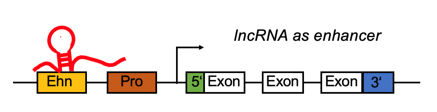
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | KCNMB1,PLIN4,USP22,TBC1D24,PADI2,CIRBP,PEBP1,MAPK8IP3,SUGP2,MDH2,ISCU,MTSS2,ACSS1,SLC25A12,ULK3,VPS39,GNAZ,PIK3R3,INPP5J,SAFB2,HIRIP3,ACAD8,RNF38,TMEM38A,NMNAT1,PBX3,TERF2IP,PYGM,SIN3A,PHYH,CDIP1,SPRN,SPG7,TACR2,CLCN6,GLRX5,RCCD1,IGIP,DUSP8,SEPTIN6,ACAD10,C3orf18,PDZD4,SGCA,TTC9,HCFC2,PAMR1,FEZ1,MYZAP,RAF1,IDH2,FGFR1,ZADH2,MAP6,GCDH,ZFP62,FOXF1,BAG1,MAP1A,RAP1GAP,PPP1R1A,COQ6,SECISBP2L,CHRDL1,ETFDH,LRRC37B,ZZEF1,IPO13,MRO,ADHFE1,DCUN1D2,ARHGEF4,HIF1AN,CBX6,NR2C2,PNMA1,LRTOMT,NT5DC3,COL4A6,NCALD,DMAC2L,DNASE1,ARL4D,ARHGEF7,HSPB7,TTC3,SIK3,AHDC1,RHCG,ATPAF1,ZFP3,CYFIP2,MEF2D,INSYN1,PITPNM2,CASTOR3,ARHGEF9,SHISAL1,SLC25A20,SH2B1,PNPLA4,UBE2G2,RRAGD,BVES,FAM168B,KLHL15,ARMCX4,MLLT6,NDUFS1,PDZRN3,NEXN,GRIK5,PHF7,COG8,ACYP2,APBB1,JPH2,SCO1,MYORG,TMEM175,DENND6A,POPDC2,ZBED3,SLC4A3,NPR2,CACNA1C,PI16,SPACA9,DACT3,CACNB1,NECAB3,MPP2,MTURN,PKD1,AMIGO1,PIDD1,CKMT2,PPM1L,CRYAB,PABIR1,TCTA,ARNT2,HAND2,ANKDD1A,EOLA1,FADS2,SLX4,FOXN3,LMOD1,CHRNB1,SMYD4,ZFYVE1,TARS3,LMF1,BRF1,PPP1R3F,UBOX5,ST3GAL3,CYB561D1,ZNF48,TMOD2,ACACB,INPP5E,PPM1K,IL11RA,CHD2,ACADSB,FOXP1,PDLIM3,GNAO1,ACTA2,CACNA1H,HERC1,GRAMD1B,TMEM35A,ZNF862,SBF1,ANXA6,BTBD6,LYRM7,SMAD4,CRAT,SFXN5,MYH11,ECI2,DENND2A,DECR1,CASQ2,ADAMTS8,CROCC,ATPAF2,ACAP3,BAG2,UNC119B,ZNF10,CPTP,ACADS,NISCH,DET1,SAR1B,IFFO1,ACTC1,PI4KA,PLCG1,CLEC3B,CCDC107,POMT1,ATXN7L2,FAM214A,UBTF,TNS1,SPARCL1,WDTC1,ADAM33,IRF2BPL,COQ3,BDH1,TTYH2,DMPK,CLK3,TOM1L2,HS6ST1,ZER1,USP11,PRKACA,KLHL25,OSBPL7,FYCO1,PBXIP1,RNF123,TCFL5,PDK2,ARHGEF3,RNF150,PKIA,ELP6,LNPEP,SEMA6C,SH3BGR,ANKRD46,FAM53B,CAVIN2,CIB2,PSKH1,ZHX3,SAP30L,MTHFR,HIRA,ACAA1,DEAF1,DBT,HERC2,ANKS3,YJEFN3,RCAN2,AOC3,USP19,DUSP15,TMEM201,SCMH1,SNTA1,EPM2A,C16orf89,R3HDM2,LYVE1,DEXI,GAS7,UBL3,RAB3A,RFX3,LETM1,CDK5R1,MOCS2,HSPB6,CDS2,POLG,NEIL1,MYO18A,ABHD10,DTX3,SELENBP1,REEP1,DYM,NDUFA5,WDR6,C3orf70,COQ9,CBX8,SMARCD3,KMT2A,TINAGL1,NCAM1,AP1S2,APBB3,TLN1,NDUFAF1,NRIP2,PARP16,RBM33,ADAL,WDR37,SEC22C,CAMKK2,TIMM21,GABPB2,MAPKBP1,SLC29A2,MAOB,VSTM4,RGMA,HSPB8,PTN,ITFG2,MAMDC2,ADGRL1,CCDC92,COQ8A,RNF10,GBA2,REEP2,TTLL7,ACO2,CILP,C1QL1,ATP5F1A,TUB,DCAF8,RPTOR,GREM2,FBXO10,RUFY3,SNX29,OGA,GORASP1,CLASP2,TBC1D17,GATA2,PPP1R3C,NRTN,CPXM2,PACS2,CFAP410,VAMP2,KAT6B,MPC1,RNF216,TECPR2,PCNT,BEND5,C21orf58,H2AZ2,RERG,NCOA5,TMEM143,PAFAH2,ANKRD9,ITIH5,TADA2B,LIPT2,DYRK1B,NNT,DGLUCY,NINL,SDR39U1,SORBS1,FABP3,NSMF,DIRAS1,STOML1,TIGD1,ADGRB2,GPS2,HADHB,USF3,MTFR1L,VEGFB,BAHCC1,FBXO31,ZMAT1,IRAG1,TAPT1,MACROD1,GAS6,PXMP2,NKTR,RIMS3,OXCT1,CASZ1,PITPNM3,ZNF84,GDPGP1,TUBA1A,NDUFV3,ALKBH5,B4GALT6,PIAS2,TSC1,ASB2,MAN2C1,USP20,ASB8,TRAPPC2 | | Exist in public source | | NA |
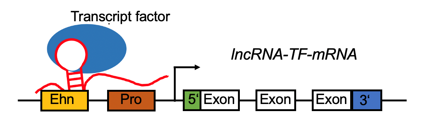
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | HSPB8,PBX3,SLC16A2,ALKBH5,RAB3A,SELENBP1,ADAM33,CHRDL1,LIPT2,DENND6A,ULK1,IDH2,ZHX3,IRF2BPL,EDNRA,TNS1,CRAT,SPEG,PPP1R12B,PRKG1,ZER1,MAOB,MLLT6,TBC1D1,TTYH2,BEND5,NTN1,CYFIP2,GRIK5,FOXF1,NPR2,KMT2A,UNC119B,GDF11,DCLK2,TCEAL2,INPP5J,RPP14,MAPK8IP3,KCTD7,VAMP2,GADD45G,PKD1,SH3BP2,SEMA6C,ARNT2,ZADH2,NEURL1B,NUDT7,TBKBP1,WWOX,PKDCC,RNF38,NT5M,NEXN,MAGEF1,SECISBP2L,HERC1,ARMCX4,KAT6B,PDZRN3,IPO13,SEPT6,ITPKB,MARK1,NAT8L,AKAP13,POMT1,ADAMTS8,CAMK1D,ETFDH,GSE1,MAMDC2,PI4KA | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
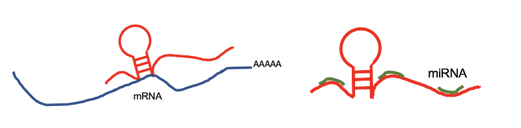
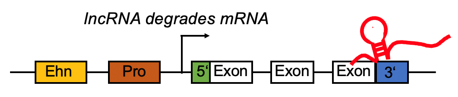
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | PSMB5,INTS7,PRDX4,DCTPP1,DENR,SEL1L3,EIF6,PTPN12,UTP18,ALG3,WDR46,PSMC4,SH2D3A,NRBP1,SRFBP1,HEXB,NFKB2,ATIC,S100A11,SLC15A4,LSR,PLAUR,GRN,NTMT1,MYDGF,YWHAZ,NMD3,ATG7,RNPEP,SURF4,SP100,YIF1B,PTPRE,HSPE1,CCT7,MOGS,PRELID1,ABCE1,NRBF2,AGTRAP,GBA,SMAGP,CASP4,MVP,VPS25,PSMD11,SLC52A2,NME1,FARSA,RAB7A,TNFRSF10A,SERPINH1,P4HB,PNO1,GLMP,TGIF1,LY6E,KLF16,AUP1,SERP1,LMAN2,THOC3,NHP2,C1orf43,TMEM41A,PYGL,PSME3,NOB1,TMSB10,RHBDF2,ORMDL2 | | Exist in public source | | NA |
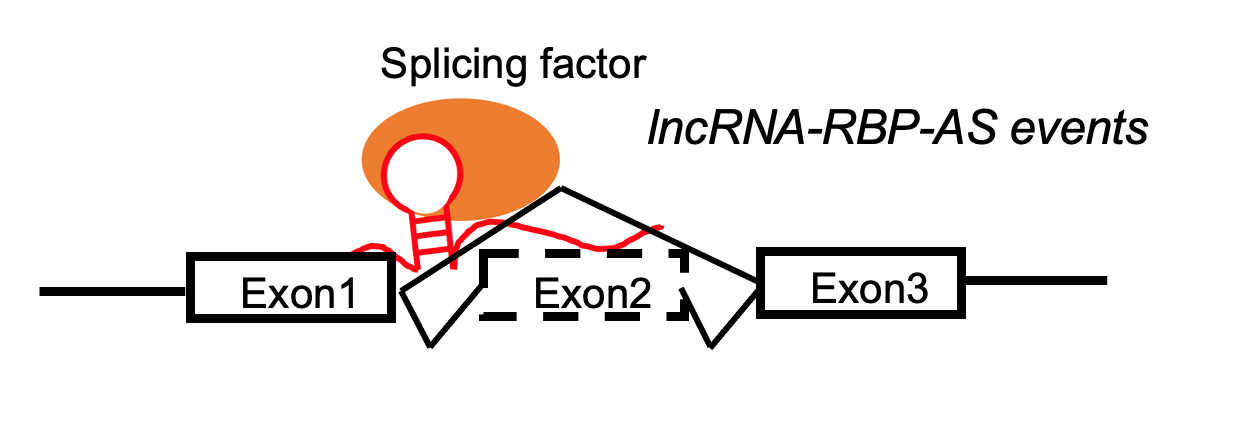
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000259291 | ENST00000558334 | exon_skip_7558 | chr1:81952986-81953025 | -6.15 | -0.1662 | ADGRL2 | ENST00000370717,ENST00000370728 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_40119 | chr10:24494499-24494604 | -11.96 | -0.1359 | KIAA1217 | ENST00000376454 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_42594 | chr10:74111908-74112112 | -25.75 | -0.1497 | VCL | ENST00000211998 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_43507 | chr10:84499874-84499959 | -15.38 | -0.2902 | CCSER2 | ENST00000224756 | Frame-shift | | ENSG00000259291 | ENST00000558334 | exon_skip_48473 | chr10:29526960-29527056 | -15.15 | -0.1702 | SVIL | ENST00000355867 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_125679 | chr15:29719776-29720016 | -22.36 | -0.1308 | TJP1 | ENST00000346128 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_293617 | chr17:76091142-76091235 | -11.68 | -0.1256 | EXOC7 | ENST00000589210 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_300058 | chr18:79933968-79934022 | -10.89 | -0.2533 | PQLC1 | ENST00000397778 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_301267 | chr19:6746028-6746196 | -27.83 | -0.1666 | TRIP10 | ENST00000313244 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_335378 | chr2:237751199-237751271 | -9.76 | -0.1457 | LRRFIP1 | ENST00000392000 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_453010 | chr6:70791688-70791769 | -12.70 | -0.1649 | SMAP1 | ENST00000370455 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_506938 | chr9:125506309-125506417 | -16.50 | -0.1964 | MAPKAP1 | ENST00000265960,ENST00000373498 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_507736 | chr9:129915965-129915980 | -4.29 | -2.1450 | FNBP1 | ENST00000446176 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_515330 | chrX:65524563-65524614 | -10.14 | -0.2305 | LAS1L | ENST00000374811 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_293641 | chr17:76091142-76091235 | -11.68 | -0.1256 | EXOC7 | ENST00000589210 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_322579 | chr19:55455635-55455845 | -28.52 | -0.1698 | ISOC2 | ENST00000425675 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_332068 | chr2:191402631-191402718 | -17.11 | -0.3111 | MYO1B | ENST00000304164,ENST00000392318 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000259291 | ENST00000558334 | exon_skip_83404 | chr12:53465931-53465973 | -8.88 | -0.2691 | PCBP2 | ENST00000439930 | In-frame | | ENSG00000259291 | ENST00000620791 | exon_skip_95538 | chr12:98634295-98634413 | -11.83 | -0.1160 | IKBIP | ENST00000342502 | Frame-shift | | ENSG00000259291 | ENST00000558334 | exon_skip_106783 | chr14:55673171-55673255 | -10.24 | -0.1625 | KTN1 | ENST00000395314 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_132355 | chr16:1786756-1786955 | -27.84 | -0.1538 | NUBP2 | ENST00000262302 | Frame-shift | | ENSG00000259291 | ENST00000558334 | exon_skip_287890 | chr17:29116443-29116455 | -5.29 | -1.3225 | MYO18A | ENST00000527372 | In-frame | | ENSG00000259291 | ENST00000558334 | exon_skip_64939 | chr11:112230207-112230230 | -6.69 | -0.8362 | PTS | ENST00000280362 | Frame-shift | | ENSG00000259291 | ENST00000558334 | exon_skip_345955 | chr2:200860124-200860215 | -18.47 | -0.2757 | CLK1 | ENST00000321356 | Frame-shift | | ENSG00000259291 | ENST00000558334 | exon_skip_446942 | chr5:178617343-178617434 | -18.39 | -0.2590 | CLK4 | ENST00000316308 | Frame-shift | | ENSG00000259291 | ENST00000558334 | exon_skip_302276 | chr19:10808568-10808580 | -7.80 | -0.8667 | DNM2 | ENST00000355667 | In-frame |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
