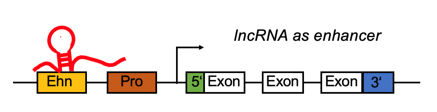
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | CEBPZ,ELF4,PUM1,TBX3,KLF9,XRN1,BCL6,CPEB2,PBX2,KLF12,ZBTB4,GLYR1,GRSF1,KLF7,RBM7,JAZF1,MYSM1,CSDE1,RBM23,ARID2,ZXDC,SUGP2,FOXF2,RARB,DMTF1,ELF1,FOSL2,VEZF1,GLI3,RBM27,ZNF70,BAZ2A,ZNF24,DDX52,RORA,RUNX1,ATF6B,TBP,GABPA,NFKB1,IKZF5,DDX24,MEF2C,GLIS3,ZMAT3,TET3,ZBTB5,SRP68,HEYL,ZFP14,ZXDA,RNPC3,TFE3,TRPS1,KDM5B,NOP9,ATMIN,FOXO1,LARP7,TRNT1,CNOT4,SON,CWC22,DDX59,PBX3,ZZZ3,PNO1,MLX,ZBED3,DDX60,KLF6,STAT3,CREB1,MEX3C,RC3H1,SP100,HIF1A,EEA1,ZNF91,ZFP91,RBMX,AGO2,RBMS3,TIGD6,RBM26,ZFP1,PURB,ADNP,GZF1,NCBP1,MYEF2,PPIL4,MBD1,IKZF4,PUS7L,ASH1L,MEIS1,CLOCK,NFIA,MEIS2,FUBP1,ARNT2,RBM43,FLI1,KLF11,PRDM4,TCF20,LIN54,AGFG1,UBR5,THOC2,ZBTB6,THAP6,RBM47,ERG,DDX17,SRRM2,RFX7,E2F6,FOXN3,SP2,ZHX3,LCOR,FMR1,MECP2,FOXJ3,KLF3,PURA,PHAX,ZNF41,KLF15,TTF1,RBM41,SART3,DZIP3,SRBD1,IRF2,ZFP62,FOXF1,KLF10,GATA6,SMAD3,PAIP1,CPSF2,ZNF45,CELF1,SCMH1,KRR1,TMF1,ELK4,NFXL1,BACH1,PRDM2,AHR,RBM18,UNC50,SALL2,MYNN,TFCP2,PATL1,QKI,ZFR,STAU1,ZFX,NAF1,NR2C2,SYNJ1,STAT6,SIX5,SALL1,GPBP1,C1D,ARNT,KDM2B,G3BP2,JDP2,PLS1,LSM11,ZFHX3,DZIP1,PUM2,XPO1,TEAD3,ELF2,SMAD4,TIGD2,RBMS1,NFYB,ZNF43,MTF1,SP3,FOXC1,SRCAP,ETS1,DDX42,BAZ2B,ZHX2,CSTF2,AHDC1,KMT2A,ZEB2,PDCD4,NACC2,SF3A1,MAF,DACH1,ZFP3,MEF2D,TEP1,MECOM,ADAT1,ZBED6,ETV1,TET2,RBM33,NOL8,ATF2 | | Exist in public source | | NA |
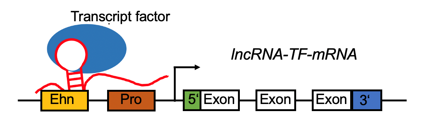
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | ZHX3,AGO1,IRF2,FOXO3,BCL6,DDX17,DACH1,ELF1,UBR5,AGO2,ZBTB4,GTF2I,RFX7,ASH1L,DDX52,ATF1,MBNL1,SMAD1,FOXO4,ADAT1,SF3A1,RBMS2,HLF,TFCP2,TERF1,REST,MYNN,NCBP1,PATL1,PPIL4,RUNX1,ZFHX3,ZNF70,AGO4,FOXF1,RBMS3,TEP1,ZMAT3,CELF2,NFIA,CELF1,RORA,FOXJ2,ZFP91,ARID2,STAU1,ZBTB6,DDX1,FLI1 | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
| Only predicted by TDF | | RRP7A,EIF3G,E4F1,RBM42,RPP25,IRF3 | | Exist in public source | | NA |
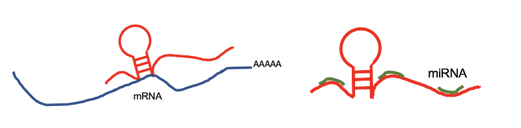
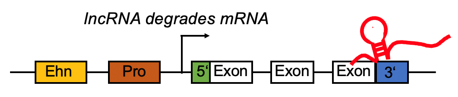
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | RPP25,RALY,LSM4,SNRPA,SLIRP,EIF3G,DDX41,THOC6,LSM7,RNPS1,DDX56,DDX49,PUF60,USF2,HSF1,CENPA,GLI4,SUGP1,KLF16,THAP7,NR2F6,IRF3,RBM42,E4F1,PA2G4,MBD3 | | Exist in public source | | NA |
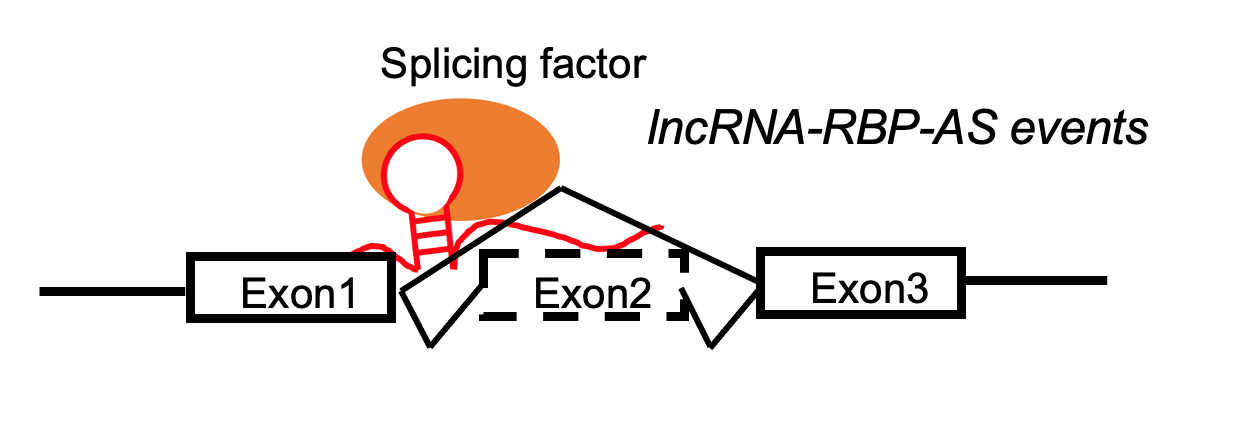
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000265519 | ENST00000582780 | exon_skip_7558 | chr1:81952986-81953025 | -6.60 | -0.5500 | ADGRL2 | ENST00000370717,ENST00000370728 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_21628 | chr1:9756480-9756510 | -6.55 | -0.5955 | CLSTN1 | ENST00000377298 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_33516 | chr1:156938417-156938513 | -13.38 | -0.1911 | ARHGEF11 | ENST00000361409 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_43507 | chr10:84499874-84499959 | -15.18 | -0.1946 | CCSER2 | ENST00000224756 | Frame-shift | | ENSG00000265519 | ENST00000582780 | exon_skip_90104 | chr12:10709907-10710114 | -32.73 | -0.1581 | YBX3 | ENST00000228251 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_287874 | chr17:29085603-29085648 | -10.45 | -0.2750 | MYO18A | ENST00000527372 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_436488 | chr5:96726793-96726859 | -7.20 | -0.1125 | CAST | ENST00000341926,ENST00000395813 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_29583 | chr1:114780109-114780215 | -8.92 | -0.1898 | SIKE1 | ENST00000060969 | Frame-shift | | ENSG00000265519 | ENST00000582780 | exon_skip_59439 | chr11:57737912-57738026 | -11.06 | -0.2698 | TMX2 | ENST00000278422 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_89597 | chr12:6672349-6672532 | -24.39 | -0.1524 | ZNF384 | ENST00000361959,ENST00000396801 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_120741 | chr15:34228428-34228589 | -14.32 | -0.1224 | EMC4 | ENST00000267750 | Frame-shift | | ENSG00000265519 | ENST00000582780 | exon_skip_140339 | chr16:635611-635774 | -18.68 | -0.1279 | METTL26 | ENST00000301686 | Frame-shift | | ENSG00000265519 | ENST00000582780 | exon_skip_143275 | chr16:24939363-24939597 | -29.35 | -0.1316 | ARHGAP17 | ENST00000289968 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_319500 | chr19:45413962-45414034 | -9.43 | -0.1310 | ERCC1 | ENST00000300853,ENST00000589165 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_326741 | chr2:73946718-73946906 | -20.06 | -0.1605 | DGUOK | ENST00000264093 | Frame-shift | | ENSG00000265519 | ENST00000582780 | exon_skip_358606 | chr20:63202286-63203807 | -39.02 | -0.1172 | YTHDF1 | ENST00000370339 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_445390 | chr5:150403120-150403312 | -18.52 | -0.1332 | CD74 | ENST00000009530 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_363747 | chr22:23877720-23877869 | -21.63 | -0.1492 | SLC2A11 | ENST00000345044 | Frame-shift | | ENSG00000265519 | ENST00000582780 | exon_skip_500083 | chr9:129127963-129128068 | -11.07 | -0.1118 | PTPA | ENST00000337738 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_390418 | chr3:183842297-183842447 | -10.67 | -0.1286 | PARL | ENST00000317096 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000265519 | ENST00000582780 | exon_skip_48713 | chr10:34336198-34336243 | -8.33 | -0.1893 | PARD3 | ENST00000374789 | In-frame | | ENSG00000265519 | ENST00000582780 | exon_skip_499771 | chr9:128592982-128593042 | -12.55 | -0.2368 | SPTAN1 | ENST00000372731 | In-frame |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000265519 | hsa-mir-182 | ENST00000582780 | chr17:15787999-15788120 | 185.00 | -120.68 | 116 | NA | | ENSG00000265519 | hsa-mir-182 | ENST00000582780 | chr17:15787788-15787901 | 153.00 | -87.85 | 110 | NA | | ENSG00000265519 | hsa-mir-183 | ENST00000582780 | chr17:15787821-15787937 | 176.00 | -95.97 | 107 | NA | | ENSG00000265519 | hsa-mir-183 | ENST00000582780 | chr17:15787975-15788096 | 170.00 | -110.33 | 114 | NA | | ENSG00000265519 | hsa-mir-210 | ENST00000582780 | chr17:15788000-15788115 | 180.00 | -133.01 | 112 | NA | | ENSG00000265519 | hsa-mir-210 | ENST00000582780 | chr17:15787838-15787947 | 158.00 | -105.46 | 88 | NA | | ENSG00000265519 | hsa-mir-3613 | ENST00000582780 | chr17:15787899-15787980 | 165.00 | -81.96 | 84 | NA | | ENSG00000265519 | hsa-mir-484 | ENST00000582780 | chr17:15788030-15788109 | 166.00 | -80.09 | 78 | NA | | ENSG00000265519 | hsa-mir-96 | ENST00000582780 | chr17:15788016-15788110 | 160.00 | -90.34 | 91 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000265519 | ENSG00000120756 | PLS1 | COAD,KICH,KIRC,KIRP,READ,TGCT,THCA,THYM | | COAD,READ |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000174306 | ZHX3 | 5 | ACC,BRCA,KICH,OV,THCA | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000174306 | ZHX3 | 6 | ACC,BRCA,KICH,LUSC,STAD,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000092847 | AGO1 | 5 | KIRP,MESO,OV,PCPG,THCA | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000092847 | AGO1 | 5 | KICH,KIRP,MESO,PRAD,THCA | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000092847 | AGO1 | 6 | CHOL,KIRP,MESO,OV,PRAD,THCA | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000092847 | AGO1 | 7 | CHOL,KICH,KIRP,MESO,OV,PRAD,THCA | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000092847 | AGO1 | 5 | KICH,KIRP,MESO,PRAD,THCA | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000092847 | AGO1 | 5 | KICH,KIRP,MESO,PRAD,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000092847 | AGO1 | 7 | CHOL,KICH,KIRP,MESO,PCPG,PRAD,THCA | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000092847 | AGO1 | 5 | CHOL,KIRP,MESO,PRAD,THCA | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000092847 | AGO1 | 6 | KICH,KIRP,MESO,PCPG,PRAD,THCA | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000168310 | IRF2 | 5 | CHOL,HNSC,KIRP,PRAD,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000168310 | IRF2 | 5 | CHOL,HNSC,KIRP,PRAD,THYM | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000118689 | FOXO3 | 5 | BLCA,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000118689 | FOXO3 | 5 | KIRP,TGCT,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000118689 | FOXO3 | 5 | BLCA,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000118689 | FOXO3 | 5 | KICH,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000118689 | FOXO3 | 6 | BLCA,KICH,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000118689 | FOXO3 | 6 | BLCA,KICH,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000100201 | DDX17 | 5 | ACC,BRCA,CHOL,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000100201 | DDX17 | 5 | ACC,BRCA,KICH,THYM,UCEC | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000100201 | DDX17 | 5 | ACC,CHOL,KICH,THYM,UCEC | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000120690 | ELF1 | 8 | BLCA,BRCA,KIRP,MESO,PCPG,THCA,UCEC,UCS | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000120690 | ELF1 | 8 | BRCA,KICH,KIRP,MESO,PCPG,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000120690 | ELF1 | 10 | ACC,BLCA,BRCA,CHOL,KIRC,KIRP,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000120690 | ELF1 | 8 | CHOL,KICH,KIRP,MESO,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000120690 | ELF1 | 9 | ACC,BLCA,BRCA,KICH,KIRP,MESO,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000120690 | ELF1 | 7 | KICH,KIRC,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000120690 | ELF1 | 5 | ACC,BRCA,KICH,PRAD,THCA | | ENSG00000265519 | ENSG00000168769 | TET2 | ENSG00000120690 | ELF1 | 5 | BRCA,KIRP,OV,PRAD,THCA | | ENSG00000265519 | ENSG00000169554 | ZEB2 | ENSG00000120690 | ELF1 | 5 | KICH,KIRP,PCPG,PRAD,SKCM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000120690 | ELF1 | 13 | ACC,BLCA,CHOL,KICH,KIRC,KIRP,MESO,PCPG,PRAD,SKCM,THCA,UCEC,UCS | | ENSG00000265519 | ENSG00000178764 | ZHX2 | ENSG00000120690 | ELF1 | 6 | ACC,KICH,KIRP,MESO,PCPG,PRAD | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000120690 | ELF1 | 10 | ACC,BLCA,CHOL,KIRC,KIRP,MESO,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000120690 | ELF1 | 8 | ACC,BRCA,KICH,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000104517 | UBR5 | 6 | KIRP,MESO,PCPG,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000104517 | UBR5 | 6 | KIRP,MESO,PCPG,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000104517 | UBR5 | 7 | ACC,KIRC,KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000104517 | UBR5 | 5 | KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000104517 | UBR5 | 6 | ACC,KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000104517 | UBR5 | 6 | ACC,KIRC,KIRP,MESO,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000104517 | UBR5 | 8 | ACC,KIRC,KIRP,MESO,PCPG,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000104517 | UBR5 | 6 | ACC,KIRC,KIRP,MESO,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000104517 | UBR5 | 5 | ACC,KIRP,MESO,PCPG,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000174282 | ZBTB4 | 6 | BRCA,KIRP,OV,PCPG,THCA,THYM | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000174282 | ZBTB4 | 7 | BRCA,KICH,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000106571 | GLI3 | ENSG00000174282 | ZBTB4 | 7 | BRCA,KIRP,PRAD,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000174282 | ZBTB4 | 10 | ACC,CHOL,KIRC,KIRP,OV,PRAD,READ,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000116016 | EPAS1 | ENSG00000174282 | ZBTB4 | 9 | COAD,KICH,LUSC,PRAD,READ,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000174282 | ZBTB4 | 5 | CHOL,KICH,KIRP,PRAD,THCA | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000174282 | ZBTB4 | 9 | BRCA,CESC,KICH,KIRP,MESO,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000174282 | ZBTB4 | 8 | ACC,KICH,KIRC,KIRP,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000174282 | ZBTB4 | 9 | BRCA,CESC,COAD,KICH,LUSC,PRAD,READ,STAD,TGCT | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000174282 | ZBTB4 | 7 | BRCA,KICH,KIRC,KIRP,STAD,THCA,THYM | | ENSG00000265519 | ENSG00000168769 | TET2 | ENSG00000174282 | ZBTB4 | 5 | KIRP,OV,PRAD,READ,THCA | | ENSG00000265519 | ENSG00000169554 | ZEB2 | ENSG00000174282 | ZBTB4 | 6 | COAD,KICH,LUSC,PRAD,TGCT,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000174282 | ZBTB4 | 9 | CHOL,KICH,KIRC,KIRP,MESO,PCPG,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000178764 | ZHX2 | ENSG00000174282 | ZBTB4 | 5 | ACC,KICH,KIRP,PCPG,PRAD | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000174282 | ZBTB4 | 5 | CHOL,KIRC,KIRP,READ,THCA | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000174282 | ZBTB4 | 5 | ACC,KICH,KIRP,PRAD,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000263001 | GTF2I | 10 | BLCA,BRCA,KIRP,MESO,OV,PCPG,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000100644 | HIF1A | ENSG00000263001 | GTF2I | 5 | ACC,KICH,KIRC,PRAD,UCS | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000263001 | GTF2I | 9 | BRCA,KICH,KIRP,MESO,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000106571 | GLI3 | ENSG00000263001 | GTF2I | 5 | BRCA,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000263001 | GTF2I | 13 | ACC,BLCA,BRCA,CHOL,KIRC,KIRP,MESO,OV,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000263001 | GTF2I | 10 | CHOL,KICH,KIRP,MESO,OV,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000263001 | GTF2I | 11 | ACC,BLCA,BRCA,KICH,KIRP,MESO,OV,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000263001 | GTF2I | 8 | ACC,KICH,KIRC,KIRP,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000263001 | GTF2I | 5 | ACC,KIRC,KIRP,THCA,THYM | | ENSG00000265519 | ENSG00000168769 | TET2 | ENSG00000263001 | GTF2I | 5 | BRCA,KIRP,OV,PRAD,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000263001 | GTF2I | 15 | ACC,BLCA,CHOL,KICH,KIRC,KIRP,MESO,OV,PCPG,PRAD,SKCM,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000263001 | GTF2I | 10 | ACC,BLCA,CHOL,KIRC,KIRP,MESO,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000263001 | GTF2I | 9 | ACC,BRCA,KICH,KIRP,MESO,PCPG,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000181827 | RFX7 | 6 | BRCA,KIRP,OV,PCPG,THCA,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000181827 | RFX7 | 7 | BRCA,KICH,KIRP,PCPG,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000181827 | RFX7 | 10 | ACC,BRCA,CHOL,HNSC,KIRP,OV,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000181827 | RFX7 | 7 | CHOL,KICH,KIRP,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000181827 | RFX7 | 9 | ACC,BRCA,HNSC,KICH,KIRP,OV,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000181827 | RFX7 | 6 | ACC,KICH,KIRP,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000140836 | ZFHX3 | ENSG00000181827 | RFX7 | 5 | HNSC,OV,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000168769 | TET2 | ENSG00000181827 | RFX7 | 5 | BRCA,KIRP,OV,PRAD,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000181827 | RFX7 | 11 | ACC,CHOL,HNSC,KICH,KIRP,OV,PCPG,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000181827 | RFX7 | 7 | ACC,CHOL,KIRP,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000181827 | RFX7 | 8 | ACC,BRCA,KICH,KIRP,PCPG,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000116539 | ASH1L | 9 | BLCA,BRCA,KIRP,MESO,OV,PCPG,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000116539 | ASH1L | 10 | BRCA,KICH,KIRP,MESO,PCPG,PRAD,TGCT,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000106571 | GLI3 | ENSG00000116539 | ASH1L | 6 | BRCA,KIRP,PRAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000116539 | ASH1L | 14 | ACC,BLCA,BRCA,HNSC,KIRC,KIRP,MESO,OV,PRAD,READ,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000116016 | EPAS1 | ENSG00000116539 | ASH1L | 5 | KICH,READ,STAD,TGCT,THYM | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000116539 | ASH1L | 8 | KICH,KIRP,MESO,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000116539 | ASH1L | 12 | ACC,BLCA,BRCA,HNSC,KICH,KIRP,MESO,OV,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000116539 | ASH1L | 9 | ACC,KICH,KIRC,KIRP,MESO,PRAD,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000140836 | ZFHX3 | ENSG00000116539 | ASH1L | 7 | HNSC,OV,PRAD,SKCM,STAD,TGCT,THCA | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000116539 | ASH1L | 8 | ACC,BRCA,COAD,KICH,PRAD,READ,TGCT,THCA | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000116539 | ASH1L | 7 | KICH,KIRC,KIRP,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000168769 | TET2 | ENSG00000116539 | ASH1L | 6 | BRCA,KIRP,OV,PRAD,READ,THCA | | ENSG00000265519 | ENSG00000169554 | ZEB2 | ENSG00000116539 | ASH1L | 7 | COAD,KICH,KIRP,PRAD,SKCM,TGCT,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000116539 | ASH1L | 14 | ACC,BLCA,HNSC,KICH,KIRC,KIRP,MESO,OV,PCPG,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000116539 | ASH1L | 10 | ACC,BLCA,KIRC,KIRP,MESO,PRAD,READ,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000116539 | ASH1L | 9 | ACC,BRCA,KICH,KIRP,MESO,PCPG,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000278053 | DDX52 | 5 | ACC,KICH,KIRC,KIRP,SKCM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000278053 | DDX52 | 5 | ACC,KICH,KIRC,KIRP,SKCM | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000123268 | ATF1 | 7 | BRCA,KIRP,MESO,PCPG,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000123268 | ATF1 | 10 | BRCA,KICH,KIRP,MESO,PCPG,PRAD,TGCT,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000123268 | ATF1 | 11 | ACC,BRCA,CESC,CHOL,KIRC,KIRP,MESO,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000123268 | ATF1 | 8 | CHOL,KICH,KIRP,MESO,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000123268 | ATF1 | 8 | BRCA,KICH,KIRP,MESO,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000123268 | ATF1 | 8 | ACC,KICH,KIRC,KIRP,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000123268 | ATF1 | 12 | ACC,CESC,CHOL,KICH,KIRC,KIRP,MESO,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000123268 | ATF1 | 8 | ACC,CHOL,KIRC,KIRP,MESO,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000123268 | ATF1 | 8 | ACC,BRCA,KICH,KIRP,MESO,PCPG,PRAD,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000170365 | SMAD1 | 6 | KIRP,PCPG,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000170365 | SMAD1 | 7 | KIRP,PCPG,PRAD,TGCT,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000170365 | SMAD1 | 7 | KIRC,KIRP,PRAD,READ,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000116016 | EPAS1 | ENSG00000170365 | SMAD1 | 5 | COAD,GBM,READ,TGCT,THCA | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000170365 | SMAD1 | 5 | KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000170365 | SMAD1 | 6 | ACC,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000170365 | SMAD1 | 6 | ACC,KIRC,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000170365 | SMAD1 | 6 | ACC,COAD,PRAD,READ,TGCT,THCA | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000170365 | SMAD1 | 5 | KIRC,KIRP,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000170365 | SMAD1 | 8 | ACC,KIRC,KIRP,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000170365 | SMAD1 | 7 | ACC,KIRC,KIRP,PRAD,READ,THCA,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000065457 | ADAT1 | 6 | CESC,HNSC,KIRC,KIRP,SKCM,THCA | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000065457 | ADAT1 | 5 | KICH,KIRC,KIRP,SKCM,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000065457 | ADAT1 | 8 | CESC,HNSC,KICH,KIRC,KIRP,PCPG,SKCM,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000099995 | SF3A1 | 7 | BLCA,KIRP,MESO,PCPG,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000099995 | SF3A1 | 9 | ACC,CHOL,HNSC,KIRC,KIRP,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000099995 | SF3A1 | 7 | CHOL,KICH,KIRP,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000099995 | SF3A1 | 8 | ACC,HNSC,KICH,KIRP,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000099995 | SF3A1 | 5 | KICH,KIRC,KIRP,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000099995 | SF3A1 | 13 | ACC,BLCA,CHOL,HNSC,KICH,KIRC,KIRP,MESO,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000178764 | ZHX2 | ENSG00000099995 | SF3A1 | 6 | ACC,KICH,KIRP,MESO,PCPG,PRAD | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000099995 | SF3A1 | 9 | ACC,BLCA,CHOL,KIRC,KIRP,MESO,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000076067 | RBMS2 | 7 | BRCA,CHOL,KIRP,OV,PRAD,THYM,UCEC | | ENSG00000265519 | ENSG00000116016 | EPAS1 | ENSG00000076067 | RBMS2 | 5 | KICH,LUSC,PRAD,TGCT,THYM | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000076067 | RBMS2 | 6 | CHOL,KICH,KIRP,PRAD,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000076067 | RBMS2 | 7 | BRCA,KICH,KIRP,OV,PRAD,THYM,UCEC | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000076067 | RBMS2 | 5 | BRCA,KICH,LUSC,PRAD,TGCT | | ENSG00000265519 | ENSG00000169554 | ZEB2 | ENSG00000076067 | RBMS2 | 5 | KICH,LUSC,PRAD,TGCT,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000076067 | RBMS2 | 6 | CHOL,KICH,KIRP,PRAD,THYM,UCEC | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000135457 | TFCP2 | 6 | BRCA,KIRP,OV,PCPG,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000135457 | TFCP2 | 6 | CHOL,KIRC,KIRP,OV,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000135457 | TFCP2 | 5 | BRCA,KIRC,KIRP,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000135457 | TFCP2 | 7 | CHOL,KICH,KIRC,KIRP,PCPG,THCA,THYM | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000135457 | TFCP2 | 5 | BRCA,KICH,KIRP,PCPG,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000084093 | REST | 7 | BLCA,BRCA,KIRP,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000084093 | REST | 6 | BRCA,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000084093 | REST | 11 | ACC,BLCA,BRCA,HNSC,KIRC,KIRP,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000084093 | REST | 6 | KIRP,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000084093 | REST | 9 | ACC,BLCA,BRCA,HNSC,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000084093 | REST | 7 | ACC,KIRC,KIRP,PRAD,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000084093 | REST | 11 | ACC,BLCA,HNSC,KIRC,KIRP,PRAD,SKCM,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000084093 | REST | 8 | ACC,BLCA,KIRC,KIRP,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000084093 | REST | 6 | ACC,BRCA,KIRP,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000085274 | MYNN | 5 | ACC,KICH,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000136937 | NCBP1 | 6 | CHOL,KICH,KIRC,KIRP,PCPG,THCA | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000166889 | PATL1 | 5 | MESO,PRAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000166889 | PATL1 | 5 | CHOL,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000166889 | PATL1 | 5 | CHOL,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000131013 | PPIL4 | 5 | KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000131013 | PPIL4 | 8 | BLCA,CHOL,KIRC,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000131013 | PPIL4 | 7 | CHOL,KICH,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000131013 | PPIL4 | 7 | BLCA,CESC,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000131013 | PPIL4 | 6 | KICH,KIRC,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000131013 | PPIL4 | 5 | KICH,KIRC,KIRP,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000131013 | PPIL4 | 10 | BLCA,CESC,CHOL,KICH,KIRC,KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000131013 | PPIL4 | 6 | BLCA,KIRC,KIRP,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000140836 | ZFHX3 | 7 | CESC,HNSC,OV,PRAD,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000140836 | ZFHX3 | 6 | CESC,HNSC,OV,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000140836 | ZFHX3 | 7 | CESC,OV,SKCM,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000169554 | ZEB2 | ENSG00000140836 | ZFHX3 | 5 | HNSC,PRAD,SKCM,TGCT,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000140836 | ZFHX3 | 7 | CESC,HNSC,OV,PRAD,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000106571 | GLI3 | ENSG00000187792 | ZNF70 | 6 | KIRP,PRAD,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000187792 | ZNF70 | 5 | KIRP,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000134698 | AGO4 | 7 | BRCA,KICH,PRAD,TGCT,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000134698 | AGO4 | 7 | BRCA,CHOL,OV,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000134698 | AGO4 | 7 | CHOL,KICH,OV,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000134698 | AGO4 | 8 | ACC,BRCA,KICH,OV,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000134698 | AGO4 | 7 | ACC,CHOL,KICH,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000106571 | GLI3 | ENSG00000144642 | RBMS3 | 5 | PRAD,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000144642 | RBMS3 | 5 | BRCA,PRAD,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000116016 | EPAS1 | ENSG00000144642 | RBMS3 | 6 | LUSC,PRAD,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000140836 | ZFHX3 | ENSG00000144642 | RBMS3 | 5 | PRAD,SKCM,STAD,TGCT,THCA | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000144642 | RBMS3 | 6 | BRCA,LUSC,PRAD,STAD,TGCT,THCA | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000172667 | ZMAT3 | 9 | ACC,CHOL,HNSC,KIRC,KIRP,PRAD,SKCM,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000172667 | ZMAT3 | 6 | ACC,HNSC,KIRP,PRAD,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000172667 | ZMAT3 | 6 | ACC,KIRC,KIRP,MESO,PRAD,THYM | | ENSG00000265519 | ENSG00000169554 | ZEB2 | ENSG00000172667 | ZMAT3 | 5 | HNSC,KIRP,PRAD,TGCT,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000172667 | ZMAT3 | 9 | ACC,CHOL,HNSC,KIRC,KIRP,PRAD,SKCM,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000172667 | ZMAT3 | 5 | CHOL,KIRC,KIRP,PRAD,UCEC | | ENSG00000265519 | ENSG00000106571 | GLI3 | ENSG00000162599 | NFIA | 5 | PRAD,STAD,TGCT,THCA,THYM | | ENSG00000265519 | ENSG00000140836 | ZFHX3 | ENSG00000162599 | NFIA | 5 | PRAD,SKCM,STAD,TGCT,THCA | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000162599 | NFIA | 6 | BRCA,KICH,PRAD,STAD,TGCT,THCA | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000149187 | CELF1 | 5 | CHOL,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000149187 | CELF1 | 5 | CHOL,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000149187 | CELF1 | 5 | CHOL,KIRP,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000149187 | CELF1 | 5 | CHOL,KIRP,SARC,THCA,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000069667 | RORA | 5 | BRCA,KICH,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000069667 | RORA | 6 | ACC,BRCA,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000069667 | RORA | 7 | ACC,BRCA,KICH,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000069667 | RORA | 6 | ACC,KICH,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000069667 | RORA | 6 | ACC,KICH,KIRP,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000069667 | RORA | 6 | ACC,BRCA,KICH,KIRP,PRAD,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000065970 | FOXJ2 | 5 | KIRP,OV,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000065970 | FOXJ2 | 5 | KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000065970 | FOXJ2 | 5 | KIRP,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000186660 | ZFP91 | 5 | BLCA,KIRP,MESO,THCA,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000186660 | ZFP91 | 5 | KIRP,MESO,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000186660 | ZFP91 | 11 | ACC,BLCA,CHOL,KIRC,KIRP,MESO,PRAD,READ,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000186660 | ZFP91 | 7 | CHOL,KIRP,MESO,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000186660 | ZFP91 | 7 | ACC,BLCA,KIRP,MESO,PRAD,THCA,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000186660 | ZFP91 | 7 | ACC,KIRC,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000186660 | ZFP91 | 10 | ACC,BLCA,CHOL,KIRC,KIRP,MESO,PRAD,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000186660 | ZFP91 | 6 | ACC,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000189079 | ARID2 | 5 | BRCA,KIRP,MESO,PCPG,THCA | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000189079 | ARID2 | 7 | BRCA,KICH,KIRP,MESO,PCPG,PRAD,THCA | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000189079 | ARID2 | 7 | ACC,BRCA,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000189079 | ARID2 | 5 | KICH,KIRP,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000189079 | ARID2 | 7 | ACC,BRCA,KICH,KIRP,MESO,PRAD,THCA | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000189079 | ARID2 | 7 | ACC,KICH,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000189079 | ARID2 | 8 | ACC,KICH,KIRP,MESO,PCPG,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000189079 | ARID2 | 6 | ACC,KIRP,MESO,PRAD,SKCM,THCA | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000124214 | STAU1 | 5 | KIRP,MESO,PCPG,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000124214 | STAU1 | 5 | KIRP,MESO,PCPG,THCA,THYM | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000124214 | STAU1 | 7 | ACC,KIRC,KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000124214 | STAU1 | 6 | ACC,KIRC,KIRP,MESO,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000124214 | STAU1 | 8 | ACC,KIRP,MESO,PCPG,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000124214 | STAU1 | 6 | ACC,KIRC,KIRP,MESO,THCA,UCEC | | ENSG00000265519 | ENSG00000076108 | BAZ2A | ENSG00000186130 | ZBTB6 | 5 | KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000186130 | ZBTB6 | 6 | KICH,KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000186130 | ZBTB6 | 8 | ACC,CHOL,KIRC,KIRP,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000117000 | RLF | ENSG00000186130 | ZBTB6 | 8 | CHOL,KICH,KIRP,MESO,SKCM,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000118058 | KMT2A | ENSG00000186130 | ZBTB6 | 7 | ACC,KICH,KIRP,MESO,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000186130 | ZBTB6 | 8 | ACC,KICH,KIRC,KIRP,MESO,SKCM,THCA,THYM | | ENSG00000265519 | ENSG00000165821 | SALL2 | ENSG00000186130 | ZBTB6 | 5 | ACC,KICH,KIRP,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000186130 | ZBTB6 | 11 | ACC,CHOL,KICH,KIRC,KIRP,MESO,SKCM,THCA,THYM,UCEC,UCS | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000186130 | ZBTB6 | 8 | ACC,CHOL,KIRC,KIRP,MESO,SKCM,THCA,UCEC | | ENSG00000265519 | ENSG00000189079 | ARID2 | ENSG00000186130 | ZBTB6 | 5 | KICH,KIRP,MESO,SKCM,THCA | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000079785 | DDX1 | 6 | ACC,CHOL,KICH,PCPG,THCA,THYM | | ENSG00000265519 | ENSG00000148516 | ZEB1 | ENSG00000151702 | FLI1 | 5 | COAD,HNSC,KICH,LUSC,READ |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000167967 | E4F1 | 5 | KIRC,PRAD,READ,THYM,UCEC | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000126254 | RBM42 | 6 | KICH,MESO,PCPG,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000126254 | RBM42 | 5 | KICH,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000126254 | RBM42 | 6 | KICH,MESO,PCPG,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000102189 | EEA1 | ENSG00000126456 | IRF3 | 7 | BRCA,MESO,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000114439 | BBX | ENSG00000126456 | IRF3 | 7 | BRCA,KIRC,MESO,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000134852 | CLOCK | ENSG00000126456 | IRF3 | 5 | KIRC,MESO,PRAD,THCA,THYM | | ENSG00000265519 | ENSG00000171634 | BPTF | ENSG00000126456 | IRF3 | 6 | MESO,PCPG,PRAD,THCA,THYM,UCEC | | ENSG00000265519 | ENSG00000186660 | ZFP91 | ENSG00000126456 | IRF3 | 5 | KIRC,MESO,PRAD,THCA,UCEC |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation |
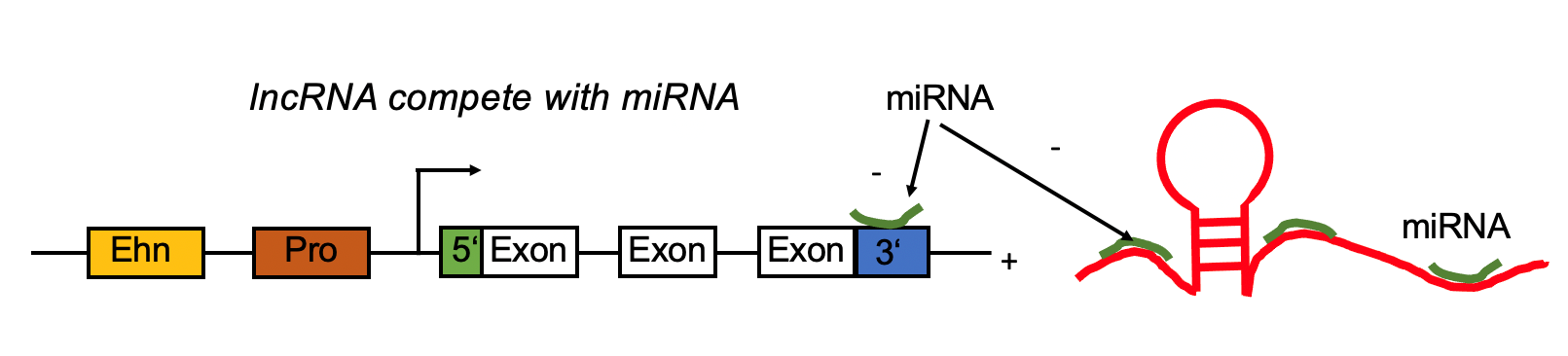
