf76,RCC2,B9D2,NOP53,WBP1,RBM33,RAD17,AVL9,NOL8,TONSL,NADSYN1,MOK,IMMP1L,CFAP44,SRRT,AGAP6,TPRA1,CHEK2,GABPB2,ZNF16,PIGA,GEMIN8,MPHOSPH10,GTPBP2,TRIM66,RBM39,EDC3,C15orf61,SLC26A6,SLC25A27,MRPS26,MSL2,IRF3,TMEM161A,ZCCHC8,NRF1,TMEM44,ITFG2,ZNF2,RRNAD1,ZXDC,ESF1,SNRPD2,KAT8,PDS5A,PIGL,PPP5C,KCTD6,ERH,RNF10,CSAD,LRRC37A3,KIAA0895L,RINL,DDX39A,ARV1,NDUFAF6,ZNF77,ATAD3B,HAUS8,SAMD10,MUTYH,RBM28,ZNF74,BANF1,DCAF8,BRI3BP,C2CD5,STAG3L5P-PVRIG2P-PILRB,LAMA5,HJURP,PHYKPL,POFUT2,CACTIN,RUFY3,OGA,ZFP69,NUP42,IFT27,PHKG2,NARF,LARP7,ZC3HC1,PIGM,PPP1R16A,COPS5,SS18L2,SWSAP1,DCAF16,ZC3H4,RICTOR,SNX16,TM2D3,CENPT,OSGEPL1,CHORDC1,ZGLP1,CRLS1,TRIM3,DPP8,MTG1,URI1,WDR55,FAM76B,POGZ,GTPBP3,ANAPC7,USP7,BRWD3,RBM4B,DTWD1,PCNT,C21orf58,TAF1C,UFD1,ADNP,ADAT2,NCOA5,GABRE,MYO19,SURF6,C12orf43,PRIMPOL,RAPGEFL1,UBL5,RPS11,PRDM4,CNKSR1,THAP9,SYNE4,MRNIP,ZNF71,THAP6,SS18L1,ELMOD3,DDX17,TRPM4,PUS7,ING4,SLC7A6,HAUS5,AQR,RSBN1L,RAD52,CEP43,HDAC7,ARVCF,THYN1,LIAS,DYRK1A,NINL,CTDSPL2,TMEM165,INO80,FBXL12,CEP290,SDR39U1,EIF2AK1,ZMYND19,PGGHG,MCM8,TELO2,CDAN1,FBXL6,TEDC1,TRAIP,PCYOX1L,ATAD5,C12orf76,ANO9,WASHC1,ABCC5,TIGD1,KLC1,GPS2,TRIM52,RPL36A,COA8,PCBP4,THAP3,NOL12,SLC22A5,NSA2,CLASRP,CCDC93,MBD6,STAC3,CPT1B,TTC32,MICAL1,MTX3,SIPA1L3,ZMAT1,GOLGA8B,MCM3AP,SPATA33,ABHD3,RPS24,STRADA,SMG9,ZNF18,TIMM29,NKRF,DIDO1,GOLGA8A,CYP2R1,PAM16,PPHLN1,SETD4,SPIRE2,CNOT2,C1orf109,TPRKB,RBM48,PSMC3IP,ASNS,AARSD1,GMEB2,DUSP28,PUM2,NACA,RBM5,USP3,ANP32A,NKTR,SLC35A2,SAP30BP,HPN,ZFP28,BCDIN3D,RAD18,TTC9C,ZCCHC7,C19orf71,ZNF17,SRCAP,DOT1L,CRBN,TTLL3,ZNF84,TOX3,RXRB,PARP6,RHBDD3,GDPGP1,CACNB3,C19orf18,MED10,PDCD6,NDUFV3,UTP15,REX1BD,PLEKHN1,PIAS2,LRP5L,KATNIP,TIPIN,ASXL1,QRICH2,TSC1,TEP1,SIRT6,NAE1,COLCA2,MACROH2A1,MAN2C1,DXO,HSPB11,PDCD7,CCDC112,USP20,DPP9,ANKRD13D,WHAMM,LIG1,NELFA,RABL2B,TRAPPC2,SIRT7,SNRPB2,MRFAP1L1 | | Exist in public source | | NA |
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | DPH7,C14orf93,MICA,ATF7IP,DNASE1,PDCD2L,POC5,C21orf58,DGCR8,OPA3,TNRC18,ULK1,TRA2A,METTL5,STX16,HEMK1,ANAPC10,PDIK1L,ZFP14,TBCC,WDR55,PARP2,PDCD6,STAT2,RGL3,SETD6,PIGM,UBXN7,TATDN2,DAGLB,HIP1R,CCDC66,VMAC,MZF1,TRMT10C,C3orf62,ANKRD54,FAM160B2,ZSWIM1,LRRC57,C2CD5,MYL5,FBXO46,POLR2K,TMEM44,STRADA,PPCDC,SPICE1,NINL,SLC38A9,MOB1B,CDK13,FAM111A,MIDN,PTCD1,ARL14EP,DCAF8,PCNT,YDJC,ITGB3BP,SEH1L,ZNF18,CBFA2T2,DMPK,PFDN4,FBXL12,HSH2D,VARS2,CDK2,EP400NL,UPF3B,KLHDC4,DYRK4,SPDL1,ANP32B,ZNF14,LBHD1,INSR,FBXL20,FAM133B,SRP14,CLASRP,RRP7BP,SDR39U1,TMEM161A,DCP1B,RAD9A,NOL3,FAM122B,MYB,QTRT1,TLK1,ACCS,ENGASE,C2orf68,XPNPEP3,MRPL42,SCNM1,MLLT1,SMG9,THAP6,CYTH2,COMMD3,UBA2,ING5,AIMP1,AARSD1,PCBD2,C8orf76,COG7,ZMYND19,MAP3K1,ATAD5,ZBTB6,MCM3AP,MDM4,TTC32,CCDC174,MSTO1,ZMAT1,LYSMD4,WDR60,HAUS5,KAT2A,RAD52,RPAP3,FASTKD1,EID2B,SMG1,FAM32A,NFKBIB,BAX,TRMU,DDX17,MSL2,IQCE,ZNF720,ZNRF1,ZNF83,GLTSCR2,EVL,STAC3,PRPF3,PIK3C3,CEP89,PRR7,DCLRE1C,SHFM1,THAP3,LCORL,LUC7L,ZFP82,DOT1L,SIN3B,RNF149,BANF1,FAM71E1,YEATS2,SAYSD1,U2AF2,ZNF45,EIF5B,ABHD18,ANKRD26,MAVS,PBX4,RAD17,CCDC130,NELFCD,ABCD4,C21orf59,MFSD8,SPATA2,TRIOBP,NR2C2AP,CCNL1,LRRC37A3,CAMSAP3,CPSF7,TMEM248,SENP5,SLX4,RBM6,TMEM50B,PDCD7,KIAA0556,CDAN1,CDK5RAP1,ANKRD39,GABPB1,GTF2H4,TEAD2,ARV1,ZCCHC10,ABHD11,OSGEPL1,HAUS1,EID2,ZNF3,CBFB,ZNF70,OXLD1,ZNF30,LPAR2,SPATA25,METTL17,TIA1,RBM5,PUS7,CEP290,BCAT2,NME3,C9orf114,ING4,BBC3,COQ6,MYNN,TPRA1,MPLKIP,ZUFSP,B4GALT7,CELF1,ENOSF1,NADSYN1,POLR2D,XRCC2,C14orf79,TMCO6,MPHOSPH10,GEMIN8,CENPJ,DPP8,MLEC,RPSA,TMEM120B,RALGPS1,KIAA0907,SRSF5,PRR14,TRAF5,TRIM27,PSMA2,MAU2,SETD4,PUS10,H2AFX,CCDC137,CCDC106,SUZ12,C19orf18,TEP1,KPTN,COQ7,HSF4,TBRG1,NUPL2,TFCP2,RC3H1,TMX3,WDR59,ODF2,CIRBP | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
| Only predicted by TDF | | DCTN1,TPP1,TINF2,CRTAP,MYH9,RNF19B,CYB5R3,TUBA1B,TSKU,DLST,HK1,GRHPR,MTCH2,ALDH2,MGST1,CD14 | | Exist in public source | | NA |
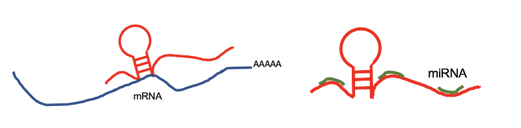
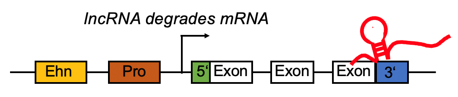
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | ACAA1,AKR7A2,TUBA1B,DLST,PSMD2,EIF3L,RAB5C,SMPD1,THTPA,ATP6V0D1,RAB31,MYH9,TANGO6,FTH1,TOLLIP,COMT,C1R,PC,PCDHGC3,METRNL,TALDO1,CYB5R3,WDR1,TINF2,EIF4G1,ALDH2,OS9,PEX19,NOMO2,MSRA | | Exist in public source | | NA |
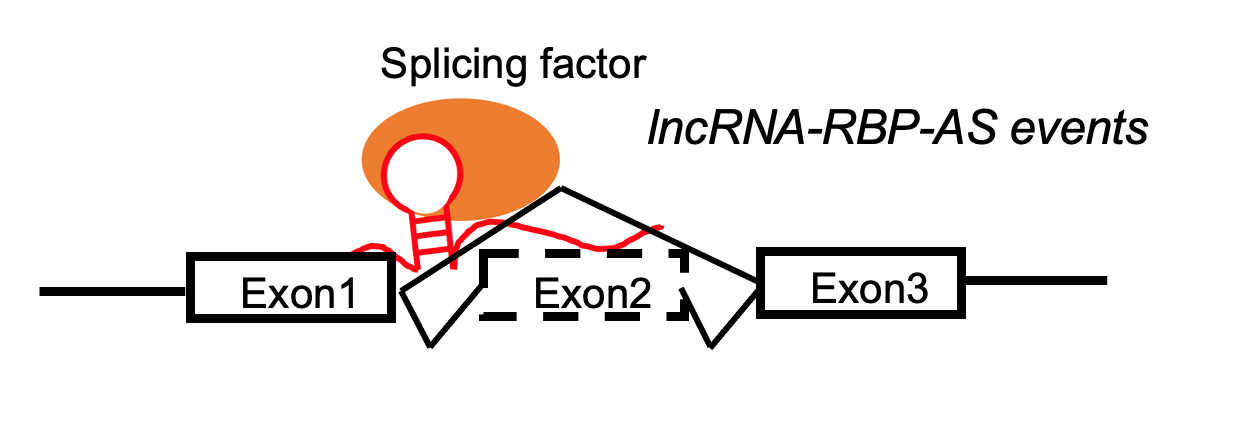
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000269867 | ENST00000597780 | exon_skip_386274 | chr3:108049618-108049651 | -5.03 | -0.2286 | CD47 | ENST00000361309 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_470077 | chr7:102397619-102397698 | -9.78 | -0.1397 | PRKRIP1 | ENST00000397912,ENST00000496391 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_423795 | chr4:55888887-55888932 | -7.59 | -0.2372 | EXOC1 | ENST00000346134,ENST00000381295 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_44998 | chr10:110132304-110132400 | -9.72 | -0.1800 | ADD3 | ENST00000356080 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_430601 | chr4:82842139-82842481 | -23.80 | -0.1221 | SEC31A | ENST00000355196,ENST00000395310,ENST00000448323 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000269867 | ENST00000597780 | exon_skip_377198 | chr3:123130479-123130616 | -14.79 | -0.1112 | PDIA5 | ENST00000316218 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_11029 | chr1:154989559-154989707 | -17.54 | -0.1319 | FLAD1 | ENST00000292180 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_309204 | chr19:49181348-49181461 | -18.92 | -0.1971 | TRPM4 | ENST00000252826 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_382458 | chr3:38131595-38131638 | -9.34 | -0.2278 | ACAA1 | ENST00000333167 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_7523 | chr1:78660553-78660654 | -9.99 | -0.1009 | IFI44 | ENST00000370747 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_9446 | chr1:147270931-147271005 | -9.15 | -0.1551 | CHD1L | ENST00000369258 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_16568 | chr1:201989382-201989531 | -20.13 | -0.1491 | RNPEP | ENST00000295640 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_41813 | chr10:68337857-68337996 | -17.64 | -0.1819 | HNRNPH3 | ENST00000265866 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_53559 | chr10:101793912-101793998 | -13.13 | -0.3202 | OGA | ENST00000361464 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_59071 | chr11:47234756-47234934 | -16.59 | -0.1327 | DDB2 | ENST00000256996 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_66482 | chr11:126274926-126275021 | -10.54 | -0.1285 | FOXRED1 | ENST00000263578 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_87733 | chr12:120571190-120571291 | -11.89 | -0.1887 | RNF10 | ENST00000325954 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_92074 | chr12:50142052-50142110 | -7.36 | -0.1363 | CERS5 | ENST00000317551 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_95407 | chr12:95040475-95040597 | -12.46 | -0.1246 | NR2C1 | ENST00000333003 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_108095 | chr14:73955450-73955509 | -9.33 | -0.1794 | COQ6 | ENST00000334571 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_114365 | chr14:74290026-74290118 | -18.64 | -0.2273 | ABCD4 | ENST00000356924 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_114376 | chr14:74293153-74293248 | -12.30 | -0.1618 | ABCD4 | ENST00000356924 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_128982 | chr15:72250844-72250954 | -17.17 | -0.2385 | PARP6 | ENST00000287196,ENST00000569795 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_133217 | chr16:8635038-8635079 | -9.34 | -0.3592 | METTL22 | ENST00000381920 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_139422 | chr16:89114338-89114487 | -19.70 | -0.1743 | ACSF3 | ENST00000317447,ENST00000406948 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_290158 | chr17:42953723-42953778 | -6.96 | -0.1450 | AARSD1 | ENST00000427569 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_300663 | chr19:1425877-1425960 | -11.74 | -0.1631 | DAZAP1 | ENST00000233078 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_302168 | chr19:10680345-10680443 | -9.85 | -0.1059 | ILF3 | ENST00000590261 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_319223 | chr19:43526200-43526349 | -17.38 | -0.1287 | ETHE1 | ENST00000292147 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_327433 | chr2:86125794-86125880 | -6.98 | -0.1091 | PTCD3 | ENST00000254630 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_353131 | chr20:58669290-58669453 | -23.38 | -0.1569 | STX16 | ENST00000371141 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_361485 | chr21:29008239-29008307 | -9.18 | -0.2700 | RWDD2B | ENST00000493196 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_370518 | chr22:50247691-50247776 | -15.34 | -0.1871 | HDAC10 | ENST00000216271 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_374497 | chr3:49999439-49999513 | -10.27 | -0.1902 | RBM6 | ENST00000266022 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_380242 | chr3:186784961-186785101 | -12.97 | -0.1365 | EIF4A2 | ENST00000323963 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_438188 | chr5:141638954-141639021 | -16.15 | -0.2991 | RELL2 | ENST00000297164,ENST00000444782 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_444499 | chr5:138164086-138164133 | -5.11 | -0.2044 | BRD8 | ENST00000254900 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_445390 | chr5:150403120-150403312 | -26.10 | -0.1684 | CD74 | ENST00000009530 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_374581 | chr3:50110378-50110463 | -12.70 | -0.1789 | RBM5 | ENST00000347869 | Frame-shift | | ENSG00000269867 | ENST00000597780 | exon_skip_75223 | chr11:72012400-72012442 | -6.74 | -0.2592 | NUMA1 | ENST00000393695 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_96112 | chr12:109945259-109945349 | -12.43 | -0.1535 | GIT2 | ENST00000355312 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_376144 | chr3:100311832-100311877 | -7.87 | -0.3279 | TBC1D23 | ENST00000394144 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_382356 | chr3:37091466-37091538 | -4.99 | -0.1085 | LRRFIP2 | ENST00000336686 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_470719 | chr7:117134123-117134192 | -7.32 | -0.1284 | ST7 | ENST00000265437 | In-frame | | ENSG00000269867 | ENST00000597780 | exon_skip_104174 | chr13:79353152-79353224 | -7.65 | -0.1159 | RBM26 | ENST00000438737 | In-frame |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000269867 | hsa-mir-152 | ENST00000597780 | chr19:57867167-57867253 | 192.00 | -82.20 | 88 | NA | | ENSG00000269867 | hsa-mir-152 | ENST00000597780 | chr19:57867400-57867482 | 154.00 | -76.08 | 87 | NA | | ENSG00000269867 | hsa-mir-22 | ENST00000597780 | chr19:57867111-57867185 | 228.00 | -86.59 | 86 | NA | | ENSG00000269867 | hsa-mir-22 | ENST00000597780 | chr19:57867193-57867276 | 178.00 | -79.51 | 81 | NA | | ENSG00000269867 | hsa-mir-22 | ENST00000597780 | chr19:57867387-57867476 | 167.00 | -67.52 | 76 | NA | | ENSG00000269867 | hsa-mir-22 | ENST00000597780 | chr19:57867468-57867545 | 165.00 | -66.77 | 83 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000269867 | ENSG00000004777 | ARHGAP33 | ACC,CESC,CHOL,COAD,DLBC,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUSC,MESO,PAAD,PCPG,PRAD,THCA,THYM,UCS | KIRC | | | ENSG00000269867 | ENSG00000104859 | CLASRP | ACC,BLCA,CHOL,DLBC,ESCA,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,OV,PAAD,PCPG,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | KIRC | | | ENSG00000269867 | ENSG00000104980 | TIMM44 | ACC,CHOL,DLBC,GBM,KICH,KIRC,LGG,LIHC,MESO,PAAD,PCPG,PRAD,THCA,UVM | KICH | | | ENSG00000269867 | ENSG00000105486 | LIG1 | ACC,CHOL,DLBC,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUSC,OV,PAAD,PCPG,PRAD,TGCT,THCA | KIRC | | | ENSG00000269867 | ENSG00000105771 | SMG9 | ACC,BLCA,CHOL,ESCA,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUSC,PAAD,PCPG,PRAD,TGCT,THCA | KICH | | | ENSG00000269867 | ENSG00000115257 | PCSK4 | ACC,BRCA,CHOL,KIRC,KIRP,PAAD,PCPG,THCA,THYM | BRCA | | | ENSG00000269867 | ENSG00000117616 | RSRP1 | ACC,BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,READ,SARC,SKCM,TGCT,THCA,THYM,UCEC,UCS,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000118162 | KPTN | ACC,BLCA,CHOL,GBM,HNSC,LGG,LIHC,LUSC,MESO,PCPG,PRAD,SARC,TGCT,THCA | BLCA | | | ENSG00000269867 | ENSG00000119559 | C19orf25 | ACC,CHOL,HNSC,KICH,LGG,LIHC,MESO,PAAD,PCPG,THCA | KICH | | | ENSG00000269867 | ENSG00000127586 | CHTF18 | ACC,BLCA,CHOL,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,PAAD,PCPG,PRAD,SARC,THCA,UCS | BLCA,KIRC | | | ENSG00000269867 | ENSG00000130299 | GTPBP3 | ACC,BLCA,CHOL,DLBC,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,PAAD,PRAD,THCA,UVM | BLCA | | | ENSG00000269867 | ENSG00000130810 | PPAN | ACC,CHOL,GBM,KICH,LGG,PRAD,READ,TGCT,THCA | KICH | | | ENSG00000269867 | ENSG00000131398 | KCNC3 | ACC,BLCA,CHOL,DLBC,OV,PCPG,THCA,UCS | BLCA | | | ENSG00000269867 | ENSG00000139631 | CSAD | ACC,BLCA,BRCA,CESC,COAD,GBM,HNSC,KICH,KIRC,KIRP,LGG,LUAD,LUSC,OV,PAAD,PCPG,PRAD,SKCM,STAD,THCA,THYM,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000142002 | DPP9 | ACC,GBM,KIRC,KIRP,LGG,LIHC,PAAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000142530 | FAM71E1 | ACC,ESCA,KICH,LIHC,MESO,PCPG,STAD,UCS | KICH | | | ENSG00000269867 | ENSG00000148824 | MTG1 | ACC,BLCA,CHOL,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,PAAD,SKCM,STAD,THCA,UCS,UVM | BLCA | | | ENSG00000269867 | ENSG00000160439 | RDH13 | ACC,BRCA,CESC,CHOL,DLBC,ESCA,KICH,KIRP,LIHC,LUAD,LUSC,OV,PAAD,PCPG,PRAD,READ,SKCM,STAD,TGCT,THCA,UCS,UVM | KICH | | | ENSG00000269867 | ENSG00000161010 | MRNIP | ACC,BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,KIRC,KIRP,LGG,OV,PAAD,SKCM,TGCT,THCA,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000162572 | SCNN1D | ACC,GBM,KIRC,KIRP,LGG,PAAD,PCPG,PRAD,THYM | KIRC | | | ENSG00000269867 | ENSG00000164403 | SHROOM1 | ACC,BLCA,CESC,ESCA,HNSC,KIRC,PAAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000166619 | BLCAP | ACC,BLCA,CHOL,KICH,LUSC,PAAD,READ,THCA | KICH | | | ENSG00000269867 | ENSG00000167615 | LENG8 | ACC,BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,OV,PAAD,PCPG,PRAD,SARC,SKCM,THCA,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000167670 | CHAF1A | ACC,CHOL,GBM,KICH,LGG,LIHC | KICH | | | ENSG00000269867 | ENSG00000167747 | C19orf48 | ACC,CHOL,GBM,HNSC,KICH,KIRC,LGG,LIHC,LUSC,MESO,OV,PRAD,SARC,TGCT,UCS | KICH | | | ENSG00000269867 | ENSG00000175265 | GOLGA8A | ACC,BLCA,CESC,COAD,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LUAD,LUSC,PAAD,PCPG,PRAD,SARC,SKCM,TGCT,THCA,THYM,UCEC,UCS,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000175309 | PHYKPL | ACC,BLCA,CESC,CHOL,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,PAAD,SKCM,THCA,THYM,UCEC,UVM | KIRC | | | ENSG00000269867 | ENSG00000177990 | DPY19L2 | ACC,KICH,KIRC,KIRP,PCPG,TGCT,THCA,UCS | KICH | | | ENSG00000269867 | ENSG00000184465 | WDR27 | ACC,BLCA,CESC,CHOL,COAD,DLBC,GBM,KIRC,KIRP,LGG,PAAD,READ,THCA,THYM,UCEC,UCS | KIRC | | | ENSG00000269867 | ENSG00000204149 | AGAP6 | ACC,BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,OV,PAAD,READ,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000204305 | AGER | ACC,BLCA,CESC,CHOL,GBM,HNSC,KIRC,KIRP,LGG,LIHC,PAAD,SARC,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000205129 | C4orf47 | ACC,KIRC,KIRP,TGCT,THCA | KIRC | | | ENSG00000269867 | ENSG00000215375 | MYL5 | ACC,BLCA,CHOL,KIRC,KIRP,LGG,LUAD,PCPG,PRAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000221978 | CCNL2 | ACC,BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,OV,PAAD,PCPG,PRAD,READ,SARC,SKCM,TGCT,THCA,THYM,UCEC,UCS,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000250067 | YJEFN3 | ACC,BLCA,ESCA,GBM,KIRC,KIRP,LAML,LGG,LUSC,MESO,OV,PAAD,PCPG,PRAD,TGCT,THYM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000013441 | CLK1 | BLCA,CESC,COAD,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,OV,PAAD,PCPG,PRAD,READ,SKCM,THCA,THYM,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000044446 | PHKA2 | BLCA,CHOL,ESCA,GBM,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,PRAD,THCA,THYM | KIRC | | | ENSG00000269867 | ENSG00000073605 | GSDMB | BLCA,CHOL,GBM,KIRC,KIRP,LGG,LIHC,LUSC,PAAD,SARC,THCA,THYM,UCS | KIRC | | | ENSG00000269867 | ENSG00000100068 | LRP5L | BLCA,CHOL,DLBC,ESCA,KICH,KIRP,LUAD,LUSC,PAAD,PRAD,TGCT,THCA,THYM,UVM | KICH | | | ENSG00000269867 | ENSG00000100288 | CHKB | BLCA,CHOL,COAD,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,READ,SKCM,STAD,TGCT,THCA,THYM,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000100429 | HDAC10 | BLCA,CHOL,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,PAAD,PCPG,PRAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000102287 | GABRE | BLCA,CHOL,COAD,HNSC,KIRC,LIHC,PAAD,THCA,UCS | KIRC | | | ENSG00000269867 | ENSG00000102996 | MMP15 | BLCA,ESCA,KICH,PAAD,UCS | KICH | | | ENSG00000269867 | ENSG00000103460 | TOX3 | BLCA,CHOL,ESCA,PAAD,UCS | BLCA | | | ENSG00000269867 | ENSG00000108352 | RAPGEFL1 | BLCA,KIRC,LIHC,PAAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000108773 | KAT2A | BLCA,CHOL,DLBC,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,PAAD,PCPG,PRAD,SARC,SKCM,TGCT,THCA,UCEC,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000113240 | CLK4 | BLCA,COAD,DLBC,GBM,HNSC,KIRC,KIRP,LGG,READ,SKCM,THCA,THYM,UCEC,UVM | KIRC | | | ENSG00000269867 | ENSG00000124839 | RAB17 | BLCA,CESC,ESCA,PRAD,READ,UCS | BLCA | | | ENSG00000269867 | ENSG00000130590 | SAMD10 | BLCA,CHOL,GBM,KIRC,KIRP,LIHC,PAAD,PCPG,PRAD,THCA | BLCA | | | ENSG00000269867 | ENSG00000130701 | RBBP8NL | BLCA,CHOL,KICH,PRAD,UCS | BLCA,KICH | | | ENSG00000269867 | ENSG00000132522 | GPS2 | BLCA,CESC,CHOL,DLBC,GBM,KIRC,KIRP,LGG,LUAD,LUSC,MESO,PAAD,PCPG,PRAD,SKCM,STAD,THCA,THYM,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000132680 | KHDC4 | BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,READ,SKCM,STAD,THCA,UCEC,UCS,UVM | BLCA | | | ENSG00000269867 | ENSG00000135722 | FBXL8 | BLCA,CHOL,HNSC,KIRC,KIRP,LUAD,LUSC,PAAD,PCPG,PRAD,THCA | BLCA,KIRC | | | ENSG00000269867 | ENSG00000139546 | TARBP2 | BLCA,CHOL,HNSC,KIRP,LGG,LIHC,MESO,PAAD,PRAD,THCA,UCS | BLCA | | | ENSG00000269867 | ENSG00000139636 | LMBR1L | BLCA,CHOL,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,TGCT,THCA,THYM,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000142675 | CNKSR1 | BLCA,CHOL,LUSC,PAAD,PRAD,THCA,UCS | BLCA | | | ENSG00000269867 | ENSG00000146083 | RNF44 | BLCA,CHOL,KICH,KIRC,LIHC | KICH | | | ENSG00000269867 | ENSG00000158106 | RHPN1 | BLCA,CHOL,HNSC,KIRC,KIRP,LGG,LIHC,PAAD,THCA,THYM,UCS | BLCA | | | ENSG00000269867 | ENSG00000160072 | ATAD3B | BLCA,CHOL,DLBC,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,OV,PAAD,PCPG,PRAD,THCA,UVM | KIRC | | | ENSG00000269867 | ENSG00000160972 | PPP1R16A | BLCA,CHOL,HNSC,PAAD,THCA | BLCA | | | ENSG00000269867 | ENSG00000161328 | LRRC56 | BLCA,BRCA,CHOL,KIRP,THCA | BRCA | | | ENSG00000269867 | ENSG00000162650 | ATXN7L2 | BLCA,CHOL,DLBC,GBM,KIRC,KIRP,LGG,LUAD,LUSC,PCPG,SARC,THCA,THYM,UVM | BLCA | | | ENSG00000269867 | ENSG00000163257 | DCAF16 | BLCA,CHOL,COAD,DLBC,GBM,HNSC,KICH,KIRC,KIRP,LAML,LIHC,LUAD,READ,SKCM,UVM | KICH | | | ENSG00000269867 | ENSG00000164855 | TMEM184A | BLCA,CHOL,KICH,KIRC,KIRP,LIHC,PAAD,UCS | KICH | | | ENSG00000269867 | ENSG00000164938 | TP53INP1 | BLCA,KICH,KIRC,KIRP,TGCT,THYM,UVM | KICH | | | ENSG00000269867 | ENSG00000167535 | CACNB3 | BLCA,CHOL,KIRC,KIRP,PAAD,TGCT,THYM | BLCA | | | ENSG00000269867 | ENSG00000167604 | NFKBID | BLCA,CHOL,DLBC,GBM,HNSC,KIRC,KIRP,LUAD,PAAD,THCA,UCS | KIRC | | | ENSG00000269867 | ENSG00000167608 | TMC4 | BLCA,BRCA,CESC,CHOL,ESCA,LIHC,PAAD,PRAD,UCS | BRCA | | | ENSG00000269867 | ENSG00000167702 | KIFC2 | BLCA,CHOL,HNSC,KICH,KIRC,KIRP,LIHC,OV,PAAD,THCA,UVM | BLCA,KICH,KIRC | | | ENSG00000269867 | ENSG00000168781 | PPIP5K1 | BLCA,CHOL,KICH,KIRP,LUAD,LUSC,PAAD,PRAD | KICH | | | ENSG00000269867 | ENSG00000172366 | MCRIP2 | BLCA,ESCA,HNSC,KICH,PRAD | KICH | | | ENSG00000269867 | ENSG00000172890 | NADSYN1 | BLCA,GBM,KICH,KIRC,KIRP,LGG,PAAD,THCA,THYM | KICH,KIRC | | | ENSG00000269867 | ENSG00000174943 | KCTD13 | BLCA,CHOL,HNSC,KICH,KIRC,KIRP,LIHC,PAAD,PRAD,THCA,UVM | KIRC | | | ENSG00000269867 | ENSG00000175455 | CCDC14 | BLCA,CESC,CHOL,COAD,DLBC,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,OV,PAAD,PCPG,PRAD,READ,SKCM,TGCT,THCA,THYM,UCEC,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000177548 | RABEP2 | BLCA,CHOL,HNSC,KICH,KIRC,LGG,LIHC,LUSC,PAAD,PRAD,THCA | KICH | | | ENSG00000269867 | ENSG00000178188 | SH2B1 | BLCA,CESC,CHOL,DLBC,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,THCA,THYM | KICH | | | ENSG00000269867 | ENSG00000183248 | PRR36 | BLCA,CHOL,KIRP,PCPG,PRAD | BLCA | | | ENSG00000269867 | ENSG00000184307 | ZDHHC23 | BLCA,COAD,ESCA,KICH,LIHC,LUAD,TGCT | BLCA,KICH | | | ENSG00000269867 | ENSG00000185101 | ANO9 | BLCA,CESC,CHOL,KIRC,KIRP,PAAD | KIRC | | | ENSG00000269867 | ENSG00000185158 | LRRC37B | BLCA,CHOL,KIRC,KIRP,LUAD,SKCM,STAD,TGCT,THCA | KIRC | | | ENSG00000269867 | ENSG00000186166 | CENATAC | BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,OV,PAAD,PCPG,PRAD,READ,SARC,SKCM,STAD,THCA,THYM,UCEC,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000187961 | KLHL17 | BLCA,CHOL,DLBC,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,THCA,THYM,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000188559 | RALGAPA2 | BLCA,CESC,CHOL,ESCA,KICH,UCS | KICH | | | ENSG00000269867 | ENSG00000196123 | KIAA0895L | BLCA,CESC,CHOL,GBM,HNSC,KICH,KIRC,KIRP,LIHC,LUAD,LUSC,MESO,OV,PAAD,PCPG,PRAD,SKCM,THCA,THYM,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000197948 | FCHSD1 | BLCA,CHOL,GBM,KIRC,KIRP,LGG,PAAD,THCA,THYM | KIRC | | | ENSG00000269867 | ENSG00000203772 | SPRN | BLCA,CHOL,HNSC,KIRP,LIHC,PAAD,READ,THCA,UCEC,UVM | BLCA | | | ENSG00000269867 | ENSG00000204991 | SPIRE2 | BLCA,CESC,CHOL,ESCA,GBM,KICH,KIRP,LIHC,LUAD,MESO,PAAD,PCPG,UCS | KICH | | | ENSG00000269867 | ENSG00000205517 | RGL3 | BLCA,CESC,CHOL,ESCA,KICH,KIRC,KIRP,LIHC,LUAD,PAAD,PCPG,THCA,UCS | BLCA,KICH | | | ENSG00000269867 | ENSG00000205560 | CPT1B | BLCA,CESC,CHOL,DLBC,KICH | BLCA,KICH | | | ENSG00000269867 | ENSG00000213139 | CRYGS | BLCA,CHOL,DLBC,KIRP,PAAD,TGCT,UCEC | BLCA | | | ENSG00000269867 | ENSG00000214021 | TTLL3 | BLCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,LUAD,LUSC,PAAD,PCPG,PRAD,READ,SKCM,THCA,THYM,UCEC | KIRC | | | ENSG00000269867 | ENSG00000215252 | GOLGA8B | BLCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,KICH,KIRC,KIRP,LIHC,LUAD,LUSC,OV,PCPG,PRAD,READ,SKCM,TGCT,THCA,THYM,UCEC,UCS,UVM | BLCA,KIRC | | | ENSG00000269867 | ENSG00000217930 | PAM16 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LGG,MESO,PAAD,PCPG,PRAD,TGCT,THCA,UVM | KIRC | | | ENSG00000269867 | ENSG00000240230 | COX19 | BLCA,CHOL,COAD,DLBC,ESCA,HNSC,KICH,KIRC,KIRP,LGG,LIHC,PAAD,READ,STAD,THCA,UCEC,UVM | KICH | | | ENSG00000269867 | ENSG00000256525 | POLG2 | BLCA,CHOL,COAD,DLBC,ESCA,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,READ,SKCM,STAD,TGCT,THCA,THYM,UVM | KIRC | | | ENSG00000269867 | ENSG00000273899 | NOL12 | BLCA,CHOL,DLBC,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,PAAD,PCPG,PRAD,TGCT,THCA,THYM | KIRC | | | ENSG00000269867 | ENSG00000168924 | LETM1 | BRCA,CHOL,DLBC,GBM,KICH,LIHC,TGCT | KICH | | | ENSG00000269867 | ENSG00000101194 | SLC17A9 | CESC,DLBC,ESCA,KIRC,KIRP,LIHC,LUAD,PAAD | KIRC | | | ENSG00000269867 | ENSG00000142102 | PGGHG | CESC,CHOL,GBM,HNSC,KIRC,KIRP,LGG,PAAD,THCA,UCS,UVM | KIRC | | | ENSG00000269867 | ENSG00000154760 | SLFN13 | CESC,CHOL,ESCA,KIRC,KIRP,PAAD | KIRC | | | ENSG00000269867 | ENSG00000171105 | INSR | CESC,CHOL,ESCA,KICH,UVM | KICH | | | ENSG00000269867 | ENSG00000196684 | HSH2D | CESC,ESCA,KIRC,KIRP,LUAD,PAAD | KIRC | | | ENSG00000269867 | ENSG00000213066 | CEP43 | CESC,COAD,DLBC,ESCA,KIRC,KIRP,LAML,READ,TGCT,THCA,THYM,UCEC | KIRC | | | ENSG00000269867 | ENSG00000013523 | ANGEL1 | CHOL,GBM,KICH,LIHC,TGCT | KICH | | | ENSG00000269867 | ENSG00000062716 | VMP1 | CHOL,COAD,ESCA,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,UVM | KIRC | | | ENSG00000269867 | ENSG00000070770 | CSNK2A2 | CHOL,COAD,KICH,READ,THYM | KICH | | | ENSG00000269867 | ENSG00000088356 | PDRG1 | CHOL,ESCA,KICH,LGG,LIHC,MESO,THCA | KICH | | | ENSG00000269867 | ENSG00000100099 | HPS4 | CHOL,DLBC,GBM,KICH,KIRC,KIRP,LIHC,LUAD,PAAD,SARC,TGCT,UVM | KICH | | | ENSG00000269867 | ENSG00000103550 | KNOP1 | CHOL,KICH,KIRC,LAML,LIHC,READ,STAD,UVM | KICH | | | ENSG00000269867 | ENSG00000105327 | BBC3 | CHOL,ESCA,HNSC,KICH,KIRC,LIHC,PCPG,THYM | KICH,KIRC | | | ENSG00000269867 | ENSG00000106462 | EZH2 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,UVM | KIRC | | | ENSG00000269867 | ENSG00000109881 | CCDC34 | CHOL,ESCA,KICH,KIRC,LAML,STAD,THCA,UVM | KICH | | | ENSG00000269867 | ENSG00000123136 | DDX39A | CHOL,HNSC,KIRC,KIRP,LGG,LIHC,PAAD,PCPG,PRAD,TGCT | KIRC | | | ENSG00000269867 | ENSG00000133398 | MED10 | CHOL,KICH,LGG,LIHC,THCA,UVM | KICH | | | ENSG00000269867 | ENSG00000134285 | FKBP11 | CHOL,ESCA,KIRC,KIRP,TGCT | KIRC | | | ENSG00000269867 | ENSG00000136270 | TBRG4 | CHOL,KICH,LIHC,PAAD,TGCT | KICH | | | ENSG00000269867 | ENSG00000143554 | SLC27A3 | CHOL,KIRC,LGG,TGCT,THCA | KIRC | | | ENSG00000269867 | ENSG00000145014 | TMEM44 | CHOL,DLBC,KIRC,KIRP,LGG,LIHC,PAAD,PCPG,PRAD,THCA,THYM | KIRC | | | ENSG00000269867 | ENSG00000147437 | GNRH1 | CHOL,KIRC,KIRP,PAAD,THYM | KIRC | | | ENSG00000269867 | ENSG00000161960 | EIF4A1 | CHOL,DLBC,GBM,KIRC,KIRP,LGG,LUAD,LUSC,SKCM,UVM | KIRC | | | ENSG00000269867 | ENSG00000164877 | MICALL2 | CHOL,ESCA,HNSC,KIRC,KIRP,LIHC,PAAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000165724 | ZMYND19 | CHOL,KICH,KIRC,KIRP,LGG,LIHC,TGCT,THCA | KICH | | | ENSG00000269867 | ENSG00000166012 | TAF1D | CHOL,COAD,KIRC,KIRP,LAML,LIHC,LUAD,PAAD,PRAD,READ,SKCM,THCA,THYM,UVM | KIRC | | | ENSG00000269867 | ENSG00000167548 | KMT2D | CHOL,KICH,KIRC,LIHC,LUAD | KICH | | | ENSG00000269867 | ENSG00000171456 | ASXL1 | CHOL,ESCA,GBM,KICH,KIRC,KIRP,LGG,LIHC,LUAD | KICH | | | ENSG00000269867 | ENSG00000177595 | PIDD1 | CHOL,GBM,HNSC,KICH,KIRC,KIRP,LGG,LIHC,PAAD,THCA | KIRC | | | ENSG00000269867 | ENSG00000181085 | MAPK15 | CHOL,HNSC,KIRC,KIRP,THCA | KIRC | | | ENSG00000269867 | ENSG00000182325 | FBXL6 | CHOL,GBM,HNSC,KIRC,KIRP,LGG,LIHC,PAAD,TGCT,THCA | KIRC | | | ENSG00000269867 | ENSG00000182472 | CAPN12 | CHOL,KIRC,KIRP,LUSC,PAAD | KIRC | | | ENSG00000269867 | ENSG00000183765 | CHEK2 | CHOL,KIRC,KIRP,LIHC,PRAD,TGCT,UVM | KIRC | | | ENSG00000269867 | ENSG00000183856 | IQGAP3 | CHOL,ESCA,GBM,KIRC,LGG,LIHC,PAAD,TGCT | KIRC | | | ENSG00000269867 | ENSG00000184445 | KNTC1 | CHOL,DLBC,GBM,KICH,KIRC,KIRP,LGG,LIHC,LUAD,PAAD,READ,TGCT | KICH,KIRC | | | ENSG00000269867 | ENSG00000185482 | STAC3 | CHOL,KIRC,KIRP,LGG,PCPG,TGCT,UVM | KIRC | | | ENSG00000269867 | ENSG00000187555 | USP7 | CHOL,COAD,KICH,READ,UVM | KICH | | | ENSG00000269867 | ENSG00000187583 | PLEKHN1 | CHOL,KIRC,KIRP,LUSC,PAAD | KIRC | | | ENSG00000269867 | ENSG00000188827 | SLX4 | CHOL,KICH,KIRC,KIRP,LUSC,PAAD | KICH | | | ENSG00000269867 | ENSG00000189227 | C15orf61 | CHOL,KICH,PRAD,TGCT,UVM | KICH | | | ENSG00000269867 | ENSG00000205268 | PDE7A | CHOL,COAD,KIRC,LGG,LIHC,TGCT,THCA,UVM | KIRC | | | ENSG00000269867 | ENSG00000213923 | CSNK1E | CHOL,KICH,KIRC,KIRP,LIHC,LUSC,PRAD,TGCT | KIRC | | | ENSG00000269867 | ENSG00000214193 | SH3D21 | CHOL,KIRC,LUSC,PAAD,PRAD,TGCT,THCA | KIRC | | | ENSG00000269867 | ENSG00000214290 | COLCA2 | CHOL,KICH,KIRP,LIHC,PAAD,THCA | KICH | | | ENSG00000269867 | ENSG00000215440 | NPEPL1 | CHOL,GBM,HNSC,KIRC,KIRP,LGG,LIHC,LUAD,LUSC,MESO,PAAD,PCPG,PRAD,THCA,UCEC | KIRC | | | ENSG00000269867 | ENSG00000240038 | AMY2B | CHOL,GBM,KIRC,KIRP,LGG,LUAD,THCA,THYM | KIRC | | | ENSG00000269867 | ENSG00000241343 | RPL36A | CHOL,KICH,PCPG,READ,SKCM | KICH | | | ENSG00000269867 | ENSG00000278259 | MYO19 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,PRAD,UVM | KICH | | | ENSG00000269867 | ENSG00000103995 | CEP152 | COAD,KICH,LAML,SKCM,TGCT | KICH | | | ENSG00000269867 | ENSG00000069493 | CLEC2D | DLBC,GBM,KIRC,KIRP,LGG,LUAD,TGCT,UCS | KIRC | | | ENSG00000269867 | ENSG00000131368 | MRPS25 | ESCA,GBM,KICH,KIRP,LUAD,PRAD,READ,THCA,UVM | KICH | | | ENSG00000269867 | ENSG00000135596 | MICAL1 | ESCA,GBM,HNSC,KIRC,KIRP,LIHC,PAAD,TGCT,THYM | KIRC | | | ENSG00000269867 | ENSG00000185189 | NRBP2 | GBM,KIRC,KIRP,LGG,THCA,UVM | KIRC | | | ENSG00000269867 | ENSG00000266338 | NBPF15 | GBM,KIRC,KIRP,LIHC,TGCT,UVM | KIRC | | | ENSG00000269867 | ENSG00000127922 | SEM1 | KICH,KIRP,LGG,READ,STAD,TGCT,THCA,UVM | KICH | | | ENSG00000269867 | ENSG00000130204 | TOMM40 | KICH,LGG,MESO,PCPG,PRAD,THCA | KICH | | | ENSG00000269867 | ENSG00000188549 | CCDC9B | KICH,KIRC,KIRP,LIHC,UVM | KICH | | | ENSG00000269867 | ENSG00000104881 | PPP1R13L | KIRC,KIRP,LIHC,LUSC,OV,UCS | KIRC | | | ENSG00000269867 | ENSG00000198604 | BAZ1A | KIRC,KIRP,LAML,LIHC,READ,STAD,UVM | KIRC | |
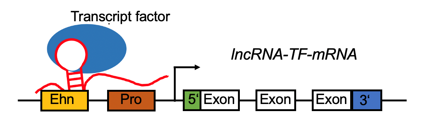
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000148399 | DPH7 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000148399 | DPH7 | 10 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000100802 | C14orf93 | 5 | CHOL,HNSC,KIRP,LIHC,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000171681 | ATF7IP | 5 | CHOL,COAD,LIHC,LUAD,READ | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000213918 | DNASE1 | 5 | BLCA,LUAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000213918 | DNASE1 | 9 | BLCA,CESC,GBM,LUAD,PRAD,SKCM,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000152359 | POC5 | 7 | CHOL,HNSC,KIRC,KIRP,LIHC,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000152359 | POC5 | 6 | ACC,KIRP,LAML,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000128191 | DGCR8 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000128191 | DGCR8 | 5 | CHOL,DLBC,GBM,KIRP,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000128191 | DGCR8 | 9 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000128191 | DGCR8 | 8 | BLCA,DLBC,GBM,KIRC,KIRP,LUSC,PRAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000182095 | TNRC18 | 5 | CHOL,GBM,LGG,LIHC,PAAD | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000177169 | ULK1 | 6 | CHOL,KIRC,KIRP,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000164548 | TRA2A | 10 | CHOL,COAD,DLBC,ESCA,GBM,KIRP,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000164548 | TRA2A | 14 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000164548 | TRA2A | 13 | ACC,BLCA,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,TGCT,THCA,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000124222 | STX16 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000124222 | STX16 | 9 | CHOL,COAD,ESCA,GBM,KIRP,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000124222 | STX16 | 13 | BLCA,CHOL,COAD,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000124222 | STX16 | 10 | BLCA,GBM,KIRP,LUAD,LUSC,PRAD,SKCM,TGCT,THCA,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000114735 | HEMK1 | 5 | KIRC,KIRP,LGG,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000114735 | HEMK1 | 5 | KIRC,KIRP,LUAD,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000114735 | HEMK1 | 6 | BLCA,KIRC,KIRP,LUAD,PRAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000142065 | ZFP14 | 5 | CHOL,GBM,KIRP,LGG,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000142065 | ZFP14 | 9 | ACC,CHOL,DLBC,ESCA,GBM,KIRP,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000142065 | ZFP14 | 9 | BLCA,CHOL,DLBC,KIRC,KIRP,LUAD,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000142065 | ZFP14 | 12 | ACC,BLCA,CESC,DLBC,GBM,KIRC,KIRP,LAML,LUAD,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000124659 | TBCC | 5 | CHOL,HNSC,LIHC,THCA,THYM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000129484 | PARP2 | 6 | CHOL,GBM,KIRC,LIHC,PRAD,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000129484 | PARP2 | 5 | GBM,KIRC,PRAD,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000170581 | STAT2 | 5 | CHOL,KIRC,KIRP,LGG,TGCT | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000205517 | RGL3 | 5 | CHOL,KIRC,KIRP,LIHC,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000205517 | RGL3 | 6 | BLCA,CHOL,KIRC,KIRP,LIHC,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000103037 | SETD6 | 7 | CHOL,GBM,KIRC,KIRP,LGG,PAAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000103037 | SETD6 | 5 | CHOL,GBM,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000103037 | SETD6 | 8 | CHOL,COAD,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000103037 | SETD6 | 7 | GBM,KIRC,KIRP,LUAD,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000180376 | CCDC66 | 8 | COAD,ESCA,GBM,KIRP,LAML,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000180376 | CCDC66 | 8 | BLCA,COAD,KIRP,LUAD,READ,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000180376 | CCDC66 | 11 | BLCA,CESC,GBM,KIRP,LAML,LUAD,PRAD,SKCM,TGCT,THCA,UCEC | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000187650 | VMAC | 9 | BLCA,CHOL,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000187650 | VMAC | 5 | BLCA,GBM,KIRC,KIRP,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000099326 | MZF1 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000099326 | MZF1 | 11 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000099326 | MZF1 | 12 | ACC,BLCA,CESC,DLBC,GBM,KIRC,KIRP,LUAD,PRAD,SKCM,THCA,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000188315 | C3orf62 | 9 | BLCA,CHOL,KIRC,KIRP,LIHC,LUAD,READ,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000188315 | C3orf62 | 8 | BLCA,KIRC,KIRP,LUAD,PRAD,SKCM,THCA,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000158863 | FAM160B2 | 6 | CHOL,GBM,KIRC,KIRP,LGG,PAAD | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000168612 | ZSWIM1 | 8 | CHOL,GBM,KIRP,LGG,LIHC,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000168612 | ZSWIM1 | 7 | CHOL,DLBC,HNSC,KIRP,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000111731 | C2CD5 | 5 | CHOL,DLBC,ESCA,KIRP,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000111731 | C2CD5 | 5 | CHOL,DLBC,KIRP,LIHC,SKCM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000177051 | FBXO46 | 10 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000177051 | FBXO46 | 7 | CHOL,DLBC,KIRP,PAAD,TGCT,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000177051 | FBXO46 | 6 | DLBC,GBM,KIRP,TGCT,THCA,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000145014 | TMEM44 | 5 | CHOL,DLBC,KIRP,LIHC,THYM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000266173 | STRADA | 6 | CHOL,GBM,KIRC,KIRP,LGG,PAAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000266173 | STRADA | 5 | CHOL,GBM,KIRP,LUAD,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000266173 | STRADA | 7 | BLCA,CHOL,KIRC,KIRP,LUAD,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000266173 | STRADA | 6 | BLCA,GBM,KIRC,KIRP,LUAD,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000163611 | SPICE1 | 7 | CHOL,COAD,GBM,KIRP,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000163611 | SPICE1 | 6 | CHOL,COAD,KIRC,KIRP,LUAD,SKCM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000163611 | SPICE1 | 6 | GBM,KIRC,KIRP,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000065883 | CDK13 | 5 | CHOL,COAD,KIRC,KIRP,LIHC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000166801 | FAM111A | 7 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000166801 | FAM111A | 5 | CHOL,COAD,KIRP,PAAD,READ | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000166801 | FAM111A | 10 | BLCA,CHOL,COAD,HNSC,KIRC,KIRP,LIHC,PAAD,READ,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000166801 | FAM111A | 6 | BLCA,CESC,GBM,KIRC,KIRP,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000132716 | DCAF8 | 5 | CHOL,KIRC,KIRP,LGG,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000132716 | DCAF8 | 10 | CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LUAD,PAAD,READ,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000132716 | DCAF8 | 8 | BLCA,DLBC,GBM,KIRC,KIRP,LUAD,THCA,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000142856 | ITGB3BP | 6 | CHOL,HNSC,KIRP,LIHC,SKCM,THYM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000154957 | ZNF18 | 6 | CHOL,HNSC,KIRC,KIRP,READ,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000078699 | CBFA2T2 | 7 | CHOL,GBM,KICH,KIRC,KIRP,LIHC,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000078699 | CBFA2T2 | 6 | CHOL,COAD,KIRC,KIRP,LIHC,READ | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000078699 | CBFA2T2 | 5 | GBM,KIRC,KIRP,LUSC,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000104936 | DMPK | 5 | CHOL,KIRC,KIRP,LIHC,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000127452 | FBXL12 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000127452 | FBXL12 | 9 | BLCA,CHOL,DLBC,KIRC,KIRP,LIHC,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000127452 | FBXL12 | 5 | BLCA,DLBC,KIRP,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000185684 | EP400P1 | 6 | BLCA,CESC,DLBC,GBM,LUAD,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000125351 | UPF3B | 5 | CHOL,KIRC,KIRP,LGG,LIHC | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000125351 | UPF3B | 7 | CHOL,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000104731 | KLHDC4 | 5 | CHOL,GBM,KIRC,LGG,PAAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000105708 | ZNF14 | 6 | ACC,CHOL,DLBC,GBM,LAML,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000105708 | ZNF14 | 5 | BLCA,CHOL,DLBC,SKCM,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000105708 | ZNF14 | 6 | ACC,BLCA,DLBC,LAML,SKCM,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000162194 | LBHD1 | 5 | BLCA,DLBC,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000108306 | FBXL20 | 5 | CHOL,KIRC,KIRP,LIHC,LUAD | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000104859 | CLASRP | 9 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000104859 | CLASRP | 11 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000104859 | CLASRP | 8 | ACC,BLCA,DLBC,GBM,KIRP,LUSC,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000100445 | SDR39U1 | 5 | CHOL,GBM,KIRC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000100445 | SDR39U1 | 6 | CHOL,DLBC,HNSC,KIRC,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000100445 | SDR39U1 | 5 | DLBC,GBM,KIRC,PRAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000172613 | RAD9A | 7 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000172613 | RAD9A | 9 | BLCA,CHOL,HNSC,KIRC,KIRP,LIHC,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000156504 | PABIR2 | 6 | CHOL,COAD,KIRC,KIRP,LIHC,PAAD | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000213339 | QTRT1 | 7 | CHOL,GBM,KICH,KIRC,KIRP,LGG,PAAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000198586 | TLK1 | 5 | CHOL,COAD,DLBC,GBM,LAML | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000110455 | ACCS | 7 | BLCA,HNSC,KIRC,KIRP,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000167280 | ENGASE | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000167280 | ENGASE | 12 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000167280 | ENGASE | 8 | BLCA,DLBC,GBM,KIRP,LUSC,PRAD,SKCM,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000168887 | C2orf68 | 6 | CHOL,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000130382 | MLLT1 | 5 | CHOL,GBM,KICH,LGG,LIHC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000105771 | SMG9 | 7 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LIHC | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000174796 | THAP6 | 5 | COAD,KIRP,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000174796 | THAP6 | 6 | COAD,KIRC,KIRP,LUAD,SKCM,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000174796 | THAP6 | 7 | GBM,KIRC,KIRP,LAML,LUAD,SKCM,TGCT | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000105443 | CYTH2 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PRAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000105443 | CYTH2 | 8 | BLCA,CHOL,DLBC,HNSC,KIRP,LIHC,TGCT,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000148444 | COMMD3 | 8 | CHOL,HNSC,KIRP,LIHC,PAAD,SKCM,TGCT,THYM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000168395 | ING5 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000168395 | ING5 | 6 | CHOL,DLBC,GBM,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000168395 | ING5 | 10 | CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000168395 | ING5 | 9 | DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000266967 | AARSD1 | 7 | CHOL,GBM,KIRC,KIRP,LGG,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000266967 | AARSD1 | 5 | CHOL,GBM,LAML,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000266967 | AARSD1 | 10 | BLCA,CHOL,DLBC,KIRC,KIRP,LUAD,PAAD,READ,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000266967 | AARSD1 | 11 | BLCA,CESC,DLBC,GBM,KIRP,LAML,LUAD,LUSC,PRAD,SKCM,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000198625 | MDM4 | 6 | CHOL,KIRC,KIRP,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000198625 | MDM4 | 8 | CHOL,COAD,DLBC,ESCA,KIRP,LUAD,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000198625 | MDM4 | 13 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000198625 | MDM4 | 11 | BLCA,CESC,DLBC,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,THCA,UCS | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000183891 | TTC32 | 6 | CHOL,COAD,DLBC,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000183891 | TTC32 | 10 | CHOL,COAD,DLBC,HNSC,KIRC,KIRP,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000183891 | TTC32 | 5 | DLBC,KIRC,KIRP,SKCM,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000154781 | CCDC174 | 6 | CHOL,DLBC,ESCA,GBM,KIRP,READ | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000154781 | CCDC174 | 8 | CHOL,DLBC,HNSC,KIRP,LIHC,READ,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000166432 | ZMAT1 | 6 | KIRC,KIRP,PRAD,TGCT,THCA,UCS | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000126870 | DYNC2I1 | 6 | CHOL,GBM,KIRC,KIRP,LGG,LIHC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000249115 | HAUS5 | 9 | CHOL,GBM,KICH,KIRC,LGG,LIHC,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000249115 | HAUS5 | 8 | CHOL,DLBC,HNSC,KIRP,LIHC,PAAD,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000249115 | HAUS5 | 7 | ACC,DLBC,GBM,KIRP,PRAD,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000108773 | KAT2A | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000108773 | KAT2A | 10 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000108773 | KAT2A | 9 | BLCA,GBM,KIRP,LUSC,PRAD,SKCM,TGCT,THCA,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000002016 | RAD52 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000002016 | RAD52 | 7 | CHOL,DLBC,GBM,KIRP,LUAD,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000002016 | RAD52 | 10 | BLCA,CHOL,DLBC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000002016 | RAD52 | 11 | BLCA,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,THCA,UCEC | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000138399 | FASTKD1 | 5 | GBM,KIRP,LAML,TGCT,UCEC | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000157106 | SMG1 | 7 | CHOL,COAD,GBM,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000157106 | SMG1 | 7 | CHOL,COAD,KIRC,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000157106 | SMG1 | 5 | GBM,KIRC,KIRP,LUAD,SKCM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000100416 | TRMU | 6 | CHOL,KIRC,KIRP,LGG,LIHC,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000100416 | TRMU | 7 | BLCA,CHOL,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000100201 | DDX17 | 5 | CHOL,GBM,KIRC,KIRP,LIHC | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000100201 | DDX17 | 7 | CHOL,COAD,DLBC,GBM,KIRP,LUAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000100201 | DDX17 | 12 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000100201 | DDX17 | 9 | BLCA,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,SKCM,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000174579 | MSL2 | 6 | CHOL,COAD,DLBC,KIRP,LUAD,READ | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000174579 | MSL2 | 9 | CHOL,COAD,DLBC,KIRP,LIHC,LUAD,READ,TGCT,THYM | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000197302 | ZNF720 | 6 | CHOL,COAD,GBM,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000197302 | ZNF720 | 5 | BLCA,GBM,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000167766 | ZNF83 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000167766 | ZNF83 | 11 | ACC,CHOL,COAD,DLBC,GBM,KIRP,LAML,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000167766 | ZNF83 | 13 | BLCA,CHOL,COAD,DLBC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000167766 | ZNF83 | 16 | ACC,BLCA,CESC,DLBC,GBM,KIRC,KIRP,LAML,LUAD,LUSC,PRAD,SKCM,TGCT,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000117360 | PRPF3 | 7 | GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000117360 | PRPF3 | 10 | COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000117360 | PRPF3 | 6 | GBM,KIRC,KIRP,LUAD,PRAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000121289 | CEP89 | 5 | CHOL,GBM,KIRP,LGG,LIHC | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000152457 | DCLRE1C | 5 | CHOL,KIRC,KIRP,READ,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000178177 | LCORL | 5 | CHOL,COAD,ESCA,LAML,READ | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000007392 | LUC7L | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000007392 | LUC7L | 6 | CHOL,COAD,GBM,LUAD,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000007392 | LUC7L | 13 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000007392 | LUC7L | 13 | ACC,BLCA,CESC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,TGCT,THCA,UCEC | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000181007 | ZFP82 | 5 | DLBC,GBM,LAML,TGCT,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000104885 | DOT1L | 6 | CHOL,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000104885 | DOT1L | 5 | GBM,KIRC,KIRP,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000127511 | SIN3B | 7 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000127511 | SIN3B | 5 | ACC,CHOL,DLBC,GBM,KIRP | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000127511 | SIN3B | 9 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000127511 | SIN3B | 7 | BLCA,DLBC,KIRC,KIRP,LUSC,TGCT,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000112167 | SAYSD1 | 6 | CHOL,HNSC,LIHC,TGCT,THCA,THYM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000063244 | U2AF2 | 7 | CHOL,GBM,KIRC,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000063244 | U2AF2 | 5 | CHOL,HNSC,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000124459 | ZNF45 | 5 | CHOL,GBM,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000124459 | ZNF45 | 6 | ACC,CHOL,DLBC,GBM,LAML,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000124459 | ZNF45 | 5 | CHOL,DLBC,LIHC,SKCM,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000124459 | ZNF45 | 6 | ACC,DLBC,GBM,LAML,SKCM,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000164074 | ABHD18 | 7 | DLBC,ESCA,KIRP,LAML,LUAD,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000164074 | ABHD18 | 8 | BLCA,DLBC,KIRC,KIRP,LUAD,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000164074 | ABHD18 | 9 | BLCA,DLBC,GBM,KIRC,KIRP,LAML,LUAD,SKCM,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000105717 | PBX4 | 6 | BLCA,KIRP,LUAD,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000104957 | CCDC130 | 7 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000104957 | CCDC130 | 9 | BLCA,CHOL,HNSC,KIRC,KIRP,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000104957 | CCDC130 | 5 | BLCA,KIRP,LUSC,THCA,UCEC | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000119688 | ABCD4 | 6 | GBM,KIRC,KIRP,LGG,PAAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000119688 | ABCD4 | 9 | BLCA,DLBC,HNSC,KIRC,KIRP,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000159079 | CFAP298 | 5 | CHOL,KIRP,LIHC,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000159079 | CFAP298 | 5 | ACC,KIRP,LAML,LUSC,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000164073 | MFSD8 | 9 | CHOL,COAD,DLBC,ESCA,KIRP,LAML,LUAD,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000164073 | MFSD8 | 9 | BLCA,CHOL,COAD,DLBC,HNSC,KIRP,LUAD,PAAD,SKCM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000164073 | MFSD8 | 6 | BLCA,DLBC,KIRP,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000163660 | CCNL1 | 7 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000163660 | CCNL1 | 9 | CHOL,COAD,DLBC,GBM,KIRP,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000163660 | CCNL1 | 14 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000163660 | CCNL1 | 9 | BLCA,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,SKCM,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000149532 | CPSF7 | 5 | DLBC,KIRC,KIRP,LIHC,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000119231 | SENP5 | 5 | CHOL,KIRC,KIRP,LIHC,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000119231 | SENP5 | 5 | CHOL,COAD,GBM,KIRP,LUAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000119231 | SENP5 | 10 | CHOL,COAD,KIRC,KIRP,LIHC,LUAD,READ,TGCT,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000119231 | SENP5 | 6 | GBM,KIRC,KIRP,LUAD,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000188827 | SLX4 | 5 | CHOL,KICH,KIRC,KIRP,PAAD | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000004534 | RBM6 | 8 | GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000004534 | RBM6 | 10 | CHOL,COAD,DLBC,ESCA,GBM,KIRP,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000004534 | RBM6 | 15 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,TGCT,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000004534 | RBM6 | 14 | BLCA,CESC,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,TGCT,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000090470 | PDCD7 | 6 | CHOL,DLBC,KIRC,KIRP,LIHC,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000140326 | CDAN1 | 7 | CHOL,KICH,KIRC,KIRP,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000140326 | CDAN1 | 10 | BLCA,CHOL,DLBC,HNSC,KIRC,KIRP,LIHC,PAAD,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000140326 | CDAN1 | 6 | BLCA,DLBC,KIRC,KIRP,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000101391 | CDK5RAP1 | 8 | CHOL,GBM,KICH,KIRC,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000101391 | CDK5RAP1 | 5 | GBM,KIRC,KIRP,PRAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000104064 | GABPB1 | 7 | CHOL,GBM,KIRP,LGG,LIHC,PRAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000104064 | GABPB1 | 6 | CHOL,COAD,DLBC,GBM,KIRP,LUAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000104064 | GABPB1 | 10 | CHOL,COAD,DLBC,KIRC,KIRP,LIHC,LUAD,TGCT,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000104064 | GABPB1 | 7 | DLBC,GBM,KIRC,KIRP,LUAD,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000213780 | GTF2H4 | 6 | CHOL,GBM,KIRC,KIRP,LGG,PAAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000213780 | GTF2H4 | 5 | CHOL,GBM,LAML,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000213780 | GTF2H4 | 6 | CHOL,KIRC,KIRP,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000213780 | GTF2H4 | 5 | BLCA,GBM,KIRP,LAML,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000074219 | TEAD2 | 5 | CHOL,HNSC,KIRP,LIHC,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000128694 | OSGEPL1 | 5 | COAD,DLBC,KIRP,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000128694 | OSGEPL1 | 5 | DLBC,LAML,PRAD,TGCT,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000166526 | ZNF3 | 7 | BLCA,CHOL,KIRC,KIRP,LIHC,READ,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000168661 | ZNF30 | 5 | GBM,KIRP,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000168661 | ZNF30 | 7 | ACC,GBM,KIRP,LAML,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000168661 | ZNF30 | 9 | BLCA,COAD,KIRC,KIRP,LIHC,PAAD,READ,SKCM,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000168661 | ZNF30 | 9 | ACC,BLCA,CESC,GBM,KIRC,KIRP,LAML,SKCM,TGCT | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000064547 | LPAR2 | 5 | CHOL,GBM,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000165792 | METTL17 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000165792 | METTL17 | 9 | CHOL,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000165792 | METTL17 | 5 | KIRC,KIRP,PRAD,SKCM,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000116001 | TIA1 | 10 | CHOL,COAD,DLBC,ESCA,KIRP,LAML,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000116001 | TIA1 | 14 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000116001 | TIA1 | 14 | BLCA,CESC,DLBC,GBM,KIRC,KIRP,LAML,LUAD,LUSC,PRAD,SKCM,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000003756 | RBM5 | 9 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000003756 | RBM5 | 10 | CHOL,COAD,DLBC,ESCA,GBM,KIRP,LUAD,PAAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000003756 | RBM5 | 14 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000003756 | RBM5 | 13 | BLCA,CESC,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,TGCT,THCA,UCEC | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000198707 | CEP290 | 7 | CHOL,COAD,KIRP,LAML,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000198707 | CEP290 | 8 | BLCA,CHOL,COAD,KIRC,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000198707 | CEP290 | 6 | BLCA,KIRC,KIRP,LAML,LUAD,SKCM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000198917 | SPOUT1 | 7 | GBM,KIRC,KIRP,LGG,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000111653 | ING4 | 5 | CHOL,HNSC,KIRP,SKCM,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000149187 | CELF1 | 5 | GBM,KIRC,KIRP,LIHC,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000149187 | CELF1 | 5 | COAD,GBM,KIRP,LUAD,READ | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000149187 | CELF1 | 7 | COAD,KIRC,KIRP,LIHC,LUAD,READ,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000149187 | CELF1 | 6 | GBM,KIRC,KIRP,LUAD,LUSC,TGCT | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000132199 | ENOSF1 | 7 | CHOL,KIRC,KIRP,LGG,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000132199 | ENOSF1 | 9 | BLCA,HNSC,KIRC,KIRP,LUAD,PAAD,TGCT,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000132199 | ENOSF1 | 11 | BLCA,DLBC,KIRC,KIRP,LUAD,LUSC,PRAD,TGCT,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000172890 | NADSYN1 | 6 | BLCA,KIRC,KIRP,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000113119 | TMCO6 | 7 | BLCA,DLBC,HNSC,KIRC,KIRP,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000113119 | TMCO6 | 5 | BLCA,DLBC,KIRP,TGCT,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000124383 | MPHOSPH10 | 5 | CHOL,DLBC,KIRP,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000124383 | MPHOSPH10 | 8 | CHOL,COAD,DLBC,KIRP,LIHC,READ,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000124383 | MPHOSPH10 | 6 | DLBC,KIRP,LAML,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000046647 | GEMIN8 | 5 | BLCA,CHOL,KIRP,LIHC,THCA | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000151849 | CENPJ | 6 | CHOL,COAD,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000151849 | CENPJ | 10 | CHOL,COAD,KIRC,KIRP,LUAD,PAAD,READ,SKCM,TGCT,THYM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000188735 | TMEM120B | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000188735 | TMEM120B | 5 | GBM,KIRC,KIRP,LUAD,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000132680 | KHDC4 | 8 | CHOL,GBM,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000132680 | KHDC4 | 8 | CHOL,COAD,ESCA,GBM,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000132680 | KHDC4 | 13 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,READ,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000132680 | KHDC4 | 13 | BLCA,CESC,DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000100650 | SRSF5 | 6 | GBM,KIRC,KIRP,LGG,PAAD,PRAD | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000100650 | SRSF5 | 6 | ACC,DLBC,GBM,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000100650 | SRSF5 | 9 | DLBC,HNSC,KIRC,KIRP,LUAD,PAAD,SKCM,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000100650 | SRSF5 | 9 | DLBC,GBM,KIRC,KIRP,LUAD,LUSC,PRAD,SKCM,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000156858 | PRR14 | 7 | CHOL,KIRC,KIRP,LGG,LIHC,PAAD,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000156858 | PRR14 | 7 | BLCA,CHOL,HNSC,KIRP,LIHC,PAAD,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000082512 | TRAF5 | 7 | COAD,HNSC,KIRC,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000129933 | MAU2 | 9 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,TGCT | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000129933 | MAU2 | 7 | ACC,CHOL,DLBC,GBM,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000129933 | MAU2 | 11 | BLCA,CHOL,DLBC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000129933 | MAU2 | 6 | BLCA,DLBC,GBM,KIRP,TGCT,THCA | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000185917 | SETD4 | 8 | CHOL,GBM,KIRC,KIRP,LIHC,PAAD,PRAD,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000185917 | SETD4 | 12 | BLCA,CHOL,COAD,HNSC,KIRC,KIRP,LIHC,LUAD,PAAD,SKCM,TGCT,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000185917 | SETD4 | 12 | ACC,BLCA,GBM,KIRC,KIRP,LUAD,PRAD,SKCM,TGCT,THCA,UCEC,UCS | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000177025 | C19orf18 | 6 | ACC,BLCA,GBM,LAML,PRAD,UCEC | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000129566 | TEP1 | 5 | CHOL,DLBC,KIRC,LIHC,PAAD | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000118162 | KPTN | 5 | CHOL,GBM,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000102878 | HSF4 | 6 | BLCA,HNSC,KIRC,KIRP,PAAD,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000102878 | HSF4 | 5 | BLCA,KIRP,PRAD,THCA,UCEC | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000154144 | TBRG1 | 8 | BLCA,KIRC,KIRP,LIHC,LUAD,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000154144 | TBRG1 | 6 | BLCA,KIRC,KIRP,LUAD,THCA,UCEC | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000136243 | NUP42 | 6 | CHOL,COAD,DLBC,KIRP,PAAD,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000136243 | NUP42 | 11 | BLCA,CHOL,COAD,DLBC,HNSC,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000136243 | NUP42 | 6 | BLCA,DLBC,KIRC,KIRP,SKCM,THCA | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000135457 | TFCP2 | 5 | CHOL,COAD,DLBC,LIHC,READ | | ENSG00000269867 | ENSG00000167232 | ZNF91 | ENSG00000135870 | RC3H1 | 8 | CHOL,COAD,DLBC,GBM,KIRP,LUAD,READ,SKCM | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000135870 | RC3H1 | 9 | CHOL,COAD,DLBC,KIRP,LIHC,LUAD,READ,SKCM,TGCT | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000135870 | RC3H1 | 7 | DLBC,GBM,KIRP,LUAD,SKCM,TGCT,UCS | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000166479 | TMX3 | 5 | CHOL,DLBC,KIRP,LIHC,TGCT | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000103091 | WDR59 | 5 | CHOL,KIRC,KIRP,LGG,PRAD | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000103091 | WDR59 | 7 | BLCA,KIRC,KIRP,LUAD,PAAD,THCA,THYM | | ENSG00000269867 | ENSG00000256223 | ZNF10 | ENSG00000103091 | WDR59 | 7 | BLCA,KIRC,KIRP,LUAD,PRAD,THCA,UCS | | ENSG00000269867 | ENSG00000104885 | DOT1L | ENSG00000136811 | ODF2 | 5 | CHOL,GBM,LGG,LIHC,TGCT | | ENSG00000269867 | ENSG00000236287 | ZBED5 | ENSG00000099622 | CIRBP | 6 | BLCA,CHOL,HNSC,KIRP,LIHC,THCA |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000269867 | ENST00000597780 | PCDHGC3 | ENSG00000240184 | ENST00000611950 | -0.1503 | 133 | - | KICH,BLCA | | ENSG00000269867 | ENST00000597780 | PCDHGC3 | ENSG00000240184 | ENST00000617222 | -0.2774 | 91 | - | KICH,BLCA | | ENSG00000269867 | ENST00000597780 | C1R | ENSG00000159403 | ENST00000536053 | -0.1057 | 74 | - | KICH,BLCA | | ENSG00000269867 | ENST00000597780 | C1R | ENSG00000159403 | ENST00000535233 | -0.1057 | 74 | - | KICH,BLCA | | ENSG00000269867 | ENST00000597780 | MSRA | ENSG00000175806 | ENST00000441698 | -0.3682 | 52 | - | KIRC,KICH,BRCA | | ENSG00000269867 | ENST00000597780 | MSRA | ENSG00000175806 | ENST00000518255 | -1.0050 | 295 | - | KIRC,KICH,BRCA | | ENSG00000269867 | ENST00000597780 | MSRA | ENSG00000175806 | ENST00000521686 | -0.1019 | 1 | - | KIRC,KICH,BRCA | | ENSG00000269867 | ENST00000597780 | AKR7A2 | ENSG00000053371 | ENST00000489286 | -0.1359 | 387 | - | KICH |
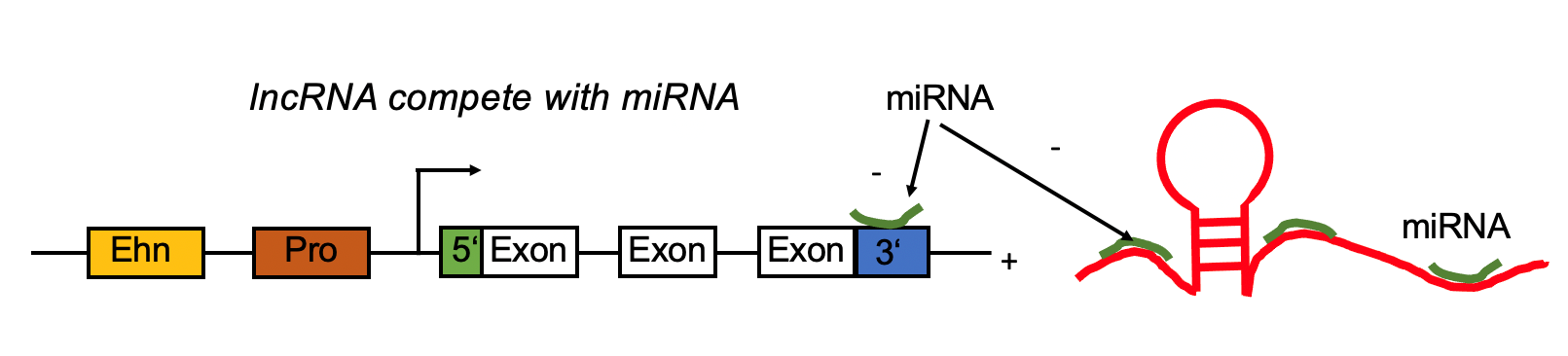
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000269867 | CTD-2583A14.8,hsa-mir-152,COX7C | KICH,TGCT,CHOL,PRAD,UCS | | ENSG00000269867 | CTD-2583A14.8,hsa-mir-152,HIGD2A | KICH,TGCT,CHOL,PRAD,UCS |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000269867.1 | ENST00000597780.1 | 89 | 130 | 399 |
|

 Gene summary
Gene summary AS events and RNA A-to-I editing events of lncRNA in TCGA based on Genvode V22 structure.
AS events and RNA A-to-I editing events of lncRNA in TCGA based on Genvode V22 structure.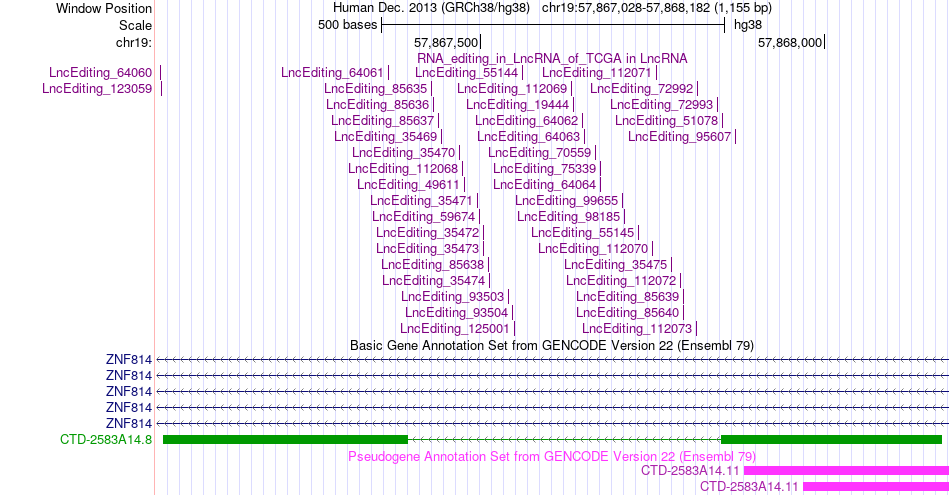
 Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).