 |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90076297-90076371 | 186.00 | -63.03 | 74 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90074788-90074864 | 167.00 | -66.93 | 74 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075135-90075211 | 164.00 | -67.34 | 75 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075549-90075618 | 163.00 | -69.05 | 68 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075996-90076056 | 160.00 | -58.31 | 69 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075030-90075101 | 159.00 | -63.21 | 64 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90076517-90076588 | 154.00 | -61.09 | 64 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075291-90075364 | 152.00 | -73.58 | 73 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075417-90075491 | 151.00 | -58.61 | 70 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90074959-90075041 | 150.00 | -68.28 | 77 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075213-90075279 | 150.00 | -57.59 | 68 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90074547-90074622 | 147.00 | -58.83 | 74 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075704-90075770 | 143.00 | -57.01 | 64 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075800-90075865 | 142.00 | -62.05 | 67 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90074652-90074736 | 141.00 | -68.35 | 75 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90075610-90075684 | 141.00 | -61.05 | 73 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000620791 | chr15:90076188-90076253 | 141.00 | -66.51 | 64 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000558334 | chr15:90075338-90075412 | 186.00 | -63.03 | 74 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000558334 | chr15:90074629-90074701 | 170.00 | -54.09 | 71 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000558334 | chr15:90074855-90074923 | 156.00 | -72.89 | 70 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000558334 | chr15:90075558-90075629 | 154.00 | -61.09 | 64 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000558334 | chr15:90075026-90075108 | 150.00 | -74.55 | 78 | NA | | ENSG00000259291 | hsa-mir-21 | ENST00000558334 | chr15:90075132-90075211 | 144.00 | -74.19 | 78 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90076447-90076526 | 214.00 | -64.66 | 79 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075637-90075713 | 196.00 | -69.34 | 73 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90074893-90074961 | 184.00 | -52.70 | 65 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075241-90075316 | 176.00 | -70.47 | 73 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075946-90076015 | 171.00 | -64.74 | 76 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90076233-90076314 | 171.00 | -67.05 | 76 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90074664-90074746 | 166.00 | -71.62 | 82 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075793-90075868 | 162.00 | -63.15 | 70 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90076078-90076168 | 161.00 | -68.17 | 84 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90074579-90074652 | 159.00 | -63.18 | 73 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075124-90075203 | 158.00 | -79.49 | 79 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90074944-90075025 | 154.00 | -61.70 | 69 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075859-90075942 | 146.00 | -62.18 | 64 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90074742-90074821 | 144.00 | -66.04 | 74 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000620791 | chr15:90075550-90075626 | 140.00 | -67.96 | 67 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90075488-90075567 | 214.00 | -64.66 | 79 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90074987-90075062 | 180.00 | -65.89 | 74 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90075111-90075188 | 177.00 | -77.32 | 75 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90075183-90075277 | 172.00 | -87.11 | 85 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90074731-90074803 | 170.00 | -68.06 | 78 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90075274-90075355 | 166.00 | -70.93 | 75 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90074626-90074716 | 155.00 | -61.08 | 78 | NA | | ENSG00000259291 | hsa-mir-27a | ENST00000558334 | chr15:90074847-90074930 | 142.00 | -84.61 | 68 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90076099-90076169 | 183.00 | -56.38 | 60 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075460-90075531 | 163.00 | -58.33 | 64 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90076245-90076313 | 160.00 | -55.88 | 59 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90076348-90076421 | 160.00 | -57.01 | 72 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075780-90075850 | 159.00 | -56.37 | 58 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075262-90075326 | 156.00 | -69.80 | 63 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90074896-90074962 | 153.00 | -50.31 | 67 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075929-90075990 | 150.00 | -56.51 | 67 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075554-90075626 | 149.00 | -65.59 | 54 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075646-90075714 | 149.00 | -63.14 | 63 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90076447-90076519 | 148.00 | -53.88 | 65 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90074742-90074822 | 146.00 | -68.45 | 77 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075169-90075238 | 146.00 | -62.33 | 67 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90074682-90074747 | 144.00 | -59.47 | 64 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90075317-90075383 | 144.00 | -61.13 | 60 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000620791 | chr15:90074585-90074665 | 142.00 | -64.50 | 79 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90074642-90074717 | 174.00 | -53.79 | 75 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90075207-90075278 | 167.00 | -68.39 | 71 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90074730-90074802 | 163.00 | -69.87 | 74 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90075286-90075354 | 160.00 | -55.88 | 59 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90075389-90075462 | 160.00 | -57.01 | 72 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90075113-90075187 | 156.00 | -71.16 | 73 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90075488-90075560 | 148.00 | -53.88 | 65 | NA | | ENSG00000259291 | hsa-mir-31 | ENST00000558334 | chr15:90074988-90075061 | 140.00 | -62.42 | 71 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90074715-90074812 | 190.00 | -80.11 | 95 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075409-90075492 | 183.00 | -72.65 | 86 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075984-90076061 | 182.00 | -79.30 | 83 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075533-90075619 | 177.00 | -81.12 | 87 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075827-90075924 | 173.00 | -90.94 | 92 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075321-90075405 | 169.00 | -77.30 | 82 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075624-90075702 | 165.00 | -72.32 | 80 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90076469-90076562 | 163.00 | -79.01 | 71 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075080-90075169 | 162.00 | -85.35 | 90 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90076064-90076163 | 161.00 | -74.98 | 97 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90074889-90074981 | 158.00 | -72.69 | 90 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90074614-90074697 | 154.00 | -82.32 | 84 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90075222-90075304 | 147.00 | -82.77 | 82 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90076214-90076306 | 145.00 | -76.31 | 93 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000620791 | chr15:90074513-90074593 | 141.00 | -79.30 | 81 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90074659-90074749 | 202.00 | -94.92 | 88 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90074928-90075019 | 171.00 | -88.55 | 84 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90075510-90075603 | 163.00 | -79.01 | 71 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90074784-90074864 | 162.00 | -76.34 | 74 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90075125-90075204 | 153.00 | -80.11 | 83 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90075045-90075126 | 142.00 | -80.04 | 85 | NA | | ENSG00000259291 | hsa-mir-3614 | ENST00000558334 | chr15:90074551-90074638 | 140.00 | -65.60 | 88 | NA | | ENSG00000259291 | hsa-mir-29a | ENST00000620791 | chr15:90075491-90075563 | 143.00 | -50.10 | 70 | NA | | ENSG00000259291 | hsa-mir-29a | ENST00000558334 | chr15:90075134-90075192 | 146.00 | -54.80 | 61 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075652-90075733 | 196.00 | -71.56 | 78 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90074815-90074885 | 190.00 | -52.59 | 77 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075159-90075241 | 190.00 | -78.04 | 85 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90076048-90076125 | 189.00 | -61.46 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90076418-90076499 | 186.00 | -53.74 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90074966-90075041 | 183.00 | -64.19 | 80 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075768-90075850 | 172.00 | -62.14 | 80 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90074660-90074747 | 170.00 | -72.47 | 86 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075571-90075652 | 170.00 | -69.90 | 79 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90074882-90074962 | 168.00 | -55.24 | 83 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075371-90075447 | 168.00 | -52.95 | 68 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075882-90075964 | 163.00 | -63.74 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90074527-90074622 | 161.00 | -68.59 | 92 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90076136-90076220 | 160.00 | -65.99 | 84 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075252-90075336 | 156.00 | -85.64 | 83 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000620791 | chr15:90075492-90075569 | 148.00 | -65.65 | 78 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000558334 | chr15:90075459-90075540 | 186.00 | -53.74 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000558334 | chr15:90075115-90075196 | 181.00 | -77.21 | 77 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000558334 | chr15:90074588-90074683 | 171.00 | -60.26 | 95 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000558334 | chr15:90074799-90074876 | 163.00 | -71.58 | 78 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000558334 | chr15:90074980-90075060 | 145.00 | -66.83 | 74 | NA | | ENSG00000259291 | hsa-mir-29b-1 | ENST00000558334 | chr15:90075276-90075354 | 142.00 | -68.19 | 76 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90074816-90074904 | 201.00 | -56.22 | 84 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075848-90075931 | 198.00 | -64.59 | 82 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075086-90075168 | 187.00 | -74.59 | 84 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075451-90075536 | 181.00 | -66.50 | 86 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075372-90075451 | 179.00 | -51.45 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90074687-90074767 | 169.00 | -52.23 | 80 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90076061-90076146 | 167.00 | -56.48 | 87 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90076417-90076502 | 165.00 | -56.25 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075196-90075288 | 163.00 | -65.64 | 91 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90076504-90076579 | 163.00 | -60.39 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075698-90075792 | 160.00 | -62.84 | 92 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90074928-90075013 | 159.00 | -58.48 | 87 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90074546-90074622 | 157.00 | -55.71 | 80 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000620791 | chr15:90075533-90075618 | 152.00 | -68.35 | 79 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90074606-90074683 | 195.00 | -57.70 | 83 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90075073-90075157 | 166.00 | -73.87 | 83 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90075458-90075543 | 165.00 | -56.25 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90075545-90075620 | 163.00 | -60.39 | 81 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90074979-90075068 | 161.00 | -65.71 | 85 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90074860-90074945 | 153.00 | -85.16 | 79 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90074768-90074848 | 148.00 | -69.95 | 79 | NA | | ENSG00000259291 | hsa-mir-29b-2 | ENST00000558334 | chr15:90074693-90074773 | 144.00 | -67.12 | 79 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000259291 | ENSG00000004776 | HSPB6 | BLCA,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000007237 | GAS7 | BLCA,COAD,DLBC,LUSC,READ,STAD | | BLCA,COAD,READ | | ENSG00000259291 | ENSG00000008710 | PKD1 | BLCA,CESC,CHOL,COAD,GBM,HNSC,KICH,LUSC,MESO,OV,PAAD,PRAD,SARC,STAD,TGCT | KICH | BLCA,GBM | | ENSG00000259291 | ENSG00000010295 | IFFO1 | BLCA,HNSC,PAAD,STAD,THYM,UVM | | BLCA | | ENSG00000259291 | ENSG00000011021 | CLCN6 | BLCA,BRCA,CHOL,COAD,GBM,LUSC,PAAD | | GBM | | ENSG00000259291 | ENSG00000053702 | NRIP2 | BLCA,BRCA,CHOL,DLBC,GBM,KIRP,LAML,LIHC,LUSC,MESO,PAAD,PCPG,PRAD,STAD,TGCT,THCA | | BLCA,GBM,KIRP | | ENSG00000259291 | ENSG00000069535 | MAOB | BLCA,COAD,DLBC,READ,SARC,STAD,TGCT,THCA | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000072952 | IRAG1 | BLCA,COAD,GBM,KIRP,LGG,PAAD,READ,SARC,STAD,TGCT | | BLCA,COAD,GBM,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000075073 | TACR2 | BLCA,COAD,PAAD,READ,STAD | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000076555 | ACACB | BLCA,COAD,ESCA,GBM,KIRC,KIRP,PAAD,PRAD,READ,STAD,TGCT,THCA,UVM | | BLCA,COAD,ESCA,GBM,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000079308 | TNS1 | BLCA,CHOL,COAD,HNSC,KIRP,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,HNSC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000088543 | C3orf18 | BLCA,KICH,KIRC,KIRP,PAAD,READ,SARC,STAD,TGCT,THCA | KICH | BLCA,READ,STAD | | ENSG00000259291 | ENSG00000090975 | PITPNM2 | BLCA,PRAD,STAD,THYM,UVM | | BLCA | | ENSG00000259291 | ENSG00000095637 | SORBS1 | BLCA,CHOL,COAD,ESCA,GBM,HNSC,PAAD,READ,SARC,STAD,TGCT | | BLCA,COAD,ESCA,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000100628 | ASB2 | BLCA,COAD,HNSC,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000101290 | CDS2 | BLCA,GBM,KICH,LAML,PAAD,PRAD,SARC,STAD,UVM | | GBM | | ENSG00000259291 | ENSG00000101400 | SNTA1 | BLCA,GBM,HNSC,KICH,LGG,PAAD,SARC,THCA | KICH | BLCA,HNSC | | ENSG00000259291 | ENSG00000101938 | CHRDL1 | BLCA,COAD,HNSC,PRAD,READ,STAD,TGCT | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000103264 | FBXO31 | BLCA,DLBC,HNSC,KICH,PAAD,PRAD,STAD | KICH | BLCA | | ENSG00000259291 | ENSG00000103657 | HERC1 | BLCA,CESC,CHOL,COAD,GBM,HNSC,KIRP,LUSC,MESO,PAAD,READ,STAD | | BLCA,GBM | | ENSG00000259291 | ENSG00000104490 | NCALD | BLCA,CHOL,COAD,PAAD,STAD,TGCT | | BLCA,STAD | | ENSG00000259291 | ENSG00000104936 | DMPK | BLCA,COAD,GBM,HNSC,KICH,MESO,PAAD,PRAD,READ,SARC,STAD,TGCT,THCA,UCS | KICH | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000107796 | ACTA2 | BLCA,COAD,SARC,STAD,TGCT,UVM | | BLCA,COAD,STAD | | ENSG00000259291 | ENSG00000108823 | SGCA | BLCA,HNSC,PAAD,PRAD,READ,SARC,STAD,UCS | | BLCA,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000108852 | MPP2 | BLCA,GBM,PAAD,PRAD,SARC,THCA | | BLCA,GBM | | ENSG00000259291 | ENSG00000109846 | CRYAB | BLCA,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000111696 | NT5DC3 | BLCA,COAD,READ,SARC,STAD,UVM | | BLCA,READ | | ENSG00000259291 | ENSG00000111727 | HCFC2 | BLCA,CHOL,READ,STAD,TGCT,UCS | | READ | | ENSG00000259291 | ENSG00000112208 | BAG2 | BLCA,COAD,PAAD,READ,SARC,STAD,UCS | | BLCA,READ,STAD | | ENSG00000259291 | ENSG00000112276 | BVES | BLCA,ESCA,HNSC,SARC,STAD,UCS | | BLCA,ESCA,STAD | | ENSG00000259291 | ENSG00000115840 | SLC25A12 | BLCA,COAD,HNSC,KIRC,PAAD,PRAD,SARC,STAD | | BLCA,HNSC | | ENSG00000259291 | ENSG00000117640 | MTFR1L | BLCA,COAD,DLBC,ESCA,HNSC,KIRC,KIRP,PAAD,READ,SARC,STAD,TGCT,THCA,UCS | | ESCA,HNSC,STAD | | ENSG00000259291 | ENSG00000118729 | CASQ2 | BLCA,COAD,HNSC,READ,SARC,STAD,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000121440 | PDZRN3 | BLCA,COAD,PAAD,READ,STAD,TGCT,UCS | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000121577 | POPDC2 | BLCA,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,THCA,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000121898 | CPXM2 | BLCA,COAD,PAAD,READ,SARC,STAD | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000126091 | ST3GAL3 | BLCA,COAD,DLBC,HNSC,PAAD,READ,STAD,TGCT,UCS | | BLCA,READ,STAD | | ENSG00000259291 | ENSG00000126950 | TMEM35A | BLCA,COAD,KICH,PRAD,READ,SARC,STAD,TGCT | KICH | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000128266 | GNAZ | BLCA,ESCA,PAAD,PRAD,READ,SARC,STAD,UVM | | BLCA,ESCA,READ,STAD | | ENSG00000259291 | ENSG00000131089 | ARHGEF9 | BLCA,CHOL,GBM,KIRP,LAML,MESO,PAAD,PRAD,READ,SARC,STAD,TGCT,THCA | | BLCA,GBM,READ | | ENSG00000259291 | ENSG00000131471 | AOC3 | BLCA,COAD,HNSC,KIRP,PAAD,PRAD,READ,SARC,STAD,TGCT | | BLCA,COAD,HNSC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000132563 | REEP2 | BLCA,GBM,KICH,PAAD,SARC,STAD,TGCT | KICH | BLCA,GBM,STAD | | ENSG00000259291 | ENSG00000133392 | MYH11 | BLCA,COAD,KIRP,PAAD,PRAD,READ,SARC,STAD,TGCT | | BLCA,COAD,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000133800 | LYVE1 | BLCA,COAD,HNSC,KIRP,PAAD,READ,STAD,TGCT | | BLCA,COAD,HNSC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000134533 | RERG | BLCA,COAD,READ,STAD,TGCT | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000136878 | USP20 | BLCA,CHOL,GBM,PAAD,SARC,STAD,THYM | | GBM | | ENSG00000259291 | ENSG00000137070 | IL11RA | BLCA,COAD,DLBC,PAAD,READ,SKCM,STAD,TGCT,THCA | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000137075 | RNF38 | BLCA,CHOL,GBM,READ,SARC,STAD | | BLCA | | ENSG00000259291 | ENSG00000137802 | MAPKBP1 | BLCA,CHOL,COAD,GBM,KICH,KIRP,PCPG,SARC,TGCT,UVM | | BLCA,GBM | | ENSG00000259291 | ENSG00000138615 | CILP | BLCA,COAD,DLBC,HNSC,READ,STAD | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000145936 | KCNMB1 | BLCA,COAD,PAAD,READ,SARC,STAD,TGCT | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000146966 | DENND2A | BLCA,KIRP,STAD,TGCT,UCS | | BLCA,KIRP,STAD | | ENSG00000259291 | ENSG00000149090 | PAMR1 | BLCA,CHOL,COAD,SKCM,TGCT,THCA | | BLCA,COAD | | ENSG00000259291 | ENSG00000149294 | NCAM1 | BLCA,GBM,PAAD,PRAD,STAD,TGCT,THCA,UCS | | BLCA,GBM,STAD | | ENSG00000259291 | ENSG00000149451 | ADAM33 | BLCA,MESO,PAAD,PRAD,STAD,TGCT | | BLCA,STAD | | ENSG00000259291 | ENSG00000149596 | JPH2 | BLCA,COAD,HNSC,KICH,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | KICH | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000152102 | FAM168B | BLCA,CHOL,COAD,GBM,KIRC,KIRP,MESO,PAAD,READ,SARC,STAD,TGCT | | BLCA,GBM | | ENSG00000259291 | ENSG00000152137 | HSPB8 | BLCA,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,THCA,UCS,UVM | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000152583 | SPARCL1 | BLCA,COAD,HNSC,LGG,PAAD,READ,STAD,TGCT,THCA | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000153446 | C16orf89 | BLCA,ESCA,KICH,KIRC,KIRP,TGCT,THCA | KICH | BLCA,ESCA,KIRC,KIRP | | ENSG00000259291 | ENSG00000154553 | PDLIM3 | BLCA,BRCA,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000156650 | KAT6B | BLCA,CHOL,COAD,LAML,READ | | BLCA | | ENSG00000259291 | ENSG00000159251 | ACTC1 | BLCA,PRAD,SARC,TGCT,UCS | | BLCA | | ENSG00000259291 | ENSG00000159884 | CCDC107 | BLCA,DLBC,SARC,STAD,THYM | | STAD | | ENSG00000259291 | ENSG00000159899 | NPR2 | BLCA,COAD,DLBC,READ,STAD,TGCT,THCA | | BLCA | | ENSG00000259291 | ENSG00000162614 | NEXN | BLCA,CHOL,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000163050 | COQ8A | BLCA,CHOL,ESCA,HNSC,KIRP | | ESCA,HNSC | | ENSG00000259291 | ENSG00000163431 | LMOD1 | BLCA,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT | | BLCA,COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000163590 | PPM1L | BLCA,COAD,GBM,HNSC,KIRC,KIRP,READ,SARC,STAD,THCA | | BLCA,GBM,HNSC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000163644 | PPM1K | BLCA,GBM,HNSC,KIRC,KIRP,LAML,LGG,STAD,THYM,UVM | | BLCA,GBM,KIRC,KIRP,STAD | | ENSG00000259291 | ENSG00000163815 | CLEC3B | BLCA,COAD,ESCA,HNSC,KIRP,PAAD,READ,STAD,THCA | | BLCA,COAD,ESCA,HNSC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000163820 | FYCO1 | BLCA,CHOL,COAD,HNSC,KIRC,PAAD,PRAD,READ,SARC,STAD,TGCT | | BLCA,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000164530 | PI16 | BLCA,COAD,READ,STAD,TGCT | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000166313 | APBB1 | BLCA,COAD,GBM,HNSC,PAAD,READ,SARC,STAD,TGCT,THYM | | BLCA,COAD,GBM,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000166963 | MAP1A | BLCA,COAD,GBM,PAAD,PRAD,READ,STAD,THYM,UVM | | BLCA,COAD,GBM,READ,STAD | | ENSG00000259291 | ENSG00000167676 | PLIN4 | BLCA,HNSC,PRAD,READ,SARC,STAD | | BLCA,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000168497 | CAVIN2 | BLCA,COAD,ESCA,PAAD,READ,SARC,STAD,TGCT | | BLCA,COAD,ESCA,READ,STAD | | ENSG00000259291 | ENSG00000171119 | NRTN | BLCA,ESCA,KICH,PAAD,STAD,TGCT | | BLCA,ESCA,STAD | | ENSG00000259291 | ENSG00000172348 | RCAN2 | BLCA,HNSC,KIRC,PAAD,READ,STAD,TGCT,THCA,UVM | | BLCA,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000173641 | HSPB7 | BLCA,COAD,HNSC,KIRC,KIRP,PAAD,PRAD,READ,STAD,TGCT,UCS | | BLCA,COAD,HNSC,KIRC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000174010 | KLHL15 | BLCA,CHOL,KIRC,KIRP,STAD | | BLCA,KIRP | | ENSG00000259291 | ENSG00000176903 | PNMA1 | BLCA,COAD,PRAD,READ,SARC,STAD | | BLCA,COAD,READ | | ENSG00000259291 | ENSG00000180354 | MTURN | BLCA,CHOL,COAD,ESCA,GBM,HNSC,KIRC,KIRP,LAML,PAAD,READ,STAD,THCA | | BLCA,ESCA,GBM,HNSC,KIRC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000181754 | AMIGO1 | BLCA,CHOL,GBM,KIRP,PAAD,READ,STAD,UCS | | GBM | | ENSG00000259291 | ENSG00000182287 | AP1S2 | BLCA,HNSC,PAAD,STAD,TGCT,THYM | | STAD | | ENSG00000259291 | ENSG00000182700 | IGIP | BLCA,CESC,CHOL,COAD,ESCA,GBM,HNSC,LGG,PAAD,READ,STAD | | BLCA,ESCA,GBM | | ENSG00000259291 | ENSG00000183741 | CBX6 | BLCA,CHOL,COAD,MESO,PAAD,PRAD,SARC,STAD,TGCT | | BLCA,COAD | | ENSG00000259291 | ENSG00000184545 | DUSP8 | BLCA,GBM,KICH,KIRC,KIRP,SARC | KICH | BLCA,GBM | | ENSG00000259291 | ENSG00000185418 | TARS3 | BLCA,BRCA,GBM,HNSC,LAML,LIHC,STAD,TGCT,UCS | | BLCA,GBM | | ENSG00000259291 | ENSG00000185437 | SH3BGR | BLCA,COAD,ESCA,HNSC,KIRC,KIRP,PAAD,PRAD,READ,SARC,STAD,THCA,UCS,UVM | | BLCA,COAD,ESCA,HNSC,KIRC,READ,STAD | | ENSG00000259291 | ENSG00000187068 | C3orf70 | BLCA,COAD,PAAD,PRAD,READ,SARC,STAD | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000196557 | CACNA1H | BLCA,COAD,KIRP,PAAD,PRAD,READ,SARC,STAD,TGCT,UCS | | BLCA,COAD,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000196663 | TECPR2 | BLCA,CHOL,GBM,PAAD,STAD,TGCT | | GBM | | ENSG00000259291 | ENSG00000197043 | ANXA6 | BLCA,HNSC,PAAD,SARC,STAD,THCA,THYM,UCS | | BLCA | | ENSG00000259291 | ENSG00000197380 | DACT3 | BLCA,COAD,MESO,PAAD,PRAD,READ,SARC,STAD,TGCT | | BLCA,COAD,READ,STAD | | ENSG00000259291 | ENSG00000197565 | COL4A6 | BLCA,PRAD,SARC,STAD,TGCT | | BLCA,STAD | | ENSG00000259291 | ENSG00000220205 | VAMP2 | BLCA,CHOL,COAD,DLBC,ESCA,GBM,KIRP,PAAD,READ,STAD,TGCT | | BLCA,COAD,ESCA,GBM,READ,STAD | | ENSG00000259291 | ENSG00000263155 | MYZAP | BLCA,ESCA,PRAD,READ,SARC | | BLCA,ESCA,READ | | ENSG00000259291 | ENSG00000112425 | EPM2A | BRCA,GBM,KIRC,KIRP,LAML,LGG,SARC,STAD,THCA,THYM,UCS | | KIRC,KIRP,STAD | | ENSG00000259291 | ENSG00000117016 | RIMS3 | BRCA,COAD,ESCA,SARC,THYM | | COAD,ESCA | | ENSG00000259291 | ENSG00000131730 | CKMT2 | BRCA,HNSC,KIRC,KIRP,LAML,PRAD,SARC,THCA,UCS | | HNSC,KIRP | | ENSG00000259291 | ENSG00000074755 | ZZEF1 | CESC,CHOL,COAD,GBM,LIHC,LUSC,PAAD,READ | | COAD,READ | | ENSG00000259291 | ENSG00000128731 | HERC2 | CESC,CHOL,GBM,LAML,LIHC,PAAD,SARC,UVM | | GBM | | ENSG00000259291 | ENSG00000137941 | TTLL7 | CESC,ESCA,GBM,PAAD,SARC,STAD | | ESCA,GBM,STAD | | ENSG00000259291 | ENSG00000138617 | PARP16 | CESC,DLBC,KICH,LAML,SARC | KICH | | | ENSG00000259291 | ENSG00000144040 | SFXN5 | CESC,DLBC,KIRC,KIRP,LGG,PAAD | | KIRC,KIRP | | ENSG00000259291 | ENSG00000147576 | ADHFE1 | CESC,GBM,KIRC,LGG,SARC | | GBM,KIRC | | ENSG00000259291 | ENSG00000160299 | PCNT | CESC,CHOL,GBM,HNSC,OV,PAAD,THYM | | GBM | | ENSG00000259291 | ENSG00000165699 | TSC1 | CESC,CHOL,COAD,GBM,PAAD | | GBM | | ENSG00000259291 | ENSG00000178188 | SH2B1 | CESC,DLBC,GBM,KICH,OV,TGCT,THCA | KICH | | | ENSG00000259291 | ENSG00000182054 | IDH2 | CESC,CHOL,DLBC,ESCA,GBM,HNSC,KICH,KIRC,KIRP,LAML,LGG,LIHC,LUAD,LUSC,MESO,OV,PAAD,PCPG,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC,UCS,UVM | KICH,PCPG | ESCA,HNSC,KIRC,KIRP | | ENSG00000259291 | ENSG00000183655 | KLHL25 | CESC,CHOL,KICH,LGG,SARC,SKCM,THYM | KICH | | | ENSG00000259291 | ENSG00000186532 | SMYD4 | CESC,COAD,KICH,PAAD,THCA | KICH | | | ENSG00000259291 | ENSG00000188827 | SLX4 | CESC,GBM,KICH,THYM,UVM | KICH | | | ENSG00000259291 | ENSG00000198408 | OGA | CESC,DLBC,GBM,LAML,THYM | | GBM | | ENSG00000259291 | ENSG00000213918 | DNASE1 | CESC,CHOL,DLBC,GBM,KICH,KIRC,KIRP,LAML,MESO,TGCT,THYM | KICH | KIRC,KIRP | | ENSG00000259291 | ENSG00000275023 | MLLT6 | CESC,GBM,HNSC,LUSC,SARC | | GBM | | ENSG00000259291 | ENSG00000005882 | PDK2 | CHOL,GBM,HNSC,KIRP,PRAD,SARC,TGCT | | GBM,HNSC | | ENSG00000259291 | ENSG00000010322 | NISCH | CHOL,DLBC,GBM,KIRC,KIRP,LUSC,MESO,SKCM,STAD | | GBM | | ENSG00000259291 | ENSG00000023171 | GRAMD1B | CHOL,GBM,KICH,KIRP,TGCT,THCA,UCS | | GBM,KIRP | | ENSG00000259291 | ENSG00000023228 | NDUFS1 | CHOL,HNSC,KIRC,KIRP,THCA | | HNSC,KIRC | | ENSG00000259291 | ENSG00000047346 | FAM214A | CHOL,COAD,LAML,PAAD,READ | | COAD,READ | | ENSG00000259291 | ENSG00000067221 | STOML1 | CHOL,KICH,SARC,TGCT,THCA,UCS | KICH | | | ENSG00000259291 | ENSG00000076864 | RAP1GAP | CHOL,ESCA,GBM,KICH,KIRC,KIRP,TGCT,THCA | | ESCA,GBM,KIRC,KIRP | | ENSG00000259291 | ENSG00000087258 | GNAO1 | CHOL,GBM,PAAD,SARC,STAD | | GBM,STAD | | ENSG00000259291 | ENSG00000088682 | COQ9 | CHOL,HNSC,KICH,KIRC,SARC,TGCT,THCA | KICH | | | ENSG00000259291 | ENSG00000095321 | CRAT | CHOL,HNSC,KICH,KIRC,SARC,TGCT,THCA | KICH | HNSC | | ENSG00000259291 | ENSG00000100412 | ACO2 | CHOL,ESCA,HNSC,KIRC,KIRP,LIHC,SARC | | ESCA,HNSC | | ENSG00000259291 | ENSG00000107262 | BAG1 | CHOL,KIRC,KIRP,SARC,TGCT | | KIRC | | ENSG00000259291 | ENSG00000114923 | SLC4A3 | CHOL,KICH,MESO,PAAD,PCPG,PRAD,UVM | PCPG | | | ENSG00000259291 | ENSG00000119669 | IRF2BPL | CHOL,KIRC,PAAD,STAD,UVM | | KIRC | | ENSG00000259291 | ENSG00000122042 | UBL3 | CHOL,ESCA,KIRC,KIRP,UCS | | ESCA | | ENSG00000259291 | ENSG00000124422 | USP22 | CHOL,GBM,MESO,PAAD,SARC,UCS | | GBM | | ENSG00000259291 | ENSG00000125375 | DMAC2L | CHOL,KIRC,KIRP,LAML,SARC,TGCT | | KIRC | | ENSG00000259291 | ENSG00000128872 | TMOD2 | CHOL,GBM,LUSC,PAAD,STAD | | GBM,STAD | | ENSG00000259291 | ENSG00000136003 | ISCU | CHOL,HNSC,KICH,KIRC,PAAD,STAD,THYM,UCS | KICH | | | ENSG00000259291 | ENSG00000138029 | HADHB | CHOL,HNSC,KIRC,KIRP,LGG | | HNSC | | ENSG00000259291 | ENSG00000138593 | SECISBP2L | CHOL,COAD,ESCA,GBM,KIRC,READ,STAD | | ESCA,GBM,READ | | ENSG00000259291 | ENSG00000146701 | MDH2 | CHOL,HNSC,KICH,KIRC,THCA | KICH | | | ENSG00000259291 | ENSG00000151498 | ACAD8 | CHOL,HNSC,KIRC,KIRP,LIHC,SARC,THYM | | HNSC,KIRC | | ENSG00000259291 | ENSG00000152234 | ATP5F1A | CHOL,HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | HNSC,KIRC | | ENSG00000259291 | ENSG00000156381 | ANKRD9 | CHOL,GBM,KICH,KIRC,KIRP,LGG,LIHC,THCA | KICH | KIRC | | ENSG00000259291 | ENSG00000159792 | PSKH1 | CHOL,DLBC,KICH,KIRC,LIHC,MESO,PAAD,TGCT,THCA | KICH | | | ENSG00000259291 | ENSG00000165861 | ZFYVE1 | CHOL,KICH,KIRC,KIRP,PAAD | KICH | | | ENSG00000259291 | ENSG00000166402 | TUB | CHOL,GBM,KIRC,PAAD,SARC,TGCT | | GBM | | ENSG00000259291 | ENSG00000168924 | LETM1 | CHOL,KICH,KIRC,KIRP,LIHC,THCA,THYM | KICH | | | ENSG00000259291 | ENSG00000170634 | ACYP2 | CHOL,HNSC,LGG,PAAD,TGCT,THCA,UCS | | HNSC | | ENSG00000259291 | ENSG00000175662 | TOM1L2 | CHOL,COAD,GBM,KIRC,KIRP,LGG,LIHC,MESO,PAAD,PCPG,READ,SARC,STAD,TGCT,THCA,UCS | | COAD,GBM,KIRC,KIRP,READ,STAD | | ENSG00000259291 | ENSG00000176542 | USF3 | CHOL,GBM,KIRC,READ,TGCT | | GBM | | ENSG00000259291 | ENSG00000177981 | ASB8 | CHOL,HNSC,KIRC,KIRP,PAAD,READ,TGCT,THCA,THYM,UCS | | HNSC | | ENSG00000259291 | ENSG00000186687 | LYRM7 | CHOL,HNSC,KIRP,STAD,THCA | | HNSC,KIRP | | ENSG00000259291 | ENSG00000197620 | EOLA1 | CHOL,KICH,PAAD,TGCT,THCA | KICH | | | ENSG00000259291 | ENSG00000203772 | SPRN | CHOL,DLBC,GBM,PAAD,UVM | | GBM | | ENSG00000259291 | ENSG00000224051 | CPTP | CHOL,KICH,KIRC,KIRP,TGCT | KICH | | | ENSG00000259291 | ENSG00000077782 | FGFR1 | COAD,DLBC,PAAD,STAD,UVM | | COAD,STAD | | ENSG00000259291 | ENSG00000082014 | SMARCD3 | COAD,HNSC,PAAD,STAD,TGCT | | HNSC,STAD | | ENSG00000259291 | ENSG00000105894 | PTN | COAD,PAAD,READ,STAD,TGCT | | COAD,READ,STAD | | ENSG00000259291 | ENSG00000119938 | PPP1R3C | COAD,HNSC,KICH,READ,SARC,STAD,TGCT,UCS | KICH | COAD,HNSC,READ,STAD | | ENSG00000259291 | ENSG00000125354 | SEPTIN6 | COAD,KICH,KIRC,KIRP,LAML,PAAD,READ,THYM | | COAD,KIRP,READ | | ENSG00000259291 | ENSG00000133943 | DGLUCY | COAD,ESCA,GBM,HNSC,KICH,KIRC,LAML,LGG,PRAD,SARC,STAD,TGCT,THCA | KICH | ESCA,HNSC | | ENSG00000259291 | ENSG00000175906 | ARL4D | COAD,KIRC,KIRP,READ,SARC,THCA | | COAD,KIRC,KIRP,READ | | ENSG00000259291 | ENSG00000180875 | GREM2 | COAD,ESCA,PAAD,READ,SARC,STAD,TGCT | | COAD,ESCA,READ,STAD | | ENSG00000259291 | ENSG00000006025 | OSBPL7 | DLBC,GBM,LUSC,PCPG,TGCT,THYM | PCPG | | | ENSG00000259291 | ENSG00000049769 | PPP1R3F | DLBC,GBM,HNSC,LAML,PAAD,TGCT | | GBM | | ENSG00000259291 | ENSG00000055163 | CYFIP2 | DLBC,ESCA,GBM,KIRC,KIRP,LAML,PAAD,SARC,THYM | | ESCA,GBM,KIRC,KIRP | | ENSG00000259291 | ENSG00000117115 | PADI2 | DLBC,HNSC,KIRC,KIRP,READ,UCS | | HNSC,KIRC,KIRP,READ | | ENSG00000259291 | ENSG00000125967 | NECAB3 | DLBC,ESCA,GBM,KICH,SARC,THCA | KICH | ESCA,GBM | | ENSG00000259291 | ENSG00000128609 | NDUFA5 | DLBC,ESCA,HNSC,KIRC,PAAD,READ,THCA | | READ | | ENSG00000259291 | ENSG00000138834 | MAPK8IP3 | DLBC,GBM,TGCT,THYM,UVM | | GBM | | ENSG00000259291 | ENSG00000147912 | FBXO10 | DLBC,GBM,KICH,PAAD,UCS,UVM | KICH | | | ENSG00000259291 | ENSG00000160445 | ZER1 | DLBC,GBM,KIRC,KIRP,PAAD,READ,SARC,STAD,TGCT | | GBM,STAD | | ENSG00000259291 | ENSG00000165802 | NSMF | DLBC,GBM,LGG,TGCT,UVM | | GBM | | ENSG00000259291 | ENSG00000182670 | TTC3 | DLBC,GBM,LAML,MESO,TGCT | | GBM | | ENSG00000259291 | ENSG00000189319 | FAM53B | DLBC,KICH,KIRP,PAAD,THCA,THYM | KICH | | | ENSG00000259291 | ENSG00000250067 | YJEFN3 | DLBC,GBM,LAML,MESO,OV | | GBM | | ENSG00000259291 | ENSG00000060762 | MPC1 | ESCA,HNSC,KIRC,KIRP,LGG,THCA,THYM,UCS | | ESCA,HNSC,KIRC | | ENSG00000259291 | ENSG00000121769 | FABP3 | ESCA,HNSC,KICH,KIRC,PRAD,THCA,UCS,UVM | KICH | ESCA,HNSC | | ENSG00000259291 | ENSG00000122971 | ACADS | ESCA,HNSC,KICH,THCA,THYM | KICH | ESCA,HNSC | | ENSG00000259291 | ENSG00000137992 | DBT | ESCA,KIRC,KIRP,READ,THCA | | ESCA,KIRC,KIRP | | ENSG00000259291 | ENSG00000158006 | PAFAH2 | ESCA,KIRC,KIRP,THCA,THYM | | ESCA,KIRC | | ENSG00000259291 | ENSG00000176894 | PXMP2 | ESCA,KIRP,LAML,TGCT,THYM | | ESCA | | ENSG00000259291 | ENSG00000185133 | INPP5J | ESCA,KICH,KIRC,KIRP,PAAD | KICH | ESCA,KIRC,KIRP | | ENSG00000259291 | ENSG00000196177 | ACADSB | ESCA,HNSC,KIRC,KIRP,TGCT,THCA | | ESCA,HNSC,KIRC,KIRP | | ENSG00000259291 | ENSG00000198721 | ECI2 | ESCA,HNSC,KIRC,KIRP,TGCT,THCA,THYM | | ESCA,KIRC,KIRP | | ENSG00000259291 | ENSG00000018189 | RUFY3 | GBM,PAAD,STAD,THYM,UVM | | GBM | | ENSG00000259291 | ENSG00000067840 | PDZD4 | GBM,PAAD,PCPG,STAD,UVM | PCPG | GBM,STAD | | ENSG00000259291 | ENSG00000068976 | PYGM | GBM,HNSC,STAD,THCA,UCS | | GBM,HNSC,STAD | | ENSG00000259291 | ENSG00000089220 | PEBP1 | GBM,KIRC,KIRP,LGG,PAAD,THCA | | GBM,KIRC | | ENSG00000259291 | ENSG00000089486 | CDIP1 | GBM,KICH,KIRC,LAML,LGG,PAAD,THCA | KICH | GBM | | ENSG00000259291 | ENSG00000100241 | SBF1 | GBM,KICH,LIHC,THYM,UCS | KICH | GBM | | ENSG00000259291 | ENSG00000102226 | USP11 | GBM,LAML,PAAD,PCPG,TGCT | PCPG | GBM | | ENSG00000259291 | ENSG00000113441 | LNPEP | GBM,READ,SARC,STAD,TGCT | | GBM | | ENSG00000259291 | ENSG00000117408 | IPO13 | GBM,KICH,KIRC,KIRP,PAAD,THCA | | GBM,KIRC | | ENSG00000259291 | ENSG00000119242 | CCDC92 | GBM,LUSC,PAAD,STAD,TGCT,UVM | | GBM | | ENSG00000259291 | ENSG00000121753 | ADGRB2 | GBM,LUSC,MESO,TGCT,THCA,THYM | | GBM | | ENSG00000259291 | ENSG00000134042 | MRO | GBM,KIRC,KIRP,LGG,THCA | | KIRC,KIRP | | ENSG00000259291 | ENSG00000136002 | ARHGEF4 | GBM,LGG,PCPG,PRAD,TGCT | PCPG | GBM | | ENSG00000259291 | ENSG00000141540 | TTYH2 | GBM,PAAD,TGCT,THCA,UCS | | GBM | | ENSG00000259291 | ENSG00000154930 | ACSS1 | GBM,KICH,KIRC,KIRP,LAML,LGG,MESO,TGCT,THCA | KICH | GBM | | ENSG00000259291 | ENSG00000162065 | TBC1D24 | GBM,KICH,KIRC,KIRP,MESO,PAAD,TGCT,THCA | KICH | GBM,KIRC,KIRP | | ENSG00000259291 | ENSG00000164976 | MYORG | GBM,KICH,KIRC,KIRP,LGG,SARC | | GBM,KIRC,KIRP | | ENSG00000259291 | ENSG00000171533 | MAP6 | GBM,KIRP,PAAD,TGCT,THCA,UCEC | | GBM,KIRP | | ENSG00000259291 | ENSG00000174669 | SLC29A2 | GBM,HNSC,KICH,KIRC,KIRP | KICH | KIRC,KIRP | | ENSG00000259291 | ENSG00000176749 | CDK5R1 | GBM,MESO,PAAD,PCPG,UCS | PCPG | GBM | | ENSG00000259291 | ENSG00000179364 | PACS2 | GBM,HNSC,KICH,PCPG,TGCT | | GBM | | ENSG00000259291 | ENSG00000182175 | RGMA | GBM,LGG,PAAD,STAD,UCS,UVM | | STAD | | ENSG00000259291 | ENSG00000196440 | ARMCX4 | GBM,PAAD,PRAD,SARC,TGCT | | GBM | | ENSG00000259291 | ENSG00000196535 | MYO18A | GBM,HNSC,LAML,TGCT,THYM,UCS | | GBM,HNSC | | ENSG00000259291 | ENSG00000205363 | INSYN1 | GBM,KICH,KIRP,LGG,UVM | KICH | GBM,KIRP | | ENSG00000259291 | ENSG00000266074 | BAHCC1 | GBM,KIRP,PAAD,TGCT,THYM | | KIRP | | ENSG00000259291 | ENSG00000025039 | RRAGD | HNSC,KICH,KIRC,KIRP,TGCT,UCS | KICH | HNSC | | ENSG00000259291 | ENSG00000072954 | TMEM38A | HNSC,KICH,KIRC,THCA,UCS | KICH | HNSC | | ENSG00000259291 | ENSG00000107537 | PHYH | HNSC,KICH,KIRC,TGCT,THCA | KICH | HNSC,KIRC | | ENSG00000259291 | ENSG00000133315 | MACROD1 | HNSC,KICH,KIRC,TGCT,UCS | | HNSC | | ENSG00000259291 | ENSG00000143434 | SEMA6C | HNSC,MESO,SKCM,TGCT,UCS | | HNSC | | ENSG00000259291 | ENSG00000161558 | TMEM143 | HNSC,KICH,KIRC,PAAD,SARC,THCA,THYM,UCS | | HNSC | | ENSG00000259291 | ENSG00000171033 | PKIA | HNSC,KICH,KIRC,STAD,TGCT,UCS | KICH | HNSC,STAD | | ENSG00000259291 | ENSG00000171503 | ETFDH | HNSC,KIRC,KIRP,READ,SARC,TGCT,THCA | | HNSC,READ | | ENSG00000259291 | ENSG00000176490 | DIRAS1 | HNSC,KICH,KIRP,MESO,PAAD,SARC,TGCT | KICH | HNSC | | ENSG00000259291 | ENSG00000178537 | SLC25A20 | HNSC,KIRC,KIRP,TGCT,THCA,THYM | | HNSC | | ENSG00000259291 | ENSG00000103227 | LMF1 | KICH,LAML,TGCT,THCA,UVM | KICH | | | ENSG00000259291 | ENSG00000132423 | COQ3 | KICH,KIRC,KIRP,THCA,THYM | KICH | | | ENSG00000259291 | ENSG00000136425 | CIB2 | KICH,MESO,SARC,TGCT,THYM | KICH | | | ENSG00000259291 | ENSG00000140519 | RHCG | KICH,KIRC,KIRP,LIHC,SARC | KICH | KIRC,KIRP | | ENSG00000259291 | ENSG00000144827 | ABHD10 | KICH,KIRC,LAML,PAAD,THCA,THYM | KICH | | | ENSG00000259291 | ENSG00000149599 | DUSP15 | KICH,KIRC,KIRP,TGCT,THCA | KICH | KIRC | | ENSG00000259291 | ENSG00000161267 | BDH1 | KICH,KIRC,KIRP,TGCT,THCA,THYM | KICH | KIRC | | ENSG00000259291 | ENSG00000173511 | VEGFB | KICH,PAAD,STAD,TGCT,THCA | KICH | | | ENSG00000259291 | ENSG00000182108 | DEXI | KICH,LGG,PAAD,STAD,TGCT | KICH | | | ENSG00000259291 | ENSG00000182512 | GLRX5 | KICH,KIRC,KIRP,THCA,THYM | | KIRC | | ENSG00000259291 | ENSG00000060971 | ACAA1 | KIRC,KIRP,LGG,TGCT,THYM | | KIRC,KIRP | | ENSG00000259291 | ENSG00000101004 | NINL | KIRC,KIRP,LUSC,PAAD,SKCM,TGCT,THYM | | KIRC | | ENSG00000259291 | ENSG00000105607 | GCDH | KIRC,KIRP,PAAD,THCA,THYM | | KIRC | | ENSG00000259291 | ENSG00000136720 | HS6ST1 | KIRC,KIRP,SKCM,TGCT,UVM | | KIRC,KIRP | | ENSG00000259291 | ENSG00000163947 | ARHGEF3 | KIRC,KIRP,TGCT,THCA,THYM | | KIRC | | ENSG00000259291 | ENSG00000165698 | SPACA9 | KIRC,LAML,PAAD,STAD,THCA,THYM | | KIRC,STAD | | ENSG00000259291 | ENSG00000170153 | RNF150 | KIRC,KIRP,PAAD,STAD,TGCT,THCA | | KIRC,KIRP,STAD | | ENSG00000259291 | ENSG00000186106 | ANKRD46 | KIRC,KIRP,LAML,THCA,UCS | | KIRP | | ENSG00000259291 | ENSG00000105737 | GRIK5 | KIRP,MESO,PAAD,PRAD,SARC,STAD,TGCT | | KIRP,STAD | | ENSG00000259291 | ENSG00000196459 | TRAPPC2 | LAML,MESO,PCPG,SARC,TGCT,THYM | PCPG | | | ENSG00000259291 | ENSG00000131094 | C1QL1 | PAAD,SARC,STAD,TGCT,UVM | | STAD | | ENSG00000259291 | ENSG00000134917 | ADAMTS8 | PAAD,PRAD,STAD,TGCT,UCS | | STAD | | ENSG00000259291 | ENSG00000138944 | SHISAL1 | PAAD,PCPG,PRAD,STAD,UCS | PCPG | STAD | | ENSG00000259291 | ENSG00000151067 | CACNA1C | PAAD,PRAD,READ,SARC,STAD,TGCT | | READ,STAD |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types | | ENSG00000259291 | ENSG00000180787 | ZFP3 | ENSG00000180011 | ZADH2 | 6 | CHOL,HNSC,KIRC,PAAD,READ,THCA | | ENSG00000259291 | ENSG00000180787 | ZFP3 | ENSG00000103657 | HERC1 | 5 | CESC,CHOL,HNSC,PAAD,READ |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000259291 | ENST00000620791 | ORMDL2 | ENSG00000123353 | ENST00000550836 | -0.1830 | 900 | GBM | - | | ENSG00000259291 | ENST00000620791 | ORMDL2 | ENSG00000123353 | ENST00000548974 | -0.1414 | 735 | GBM | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000530047 | -0.1230 | 1257 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000527078 | -1.7750 | 1818 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000402965 | -0.1230 | 1257 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000532887 | -0.1414 | 1286 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000329994 | -0.1230 | 1255 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000526779 | -0.1245 | 498 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | SLC52A2 | ENSG00000185803 | ENST00000526752 | -0.1212 | 1256 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | NME1 | ENSG00000239672 | ENST00000480143 | -0.1041 | 1112 | GBM,BLCA,KIRP,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | NME1 | ENSG00000239672 | ENST00000013034 | -0.1845 | 1775 | GBM,BLCA,KIRP,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000372486 | -0.1167 | 63 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000372483 | -0.1167 | 63 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000481189 | -0.1175 | 458 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000459968 | -0.1457 | 46 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000482347 | -0.1167 | 63 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000372481 | -0.1275 | 725 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | NTMT1 | ENSG00000148335 | ENST00000372480 | -0.7157 | 1338 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000520531 | -0.1243 | 1545 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000521003 | -0.1342 | 514 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000522528 | -0.1061 | 1405 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000522971 | -0.1074 | 381 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000519611 | -0.1141 | 241 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000523847 | -0.1365 | 429 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | LY6E | ENSG00000160932 | ENST00000522024 | -0.1928 | 1025 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000620791 | PRDX4 | ENSG00000123131 | ENST00000379341 | -0.1250 | 1689 | GBM,KIRC,COAD | - | | ENSG00000259291 | ENST00000620791 | PRDX4 | ENSG00000123131 | ENST00000439422 | -0.2236 | 1 | GBM,KIRC,COAD | - | | ENSG00000259291 | ENST00000620791 | P4HB | ENSG00000185624 | ENST00000576390 | -0.2342 | 627 | GBM,KIRC,KIRP | PCPG | | ENSG00000259291 | ENST00000620791 | P4HB | ENSG00000185624 | ENST00000439918 | -0.7600 | 1129 | GBM,KIRC,KIRP | PCPG | | ENSG00000259291 | ENST00000620791 | PTPN12 | ENSG00000127947 | ENST00000523952 | -0.1005 | 1070 | GBM,ESCA | - | | ENSG00000259291 | ENST00000620791 | PTPN12 | ENSG00000127947 | ENST00000447995 | -0.1607 | 1447 | GBM,ESCA | - | | ENSG00000259291 | ENST00000558334 | MOGS | ENSG00000115275 | ENST00000233616 | -0.1385 | 173 | BLCA | - | | ENSG00000259291 | ENST00000558334 | MOGS | ENSG00000115275 | ENST00000452063 | -0.1218 | 174 | BLCA | - | | ENSG00000259291 | ENST00000558334 | ORMDL2 | ENSG00000123353 | ENST00000550836 | -0.2692 | 382 | GBM | - | | ENSG00000259291 | ENST00000558334 | ORMDL2 | ENSG00000123353 | ENST00000548974 | -0.1752 | 382 | GBM | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000530047 | -0.1196 | 168 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000527078 | -1.7750 | 856 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000402965 | -0.1196 | 168 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000532887 | -0.1780 | 663 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000329994 | -0.1222 | 510 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000526779 | -0.1153 | 288 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | SLC52A2 | ENSG00000185803 | ENST00000526752 | -0.1184 | 503 | GBM,BLCA,HNSC,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | NME1 | ENSG00000239672 | ENST00000336097 | -0.1242 | 419 | GBM,BLCA,KIRP,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | NME1 | ENSG00000239672 | ENST00000013034 | -0.3384 | 336 | GBM,BLCA,KIRP,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | NTMT1 | ENSG00000148335 | ENST00000481189 | -0.1481 | 353 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000558334 | NTMT1 | ENSG00000148335 | ENST00000372481 | -0.1344 | 289 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000558334 | NTMT1 | ENSG00000148335 | ENST00000372480 | -0.3955 | 706 | COAD,READ | PCPG | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000520531 | -0.1507 | 599 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000521003 | -0.2675 | 304 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000522528 | -0.1169 | 371 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000522971 | -0.1119 | 192 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000519611 | -0.1341 | 339 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000523847 | -0.1392 | 644 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | LY6E | ENSG00000160932 | ENST00000522024 | -0.2540 | 314 | KIRC,BLCA,HNSC,ESCA,COAD,STAD,READ | KICH | | ENSG00000259291 | ENST00000558334 | CLIC1 | ENSG00000213719 | ENST00000375784 | -0.1238 | 275 | GBM | - | | ENSG00000259291 | ENST00000558334 | CLIC1 | ENSG00000213719 | ENST00000375779 | -0.1238 | 275 | GBM | - | | ENSG00000259291 | ENST00000558334 | CLIC1 | ENSG00000213719 | ENST00000375780 | -0.1238 | 275 | GBM | - | | ENSG00000259291 | ENST00000558334 | CLIC1 | ENSG00000213719 | ENST00000395892 | -0.1238 | 275 | GBM | - | | ENSG00000259291 | ENST00000558334 | CLIC1 | ENSG00000213719 | ENST00000616760 | -0.1408 | 277 | GBM | - | | ENSG00000259291 | ENST00000558334 | PRDX4 | ENSG00000123131 | ENST00000379341 | -0.1682 | 312 | GBM,KIRC,COAD | - | | ENSG00000259291 | ENST00000558334 | PRDX4 | ENSG00000123131 | ENST00000439422 | -0.1349 | 596 | GBM,KIRC,COAD | - | | ENSG00000259291 | ENST00000558334 | PRELID1 | ENSG00000169230 | ENST00000503216 | -0.1004 | 505 | GBM | - | | ENSG00000259291 | ENST00000558334 | P4HB | ENSG00000185624 | ENST00000576390 | -0.2086 | 306 | GBM,KIRC,KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | P4HB | ENSG00000185624 | ENST00000439918 | -0.6483 | 84 | GBM,KIRC,KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | PTPN12 | ENSG00000127947 | ENST00000447995 | -0.1142 | 689 | GBM,ESCA | - | | ENSG00000259291 | ENST00000620791 | ALG3 | ENSG00000214160 | ENST00000397676 | -0.1605 | 519 | GBM,BLCA,HNSC,ESCA,READ | PCPG | | ENSG00000259291 | ENST00000620791 | ALG3 | ENSG00000214160 | ENST00000445626 | -0.1790 | 542 | GBM,BLCA,HNSC,ESCA,READ | PCPG | | ENSG00000259291 | ENST00000620791 | ALG3 | ENSG00000214160 | ENST00000455059 | -0.3671 | 654 | GBM,BLCA,HNSC,ESCA,READ | PCPG | | ENSG00000259291 | ENST00000620791 | PLAUR | ENSG00000011422 | ENST00000339082 | -0.1657 | 1263 | GBM,KIRC,HNSC,KIRP,ESCA | - | | ENSG00000259291 | ENST00000620791 | PLAUR | ENSG00000011422 | ENST00000595038 | -0.3098 | 1213 | GBM,KIRC,HNSC,KIRP,ESCA | - | | ENSG00000259291 | ENST00000620791 | GRN | ENSG00000030582 | ENST00000053867 | -0.1085 | 19 | GBM,KIRP | - | | ENSG00000259291 | ENST00000620791 | GRN | ENSG00000030582 | ENST00000589265 | -0.1085 | 39 | GBM,KIRP | - | | ENSG00000259291 | ENST00000620791 | GRN | ENSG00000030582 | ENST00000586242 | -0.1085 | 19 | GBM,KIRP | - | | ENSG00000259291 | ENST00000620791 | HEXB | ENSG00000049860 | ENST00000511181 | -0.1308 | 143 | GBM | - | | ENSG00000259291 | ENST00000620791 | HEXB | ENSG00000049860 | ENST00000261416 | -0.1308 | 143 | GBM | - | | ENSG00000259291 | ENST00000620791 | HEXB | ENSG00000049860 | ENST00000513336 | -0.1308 | 143 | GBM | - | | ENSG00000259291 | ENST00000620791 | HEXB | ENSG00000049860 | ENST00000509579 | -0.1308 | 143 | GBM | - | | ENSG00000259291 | ENST00000620791 | MYDGF | ENSG00000074842 | ENST00000599761 | -0.1121 | 1110 | GBM,KIRC | - | | ENSG00000259291 | ENST00000620791 | MYDGF | ENSG00000074842 | ENST00000599630 | -0.1382 | 182 | GBM,KIRC | - | | ENSG00000259291 | ENST00000620791 | PYGL | ENSG00000100504 | ENST00000532462 | -0.1046 | 596 | GBM,KIRC,KIRP | KICH,PCPG | | ENSG00000259291 | ENST00000558334 | ANXA2 | ENSG00000182718 | ENST00000558503 | -0.1120 | 307 | GBM,KIRP | - | | ENSG00000259291 | ENST00000558334 | ALG3 | ENSG00000214160 | ENST00000397676 | -0.1204 | 326 | GBM,BLCA,HNSC,ESCA,READ | PCPG | | ENSG00000259291 | ENST00000558334 | ALG3 | ENSG00000214160 | ENST00000445626 | -0.1604 | 293 | GBM,BLCA,HNSC,ESCA,READ | PCPG | | ENSG00000259291 | ENST00000558334 | ALG3 | ENSG00000214160 | ENST00000455059 | -0.2797 | 362 | GBM,BLCA,HNSC,ESCA,READ | PCPG | | ENSG00000259291 | ENST00000558334 | PLAUR | ENSG00000011422 | ENST00000595038 | -0.1460 | 145 | GBM,KIRC,HNSC,KIRP,ESCA | - | | ENSG00000259291 | ENST00000558334 | GRN | ENSG00000030582 | ENST00000053867 | -0.1088 | 219 | GBM,KIRP | - | | ENSG00000259291 | ENST00000558334 | GRN | ENSG00000030582 | ENST00000589265 | -0.1196 | 239 | GBM,KIRP | - | | ENSG00000259291 | ENST00000558334 | GRN | ENSG00000030582 | ENST00000586782 | -0.1317 | 429 | GBM,KIRP | - | | ENSG00000259291 | ENST00000558334 | GRN | ENSG00000030582 | ENST00000586443 | -0.1124 | 411 | GBM,KIRP | - | | ENSG00000259291 | ENST00000558334 | GRN | ENSG00000030582 | ENST00000586242 | -0.1015 | 219 | GBM,KIRP | - | | ENSG00000259291 | ENST00000558334 | TMSB10 | ENSG00000034510 | ENST00000233143 | -0.1189 | 303 | KIRC,HNSC,KIRP,ESCA | - | | ENSG00000259291 | ENST00000558334 | MYDGF | ENSG00000074842 | ENST00000599761 | -0.1653 | 465 | GBM,KIRC | - | | ENSG00000259291 | ENST00000558334 | MYDGF | ENSG00000074842 | ENST00000262947 | -0.1002 | 220 | GBM,KIRC | - | | ENSG00000259291 | ENST00000558334 | MYDGF | ENSG00000074842 | ENST00000599630 | -0.2365 | 497 | GBM,KIRC | - | | ENSG00000259291 | ENST00000558334 | PYGL | ENSG00000100504 | ENST00000532462 | -0.1048 | 604 | GBM,KIRC,KIRP | KICH,PCPG | | ENSG00000259291 | ENST00000558334 | PYGL | ENSG00000100504 | ENST00000544180 | -0.1057 | 243 | GBM,KIRC,KIRP | KICH,PCPG | | ENSG00000259291 | ENST00000620791 | SERPINH1 | ENSG00000149257 | ENST00000530284 | -0.1240 | 522 | GBM,KIRC,HNSC,KIRP,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000620791 | S100A11 | ENSG00000163191 | ENST00000271638 | -0.1045 | 550 | GBM,KIRP,COAD,READ | - | | ENSG00000259291 | ENST00000620791 | LMAN2 | ENSG00000169223 | ENST00000515209 | -0.1484 | 599 | GBM | - | | ENSG00000259291 | ENST00000620791 | SMAGP | ENSG00000170545 | ENST00000605627 | -0.1130 | 606 | GBM | - | | ENSG00000259291 | ENST00000620791 | SMAGP | ENSG00000170545 | ENST00000603838 | -0.1023 | 1239 | GBM | - | | ENSG00000259291 | ENST00000620791 | SMAGP | ENSG00000170545 | ENST00000604188 | -0.1357 | 1372 | GBM | - | | ENSG00000259291 | ENST00000620791 | CASP4 | ENSG00000196954 | ENST00000444739 | -0.1617 | 1507 | GBM,KIRC,KIRP | - | | ENSG00000259291 | ENST00000620791 | CASP4 | ENSG00000196954 | ENST00000393150 | -0.1617 | 1509 | GBM,KIRC,KIRP | - | | ENSG00000259291 | ENST00000620791 | CASP4 | ENSG00000196954 | ENST00000613196 | -0.1617 | 1509 | GBM,KIRC,KIRP | - | | ENSG00000259291 | ENST00000620791 | EIF6 | ENSG00000242372 | ENST00000621148 | -0.1009 | 463 | READ | - | | ENSG00000259291 | ENST00000620791 | EIF6 | ENSG00000242372 | ENST00000374436 | -0.1038 | 467 | READ | - | | ENSG00000259291 | ENST00000620791 | EIF6 | ENSG00000242372 | ENST00000374443 | -0.1038 | 467 | READ | - | | ENSG00000259291 | ENST00000620791 | EIF6 | ENSG00000242372 | ENST00000374450 | -0.1038 | 467 | READ | - | | ENSG00000259291 | ENST00000620791 | EIF6 | ENSG00000242372 | ENST00000415116 | -0.1687 | 569 | READ | - | | ENSG00000259291 | ENST00000620791 | SEL1L3 | ENSG00000091490 | ENST00000507618 | -0.8344 | 312 | KIRC,KIRP | KICH | | ENSG00000259291 | ENST00000620791 | PTPRE | ENSG00000132334 | ENST00000471218 | -3.6650 | 468 | KIRC | PCPG | | ENSG00000259291 | ENST00000620791 | PTPRE | ENSG00000132334 | ENST00000467366 | -3.6650 | 467 | KIRC | PCPG | | ENSG00000259291 | ENST00000558334 | SERPINH1 | ENSG00000149257 | ENST00000530284 | -0.1217 | 301 | GBM,KIRC,HNSC,KIRP,ESCA,COAD,STAD,READ | - | | ENSG00000259291 | ENST00000558334 | S100A11 | ENSG00000163191 | ENST00000271638 | -0.1846 | 455 | GBM,KIRP,COAD,READ | - | | ENSG00000259291 | ENST00000558334 | LMAN2 | ENSG00000169223 | ENST00000515209 | -0.1563 | 285 | GBM | - | | ENSG00000259291 | ENST00000558334 | SMAGP | ENSG00000170545 | ENST00000605627 | -0.1156 | 630 | GBM | - | | ENSG00000259291 | ENST00000558334 | SMAGP | ENSG00000170545 | ENST00000604188 | -0.1161 | 720 | GBM | - | | ENSG00000259291 | ENST00000558334 | CASP4 | ENSG00000196954 | ENST00000444739 | -0.1424 | 574 | GBM,KIRC,KIRP | - | | ENSG00000259291 | ENST00000558334 | CASP4 | ENSG00000196954 | ENST00000393150 | -0.2004 | 652 | GBM,KIRC,KIRP | - | | ENSG00000259291 | ENST00000558334 | CASP4 | ENSG00000196954 | ENST00000613196 | -0.2004 | 652 | GBM,KIRC,KIRP | - | | ENSG00000259291 | ENST00000558334 | EIF6 | ENSG00000242372 | ENST00000621148 | -0.1100 | 499 | READ | - | | ENSG00000259291 | ENST00000558334 | EIF6 | ENSG00000242372 | ENST00000374436 | -0.1100 | 503 | READ | - | | ENSG00000259291 | ENST00000558334 | EIF6 | ENSG00000242372 | ENST00000374443 | -0.1100 | 503 | READ | - | | ENSG00000259291 | ENST00000558334 | EIF6 | ENSG00000242372 | ENST00000374450 | -0.1100 | 503 | READ | - | | ENSG00000259291 | ENST00000558334 | EIF6 | ENSG00000242372 | ENST00000415116 | -0.1616 | 434 | READ | - | | ENSG00000259291 | ENST00000558334 | SEL1L3 | ENSG00000091490 | ENST00000502949 | -0.1062 | 117 | KIRC,KIRP | KICH | | ENSG00000259291 | ENST00000558334 | SEL1L3 | ENSG00000091490 | ENST00000507618 | -0.3115 | 287 | KIRC,KIRP | KICH | | ENSG00000259291 | ENST00000558334 | PTPRE | ENSG00000132334 | ENST00000471218 | -0.7983 | 231 | KIRC | PCPG | | ENSG00000259291 | ENST00000558334 | PTPRE | ENSG00000132334 | ENST00000467366 | -0.7983 | 230 | KIRC | PCPG | | ENSG00000259291 | ENST00000558334 | SLC15A4 | ENSG00000139370 | ENST00000376740 | -0.1079 | 358 | KIRC,KIRP | - | | ENSG00000259291 | ENST00000620791 | MVP | ENSG00000013364 | ENST00000395353 | -0.3468 | 1392 | KIRP | PCPG | | ENSG00000259291 | ENST00000620791 | MVP | ENSG00000013364 | ENST00000357402 | -0.3468 | 1392 | KIRP | PCPG | | ENSG00000259291 | ENST00000620791 | MVP | ENSG00000013364 | ENST00000566859 | -0.1024 | 986 | KIRP | PCPG | | ENSG00000259291 | ENST00000620791 | MVP | ENSG00000013364 | ENST00000562463 | -0.1325 | 1182 | KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | RHBDF2 | ENSG00000129667 | ENST00000591885 | -0.1001 | 270 | KIRC,BLCA,KIRP,ESCA,COAD,READ | - | | ENSG00000259291 | ENST00000558334 | RHBDF2 | ENSG00000129667 | ENST00000591860 | -0.1294 | 247 | KIRC,BLCA,KIRP,ESCA,COAD,READ | - | | ENSG00000259291 | ENST00000558334 | MVP | ENSG00000013364 | ENST00000395353 | -0.1520 | 272 | KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | MVP | ENSG00000013364 | ENST00000357402 | -0.1520 | 272 | KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | MVP | ENSG00000013364 | ENST00000566859 | -0.1111 | 378 | KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | MVP | ENSG00000013364 | ENST00000562463 | -0.1226 | 348 | KIRP | PCPG | | ENSG00000259291 | ENST00000558334 | MVP | ENSG00000013364 | ENST00000569887 | -0.1056 | 195 | KIRP | PCPG |
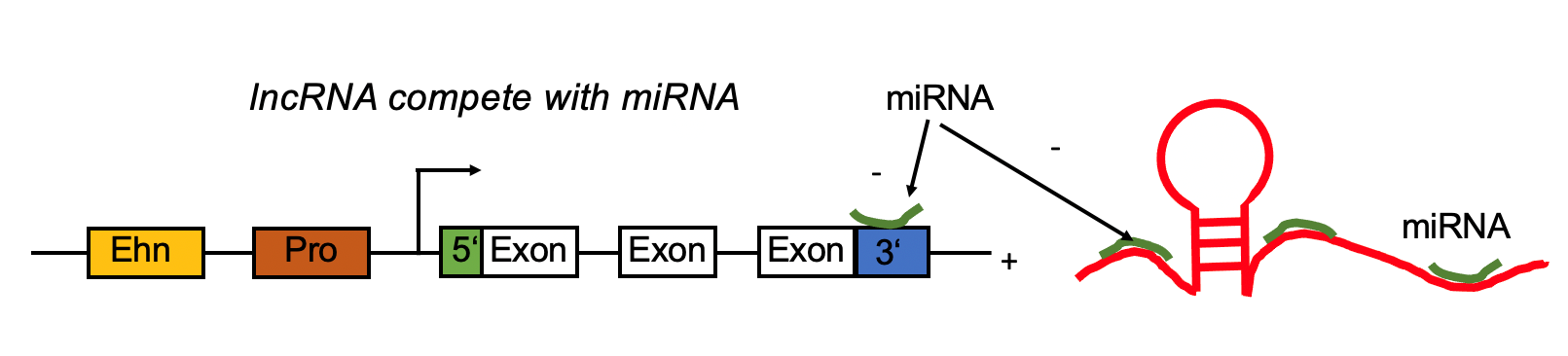
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,GDAP1 | TGCT,THCA,LGG,UCS,PAAD | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ANKRD46 | TGCT,THCA,LGG,UCS,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,CNTFR | HNSC,TGCT,THCA,LGG,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,TNS2 | KICH,HNSC,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,VAMP2 | THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,SLC25A4 | HNSC,THCA,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ARHGEF9 | TGCT,THCA,LGG,READ,PAAD | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,TAPT1 | KICH,THCA,READ,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ACACB | KICH,THCA,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,TSPYL4 | TGCT,LGG,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,UBE2D4 | KICH,HNSC,THCA,UCS,PAAD | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ATP5F1A | KICH,THCA,READ,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,TSPAN7 | KICH,THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,LDHD | HNSC,THCA,LGG,READ,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,APBB1 | HNSC,TGCT,LGG,READ,PAAD,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,MTFR1L | HNSC,TGCT,THCA,READ,UCS,PAAD | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,CNST | TGCT,THCA,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,PDK2 | HNSC,TGCT,LGG,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,NDRG2 | KICH,HNSC,TGCT,THCA,LGG,READ,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ALDH6A1 | HNSC,TGCT,THCA,LGG,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,TOM1L2 | THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,NDN | TGCT,LGG,READ,PAAD,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,CIPC | HNSC,TGCT,THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,FGFBP3 | TGCT,THCA,LGG,UCS,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,MAGEH1 | TGCT,THCA,LGG,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,FBXL17 | HNSC,THCA,LGG,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ATP1B2 | TGCT,THCA,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,WASF3 | TGCT,THCA,LGG,UCS,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,RECK | TGCT,THCA,READ,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,CNNM2 | THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,DMAC2L | KICH,TGCT,THCA,LGG,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,PKIA | HNSC,TGCT,THCA,LGG,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,TEF | KICH,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,FAXDC2 | KICH,HNSC,TGCT,THCA,READ | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,FAM107A | HNSC,TGCT,THCA,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,GPRASP1 | TGCT,THCA,UCS,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ALDH5A1 | TGCT,LGG,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,GPRASP2 | TGCT,THCA,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,METTL7A | HNSC,THCA,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,NAP1L5 | THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,LYRM7 | HNSC,THCA,LGG,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ST3GAL3 | TGCT,READ,UCS,PAAD,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,CRY2 | HNSC,THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,RALGPS1 | KICH,TGCT,THCA,LGG,PAAD | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,MOAP1 | TGCT,THCA,LGG,PAAD,THYM,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,ABAT | TGCT,THCA,LGG,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,PPM1L | HNSC,THCA,LGG,READ,PAAD,KIRP | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,GKAP1 | KICH,THCA,LGG,UCS,PAAD,THYM | | ENSG00000259291 | ZNF710-AS1,hsa-mir-21,KLHL15 | THCA,UCS,PAAD,THYM,KIRP |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000259291.2 | ENST00000620791.1 | 78 | 204 | 440 | | ENSG00000259291.2 | ENST00000558334.1 | 107 | 1979 | 1656 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).