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,MFAP4 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,VCL | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CRISPLD2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,FERMT2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,MYLK | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CALD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,SSPN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PODN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,OLFML1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PTGIS | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,TGFB1I1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,DIXDC1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,DNAJB5 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PER1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,MRGPRF | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,ANTXR2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CYBRD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,GEM | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,RASL12 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PCDHGC3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,TCEAL7 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,SLIT3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,SMIM10 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,COPZ2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,ADGRA2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,MSRB3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,S1PR3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,AOC3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,DPT | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PRELP | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PLN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,OGN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,MYH11 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CNRIP1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CD34 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CPED1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,EHD2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,TAGLN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,TSC22D3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,SPART | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,MAP3K3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CAVIN2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,DCN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CLMP | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,EMILIN1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,DDR2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,SGCE | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PI16 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,EMCN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,PMP22 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CYYR1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,RHOJ | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,LMOD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,CAVIN1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,SH3BP5 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,TUBB6 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,ZCCHC24 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,LAYN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-96,JAM2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-3613,SLMAP | UCEC,THCA,PRAD,READ,KIRP | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-484,CRIM1 | UCEC,PRAD,ACC,GBM,THYM | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-484,MAP3K20 | UCEC,PRAD,ACC,GBM,THYM | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-484,ZYG11B | UCEC,PRAD,ACC,GBM,THYM | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,MYH11 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,CALD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,TGFB1I1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,MSRB3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,TNS1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,TSC22D3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,MRGPRF | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,RECK | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PER1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,CNRIP1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PLN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,FBXL7 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,ANTXR2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,ZCCHC24 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,DCN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,TCEAL7 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PRELP | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,SH3BP5 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,MFAP4 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,ADGRA2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,MYLK | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,EMP3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,SPART | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,EHD2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PMP22 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,SLIT3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PODN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,ACVRL1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,LMOD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,OLFML1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,TNFSF12 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,COPZ2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,LIMS2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PTGIS | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,DPT | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,COL6A2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,CPED1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,MMRN2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,TAGLN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,AOC3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,CRISPLD2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,COL14A1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,DNAJB5 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,FERMT2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,CAVIN2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,S1PR1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,JAM2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,SSPN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,LAYN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,PI16 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,CLMP | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,OGN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,EMILIN1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,DDR2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,SGCE | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-182,GEM | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,KANK2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,OGN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,TNFSF12 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,TSC22D3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,MRGPRF | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,COL14A1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,ADGRA2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,OLFML1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PTGIS | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,DPT | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,COL6A2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CYYR1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,TAGLN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,AOC3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,DNAJB5 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PER1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,JAM2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,GEM | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,FBXL7 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,TNS1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CALD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,COPZ2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,S1PR3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,MYLK | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,HIC1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,SSPN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,MYH11 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CPED1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,LIMS2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,SLIT3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,INMT | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,GPRASP2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CNRIP1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,SGCE | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,ANTXR2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,ZCCHC24 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,MFAP4 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,TGFB1I1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,SPART | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,EHD2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PODN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,VSIR | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,ACVRL1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,EMCN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,LMOD1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PI16 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,MMRN2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,SERPINF1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,EMILIN1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CRISPLD2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,FERMT2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CAVIN2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,S1PR1 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,TCEAL7 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,DCN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,CLMP | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,DDR2 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,SH3BP5 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,MSRB3 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,RECK | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PLN | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PRELP | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,PMP22 | UCEC,KICH,TGCT,PRAD,READ | | ENSG00000265519 | CTD-3157E16.1,hsa-mir-183,LAYN | UCEC,KICH,TGCT,PRAD,READ |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000265519.1 | ENST00000582780.1 | 57 | 216 | 389 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).