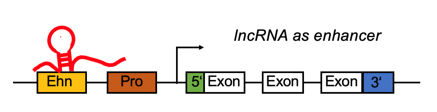
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
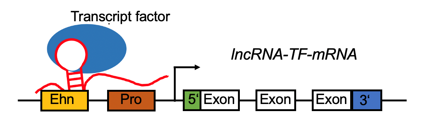
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
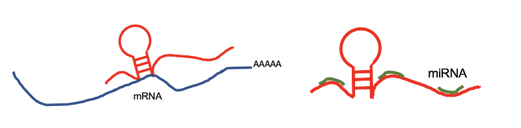
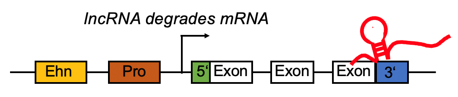
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | TBC1D14,CNOT1,IDH3A,STAU1,VPS26C,TMED5,UBR3,CDC5L,PNPO,EXOC5,AP5M1,LRRFIP1,GDE1,WDR47,MARS2,UHRF1BP1L,CHP1,ATP6V1B2,LNX2,DUSP3,HIPK1,UBQLN1,DNAJC16,TMEM245,ACTR3,TRAPPC10,GNL3L,PSEN1,DAB2,TAOK3,RAB3GAP1,APC,VTI1B,PTPN11,STRN3,EFCAB14,MTREX,BZW1,HECTD1,AP1G1,SPTY2D1,LMBRD1,NAA30,SLC17A5,SLAIN2,MSH3,SLC35D1,MTM1,OS9,NCKAP1,SGTB,PSMC1,SEC24B,CLTC,ATRN,RNF14,RBM18,PCYOX1,TGFBR2,SEC24A,SDHC,MRPL19,SMIM14,ATP10D,CCDC47,NUMB,BCL2L13,MFN2,TOR1AIP1,RBBP9,HSPA8,RAPGEF2,WWP1,SNX29,TRAK2,CTSS,CPEB4,ILK,RNF111,PCNX1,TPP1,SDHD,CTTNBP2NL,CCPG1,ACO2,TMEM33,EIF4EBP2,USP32,SEC24D,UBE2D3,YIPF5,GIGYF2,SELENOP,USP34,ACOX1,CLIP1,MMADHC,USP12,SEL1L,FBXW11,BCKDHA,MAN2A1,FBXL17,SEC23A,GIT2,IQGAP2,LACTB,PINK1,SNAP23,LUZP1,AFF4,CCSER2,ASXL2,G3BP2,MAPK1,FTO,CNIH1,HSD17B4,CLPX,EXOC8,ARHGEF12,ARFGEF2,LPCAT3,MIER3,LARP4,SPPL2A,USP9X,PHAX,ACVR1,ARSB,RETSAT,TANGO6,FNIP2,SCARB2,PDLIM5,WDFY1,MTCH2,CRK,EIF3A,ZNF106,DLAT,FOXN3,CERT1,SRBD1,MAIP1,UGP2,DNAJC3,GALK2,WDR7,SWAP70,NCOA4,CHURC1,ADH5,KIAA0232,C2orf69,SLC31A1,KPNA3,EPS15,PPTC7,DTWD2,CCNYL1,BMPR2,SERINC1,LYSMD3,MSN,SERINC3,ACTR2,FBXW2,SRPRA,UTP14C,ZYG11B,SYNJ1,PPARA,TRIP12,ITGA6,VAMP7,IFNAR1,EPB41L2,GGCX,SNX2,CTSO,MYO18A,GTF2I,IMMT,EIF3L,TGOLN2,ATE1,EIF4G2,CSNK1A1,SPTLC1,NDFIP2,SETX,CS,POLR3B,CAB39,SETD7,CYB5R4,GNG12,PRKAR1A,ACADM,KIF1B,ELF1,BACH1,IL6ST,AFF1,UGGT1,HAT1,VRK2,B3GNT2,ARL2BP,PRCP,USP8,SPTBN1,STAM2,FAM114A2,ATP11C,VAMP3,ITFG1,BTD,ITCH,SLC30A7,MAGT1,CPSF2,TMBIM6,SEC24C,ARCN1,CANX,SPG11,TMED10,AFG3L2,RECQL,FYCO1,LAP3,DLD,USP38,SPOPL,CD302,NDUFS1,SDAD1,AKAP11,PGM2 | | Exist in public source | | NA |
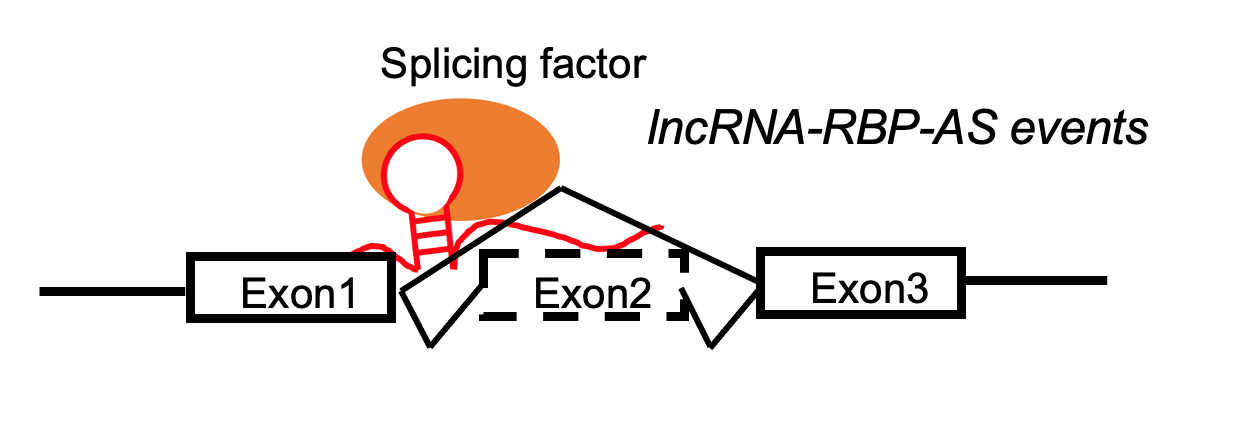
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000228109 | ENST00000414354 | exon_skip_62840 | chr11:70421469-70421580 | -17.26 | -0.1939 | CTTN | ENST00000301843 | In-frame | | ENSG00000228109 | ENST00000414354 | exon_skip_517627 | chrX:154357250-154357274 | -6.35 | -1.0583 | FLNA | ENST00000369850 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000228109 | ENST00000414354 | exon_skip_7523 | chr1:78660553-78660654 | -12.49 | -0.1329 | IFI44 | ENST00000370747 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_62623 | chr11:69079833-69079883 | -9.15 | -0.5719 | TPCN2 | ENST00000294309 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_114376 | chr14:74293153-74293248 | -15.78 | -0.1856 | ABCD4 | ENST00000356924 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_153096 | chr17:44074968-44075087 | -16.68 | -0.1402 | G6PC3 | ENST00000269097 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_370518 | chr22:50247691-50247776 | -14.20 | -0.1732 | HDAC10 | ENST00000216271 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_382458 | chr3:38131595-38131638 | -8.57 | -0.2090 | ACAA1 | ENST00000333167 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_5818 | chr1:45568484-45568684 | -20.64 | -0.1749 | AKR1A1 | ENST00000351829,ENST00000372070 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_9446 | chr1:147270931-147271005 | -12.03 | -0.2228 | CHD1L | ENST00000369258 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_11029 | chr1:154989559-154989707 | -21.55 | -0.1874 | FLAD1 | ENST00000292180 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_41813 | chr10:68337857-68337996 | -14.94 | -0.1311 | HNRNPH3 | ENST00000265866 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_41841 | chr10:68756272-68756483 | -19.28 | -0.1065 | CCAR1 | ENST00000265872 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_66482 | chr11:126274926-126275021 | -15.73 | -0.1656 | FOXRED1 | ENST00000263578 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_74009 | chr11:64773429-64773505 | -12.27 | -0.1636 | SF1 | ENST00000377390 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_87733 | chr12:120571190-120571291 | -13.92 | -0.1513 | RNF10 | ENST00000325954 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_92074 | chr12:50142052-50142110 | -4.78 | -1.5933 | CERS5 | ENST00000317551 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_111863 | chr14:23565815-23565892 | -16.11 | -0.2778 | AP1G2 | ENST00000308724,ENST00000397120 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_111870 | chr14:23566561-23566686 | -15.71 | -0.1354 | AP1G2 | ENST00000397120 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_114365 | chr14:74290026-74290118 | -15.29 | -0.1986 | ABCD4 | ENST00000356924 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_139422 | chr16:89114338-89114487 | -21.80 | -0.1772 | ACSF3 | ENST00000317447,ENST00000406948 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_153767 | chr17:48057031-48057121 | -20.67 | -0.2552 | NFE2L1 | ENST00000362042 | In-frame | | ENSG00000228109 | ENST00000414354 | exon_skip_302168 | chr19:10680345-10680443 | -13.61 | -0.1601 | ILF3 | ENST00000590261 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_309204 | chr19:49181348-49181461 | -18.28 | -0.1708 | TRPM4 | ENST00000252826 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_313557 | chr19:5786748-5786845 | -20.49 | -0.2499 | DUS3L | ENST00000309061 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_319223 | chr19:43526200-43526349 | -24.21 | -0.1891 | ETHE1 | ENST00000292147 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_326885 | chr2:74490024-74490127 | -17.09 | -0.1818 | TTC31 | ENST00000233623 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_339830 | chr2:69423581-69423717 | -14.94 | -0.3248 | NFU1 | ENST00000410022 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_370525 | chr22:50250426-50250523 | -15.89 | -0.2177 | HDAC10 | ENST00000216271 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_374497 | chr3:49999439-49999513 | -8.65 | -0.1632 | RBM6 | ENST00000266022 | Frame-shift | | ENSG00000228109 | ENST00000446695 | exon_skip_380242 | chr3:186784961-186785101 | -13.51 | -0.1485 | EIF4A2 | ENST00000323963 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_442657 | chr5:74730259-74730398 | -12.44 | -0.1398 | GFM2 | ENST00000296805,ENST00000509430 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_453169 | chr6:75702644-75703072 | -16.52 | -0.1558 | SENP6 | ENST00000447266 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_466113 | chr7:48099669-48099787 | -7.44 | -0.1261 | UPP1 | ENST00000331803,ENST00000395564 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_470226 | chr7:103307595-103307708 | -14.65 | -0.1308 | PMPCB | ENST00000249269 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_500083 | chr9:129127963-129128068 | -13.15 | -0.1623 | PTPA | ENST00000337738 | In-frame | | ENSG00000228109 | ENST00000414354 | exon_skip_309977 | chr19:49667018-49667161 | -18.48 | -0.1339 | BCL2L12 | ENST00000246785 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_356468 | chr20:35280653-35280829 | -22.69 | -0.1384 | EIF6 | ENST00000374436,ENST00000374450 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_439146 | chr5:154950820-154950938 | -9.50 | -0.1377 | MRPL22 | ENST00000523037 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_13857 | chr1:168038370-168038488 | -9.82 | -0.1383 | DCAF6 | ENST00000312263 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_95500 | chr12:96003837-96003920 | -9.67 | -0.1343 | LTA4H | ENST00000228740 | Frame-shift | | ENSG00000228109 | ENST00000446695 | exon_skip_123601 | chr15:74897010-74897219 | -15.93 | -0.1659 | MPI | ENST00000352410 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_129597 | chr15:75363998-75364188 | -24.67 | -0.1468 | MAN2C1 | ENST00000267978 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_374552 | chr3:50104247-50104308 | -7.83 | -0.7830 | RBM5 | ENST00000347869 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_462160 | chr6:127316449-127316502 | -7.84 | -0.2306 | ECHDC1 | ENST00000531967 | Frame-shift | | ENSG00000228109 | ENST00000414354 | exon_skip_424741 | chr4:83458345-83458429 | -7.37 | -0.2106 | MRPS18C | ENST00000295491 | In-frame | | ENSG00000228109 | ENST00000414354 | exon_skip_128281 | chr15:64081101-64081151 | -6.12 | -0.3060 | CIAO2A | ENST00000300030 | Frame-shift |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
 |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000228109 | hsa-mir-378a | ENST00000424769 | chr3:196999506-196999573 | 142.00 | -61.05 | 66 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000415244 | chr3:197003017-197003087 | 162.00 | -70.21 | 71 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000415244 | chr3:197003401-197003471 | 157.00 | -59.20 | 69 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000415244 | chr3:197002957-197003023 | 153.00 | -78.62 | 66 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000415244 | chr3:197003168-197003234 | 140.00 | -76.53 | 65 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000446695 | chr3:197003250-197003320 | 162.00 | -70.21 | 71 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000446695 | chr3:197003133-197003205 | 149.00 | -94.29 | 72 | NA | | ENSG00000228109 | hsa-mir-378a | ENST00000414354 | chr3:197003796-197003862 | 140.00 | -76.53 | 65 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000424769 | chr3:196999600-196999680 | 160.00 | -79.32 | 86 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000424769 | chr3:196999461-196999519 | 157.00 | -70.53 | 64 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000424769 | chr3:196999667-196999741 | 150.00 | -71.08 | 76 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003491-197003576 | 187.00 | -85.11 | 84 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003306-197003392 | 182.00 | -80.03 | 88 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003570-197003655 | 174.00 | -74.63 | 85 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003011-197003100 | 155.00 | -82.81 | 80 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003660-197003743 | 155.00 | -76.44 | 87 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003215-197003296 | 152.00 | -83.79 | 82 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197002925-197003015 | 144.00 | -89.78 | 91 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000415244 | chr3:197003390-197003466 | 142.00 | -66.10 | 81 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000446695 | chr3:197003244-197003333 | 155.00 | -82.81 | 80 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000446695 | chr3:197003133-197003209 | 152.00 | -99.69 | 81 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000437064 | chr3:197003725-197003805 | 160.00 | -79.32 | 86 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000437064 | chr3:197003792-197003866 | 150.00 | -71.08 | 76 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000414354 | chr3:197003998-197004078 | 160.00 | -79.32 | 86 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000414354 | chr3:197003843-197003924 | 152.00 | -83.79 | 82 | NA | | ENSG00000228109 | hsa-mir-22 | ENST00000414354 | chr3:197004065-197004139 | 150.00 | -71.08 | 76 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type | | ENSG00000228109 | ENSG00000004534 | RBM6 | exon_skip_370518 | ENSG00000100429:chr22:50247691-50247776 | HDAC10 | ENST00000216271 | Frame-shift | SARC,UVM,READ,PRAD,THCA | | ENSG00000228109 | ENSG00000004534 | RBM6 | exon_skip_382458 | ENSG00000060971:chr3:38131595-38131638 | ACAA1 | ENST00000333167 | Frame-shift | SARC,UVM,READ,PRAD,PCPG,THCA |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000228109 | ENST00000424769 | NCOA4 | ENSG00000266412 | ENST00000579039 | -0.1049 | 59 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000424769 | CNIH1 | ENSG00000100528 | ENST00000556113 | -0.1482 | 5 | - | CHOL | | ENSG00000228109 | ENST00000424769 | SEC23A | ENSG00000100934 | ENST00000548032 | -0.1280 | 1 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000424769 | CERT1 | ENSG00000113163 | ENST00000261415 | -0.5093 | 2 | - | GBM,LUSC | | ENSG00000228109 | ENST00000424769 | TMBIM6 | ENSG00000139644 | ENST00000552635 | -0.1154 | 136 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000424769 | TMBIM6 | ENSG00000139644 | ENST00000552370 | -0.1364 | 65 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000415244 | CNIH1 | ENSG00000100528 | ENST00000556113 | -0.1596 | 16 | - | CHOL | | ENSG00000228109 | ENST00000415244 | SEC23A | ENSG00000100934 | ENST00000548032 | -0.1481 | 1 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000415244 | SEC23A | ENSG00000100934 | ENST00000557280 | -0.4069 | 597 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000415244 | SEC23A | ENSG00000100934 | ENST00000553970 | -0.4069 | 597 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000415244 | CERT1 | ENSG00000113163 | ENST00000261415 | -0.3670 | 460 | - | GBM,LUSC | | ENSG00000228109 | ENST00000415244 | TMBIM6 | ENSG00000139644 | ENST00000552635 | -0.1845 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000415244 | TMBIM6 | ENSG00000139644 | ENST00000552370 | -0.2515 | 307 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | LRRFIP1 | ENSG00000124831 | ENST00000308482 | -0.1690 | 1 | - | BLCA,LUSC | | ENSG00000228109 | ENST00000446695 | LRRFIP1 | ENSG00000124831 | ENST00000289175 | -0.1331 | 1 | - | BLCA,LUSC | | ENSG00000228109 | ENST00000446695 | LRRFIP1 | ENSG00000124831 | ENST00000244815 | -0.1331 | 1 | - | BLCA,LUSC | | ENSG00000228109 | ENST00000446695 | BMPR2 | ENSG00000204217 | ENST00000374580 | -0.1450 | 1 | - | GBM,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | BMPR2 | ENSG00000204217 | ENST00000374574 | -0.1450 | 1 | - | GBM,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | NCOA4 | ENSG00000266412 | ENST00000578454 | -0.1022 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | CNIH1 | ENSG00000100528 | ENST00000216416 | -0.1358 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | CNIH1 | ENSG00000100528 | ENST00000556113 | -0.1283 | 4 | - | CHOL | | ENSG00000228109 | ENST00000446695 | SEC23A | ENSG00000100934 | ENST00000548032 | -0.1286 | 20 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000446695 | SEC23A | ENSG00000100934 | ENST00000557280 | -0.1509 | 34 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000446695 | SEC23A | ENSG00000100934 | ENST00000553970 | -0.1509 | 34 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000446695 | TMEM245 | ENSG00000106771 | ENST00000374586 | -0.1146 | 1 | - | GBM,KIRC,BLCA,LUSC | | ENSG00000228109 | ENST00000446695 | TMEM245 | ENSG00000106771 | ENST00000413712 | -0.1035 | 1 | - | GBM,KIRC,BLCA,LUSC | | ENSG00000228109 | ENST00000446695 | TMEM245 | ENSG00000106771 | ENST00000491854 | -0.1036 | 1 | - | GBM,KIRC,BLCA,LUSC | | ENSG00000228109 | ENST00000446695 | SERINC1 | ENSG00000111897 | ENST00000339697 | -0.1321 | 1 | - | GBM,CHOL,BLCA,LUSC,LUAD,READ | | ENSG00000228109 | ENST00000446695 | CERT1 | ENSG00000113163 | ENST00000380494 | -0.1164 | 1 | - | GBM,LUSC | | ENSG00000228109 | ENST00000446695 | CERT1 | ENSG00000113163 | ENST00000261415 | -0.5112 | 170 | - | GBM,LUSC | | ENSG00000228109 | ENST00000446695 | CERT1 | ENSG00000113163 | ENST00000508809 | -0.1156 | 1 | - | GBM,LUSC | | ENSG00000228109 | ENST00000446695 | HSD17B4 | ENSG00000133835 | ENST00000515320 | -0.1995 | 1 | - | CHOL,PCPG,LUSC | | ENSG00000228109 | ENST00000446695 | HSD17B4 | ENSG00000133835 | ENST00000442060 | -0.1210 | 1 | - | CHOL,PCPG,LUSC | | ENSG00000228109 | ENST00000446695 | HSD17B4 | ENSG00000133835 | ENST00000504811 | -0.1995 | 1 | - | CHOL,PCPG,LUSC | | ENSG00000228109 | ENST00000446695 | HSD17B4 | ENSG00000133835 | ENST00000513628 | -0.1645 | 1 | - | CHOL,PCPG,LUSC | | ENSG00000228109 | ENST00000446695 | HSD17B4 | ENSG00000133835 | ENST00000509514 | -0.1995 | 1 | - | CHOL,PCPG,LUSC | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000552635 | -0.1947 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000552699 | -0.1417 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000267115 | -0.1417 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000549385 | -0.1417 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000423828 | -0.1417 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000552370 | -0.2015 | 10 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000395006 | -0.1417 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMBIM6 | ENSG00000139644 | ENST00000547798 | -0.1417 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | SETD7 | ENSG00000145391 | ENST00000506866 | -0.1036 | 1 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000446695 | SETD7 | ENSG00000145391 | ENST00000274031 | -0.1757 | 1 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000446695 | SETD7 | ENSG00000145391 | ENST00000608958 | -0.1508 | 1 | - | CHOL,BLCA | | ENSG00000228109 | ENST00000446695 | FYCO1 | ENSG00000163820 | ENST00000535325 | -0.1280 | 1 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000437064 | NCOA4 | ENSG00000266412 | ENST00000583565 | -0.1147 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000437064 | NCOA4 | ENSG00000266412 | ENST00000580070 | -0.1147 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000437064 | NCOA4 | ENSG00000266412 | ENST00000578454 | -0.1045 | 12 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000437064 | CNIH1 | ENSG00000100528 | ENST00000557659 | -0.1001 | 1 | - | CHOL | | ENSG00000228109 | ENST00000437064 | CNIH1 | ENSG00000100528 | ENST00000556113 | -0.1133 | 111 | - | CHOL | | ENSG00000228109 | ENST00000437064 | CERT1 | ENSG00000113163 | ENST00000261415 | -0.1904 | 141 | - | GBM,LUSC | | ENSG00000228109 | ENST00000437064 | TMBIM6 | ENSG00000139644 | ENST00000552635 | -0.1154 | 84 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000437064 | TMBIM6 | ENSG00000139644 | ENST00000552370 | -0.1310 | 125 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000414354 | LRRFIP1 | ENSG00000124831 | ENST00000308482 | -0.1238 | 1 | - | BLCA,LUSC | | ENSG00000228109 | ENST00000414354 | NCOA4 | ENSG00000266412 | ENST00000578454 | -0.1214 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000414354 | CNIH1 | ENSG00000100528 | ENST00000556113 | -0.1008 | 1 | - | CHOL | | ENSG00000228109 | ENST00000414354 | CERT1 | ENSG00000113163 | ENST00000261415 | -0.2000 | 105 | - | GBM,LUSC | | ENSG00000228109 | ENST00000414354 | TMBIM6 | ENSG00000139644 | ENST00000552635 | -0.1115 | 90 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000414354 | TMBIM6 | ENSG00000139644 | ENST00000552370 | -0.2515 | 91 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000424769 | ACTR3 | ENSG00000115091 | ENST00000535589 | -0.1036 | 150 | - | PCPG | | ENSG00000228109 | ENST00000424769 | RAPGEF2 | ENSG00000109756 | ENST00000512056 | -0.1235 | 1 | - | GBM,CHOL,LUSC | | ENSG00000228109 | ENST00000415244 | BCKDHA | ENSG00000248098 | ENST00000269980 | -0.1182 | 169 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000415244 | BCKDHA | ENSG00000248098 | ENST00000457836 | -0.1182 | 173 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000415244 | BCKDHA | ENSG00000248098 | ENST00000544905 | -0.1351 | 172 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000415244 | ACTR3 | ENSG00000115091 | ENST00000535589 | -0.1281 | 168 | - | PCPG | | ENSG00000228109 | ENST00000415244 | RAPGEF2 | ENSG00000109756 | ENST00000512056 | -0.1414 | 440 | - | GBM,CHOL,LUSC | | ENSG00000228109 | ENST00000446695 | DAB2 | ENSG00000153071 | ENST00000545653 | -0.1044 | 1 | - | PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | DAB2 | ENSG00000153071 | ENST00000339788 | -0.1044 | 1 | - | PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | DAB2 | ENSG00000153071 | ENST00000320816 | -0.1044 | 1 | - | PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | DAB2 | ENSG00000153071 | ENST00000509337 | -0.1036 | 1 | - | PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | MARS2 | ENSG00000247626 | ENST00000282276 | -0.1300 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | BCKDHA | ENSG00000248098 | ENST00000269980 | -0.1569 | 1 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000446695 | BCKDHA | ENSG00000248098 | ENST00000457836 | -0.1569 | 1 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000446695 | BCKDHA | ENSG00000248098 | ENST00000544905 | -0.1948 | 1 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000446695 | ACTR3 | ENSG00000115091 | ENST00000263238 | -0.1210 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | ACTR3 | ENSG00000115091 | ENST00000535589 | -0.1567 | 22 | - | PCPG | | ENSG00000228109 | ENST00000446695 | RAPGEF2 | ENSG00000109756 | ENST00000264431 | -0.1602 | 1 | - | GBM,CHOL,LUSC | | ENSG00000228109 | ENST00000446695 | POLR3B | ENSG00000013503 | ENST00000228347 | -0.1053 | 1 | - | KIRC,PCPG | | ENSG00000228109 | ENST00000446695 | POLR3B | ENSG00000013503 | ENST00000539066 | -0.1053 | 1 | - | KIRC,PCPG | | ENSG00000228109 | ENST00000437064 | ACTR3 | ENSG00000115091 | ENST00000535589 | -0.1036 | 98 | - | PCPG | | ENSG00000228109 | ENST00000414354 | DAB2 | ENSG00000153071 | ENST00000509337 | -0.1092 | 1 | - | PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000414354 | BCKDHA | ENSG00000248098 | ENST00000269980 | -0.1094 | 1 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000414354 | BCKDHA | ENSG00000248098 | ENST00000457836 | -0.1094 | 1 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000414354 | BCKDHA | ENSG00000248098 | ENST00000544905 | -0.1432 | 1 | - | CHOL,PCPG,STAD | | ENSG00000228109 | ENST00000414354 | ACTR3 | ENSG00000115091 | ENST00000535589 | -0.2302 | 129 | - | PCPG | | ENSG00000228109 | ENST00000424769 | FYCO1 | ENSG00000163820 | ENST00000438446 | -0.1226 | 13 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000424769 | CHP1 | ENSG00000187446 | ENST00000560397 | -0.5840 | 84 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000424769 | CHP1 | ENSG00000187446 | ENST00000560965 | -0.1078 | 107 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000424769 | UHRF1BP1L | ENSG00000111647 | ENST00000548712 | -0.1846 | 1 | - | GBM,CHOL | | ENSG00000228109 | ENST00000424769 | GGCX | ENSG00000115486 | ENST00000430215 | -0.2536 | 1 | - | CHOL | | ENSG00000228109 | ENST00000424769 | GGCX | ENSG00000115486 | ENST00000421496 | -0.1231 | 1 | - | CHOL | | ENSG00000228109 | ENST00000424769 | GGCX | ENSG00000115486 | ENST00000428479 | -0.1107 | 7 | - | CHOL | | ENSG00000228109 | ENST00000424769 | DNAJC16 | ENSG00000116138 | ENST00000472665 | -0.1359 | 1 | - | CHOL | | ENSG00000228109 | ENST00000424769 | ACADM | ENSG00000117054 | ENST00000420607 | -0.1977 | 143 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000424769 | TMED5 | ENSG00000117500 | ENST00000370280 | -0.1300 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000424769 | SEC24D | ENSG00000150961 | ENST00000506622 | -0.1043 | 1 | - | CHOL | | ENSG00000228109 | ENST00000415244 | FYCO1 | ENSG00000163820 | ENST00000438446 | -0.1745 | 191 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000415244 | LYSMD3 | ENSG00000176018 | ENST00000453259 | -0.1153 | 580 | - | CHOL,LUSC | | ENSG00000228109 | ENST00000415244 | CHP1 | ENSG00000187446 | ENST00000560784 | -0.1072 | 224 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000415244 | CHP1 | ENSG00000187446 | ENST00000560397 | -1.0300 | 308 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000415244 | PNPO | ENSG00000108439 | ENST00000584061 | -0.1017 | 1 | - | CHOL | | ENSG00000228109 | ENST00000415244 | PNPO | ENSG00000108439 | ENST00000585320 | -0.1076 | 273 | - | CHOL | | ENSG00000228109 | ENST00000415244 | GGCX | ENSG00000115486 | ENST00000430215 | -0.2126 | 268 | - | CHOL | | ENSG00000228109 | ENST00000415244 | GGCX | ENSG00000115486 | ENST00000428479 | -0.1217 | 299 | - | CHOL | | ENSG00000228109 | ENST00000415244 | PLEKHB2 | ENSG00000115762 | ENST00000404460 | -0.3367 | 1 | - | GBM | | ENSG00000228109 | ENST00000415244 | DNAJC16 | ENSG00000116138 | ENST00000472665 | -0.1121 | 198 | - | CHOL | | ENSG00000228109 | ENST00000415244 | ACADM | ENSG00000117054 | ENST00000420607 | -0.1977 | 730 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000415244 | ACADM | ENSG00000117054 | ENST00000528016 | -0.1018 | 244 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000415244 | IQGAP2 | ENSG00000145703 | ENST00000396234 | -0.1446 | 1 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000415244 | IQGAP2 | ENSG00000145703 | ENST00000502745 | -0.1446 | 1 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000415244 | IQGAP2 | ENSG00000145703 | ENST00000513534 | -0.1129 | 63 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000415244 | SEC24D | ENSG00000150961 | ENST00000506622 | -0.1066 | 603 | - | CHOL | | ENSG00000228109 | ENST00000446695 | FYCO1 | ENSG00000163820 | ENST00000433878 | -0.1280 | 1 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000446695 | FYCO1 | ENSG00000163820 | ENST00000296137 | -0.1280 | 1 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000446695 | FYCO1 | ENSG00000163820 | ENST00000438446 | -0.1020 | 31 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000446695 | GNG12 | ENSG00000172380 | ENST00000370982 | -0.1254 | 1 | - | BLCA,PCPG,COAD | | ENSG00000228109 | ENST00000446695 | LYSMD3 | ENSG00000176018 | ENST00000509384 | -0.1513 | 1 | - | CHOL,LUSC | | ENSG00000228109 | ENST00000446695 | CHP1 | ENSG00000187446 | ENST00000560411 | -0.1347 | 1 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | CHP1 | ENSG00000187446 | ENST00000334660 | -0.1241 | 1 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | CHP1 | ENSG00000187446 | ENST00000560397 | -1.8550 | 187 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | CTSO | ENSG00000256043 | ENST00000433477 | -0.1182 | 1 | - | CHOL,LUSC,READ | | ENSG00000228109 | ENST00000446695 | MSN | ENSG00000147065 | ENST00000360270 | -0.1259 | 1 | - | LUSC | | ENSG00000228109 | ENST00000446695 | RETSAT | ENSG00000042445 | ENST00000429806 | -0.1163 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | RETSAT | ENSG00000042445 | ENST00000295802 | -0.1163 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | RETSAT | ENSG00000042445 | ENST00000449375 | -0.1110 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | RETSAT | ENSG00000042445 | ENST00000438611 | -0.1702 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | ATP11C | ENSG00000101974 | ENST00000370557 | -0.1293 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | ATP11C | ENSG00000101974 | ENST00000433868 | -0.1293 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | ATP11C | ENSG00000101974 | ENST00000327569 | -0.1293 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | ATP11C | ENSG00000101974 | ENST00000361648 | -0.1293 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | ATP11C | ENSG00000101974 | ENST00000450801 | -0.1293 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | TANGO6 | ENSG00000103047 | ENST00000261778 | -0.1596 | 26 | - | CHOL | | ENSG00000228109 | ENST00000446695 | TANGO6 | ENSG00000103047 | ENST00000565852 | -0.1027 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | TANGO6 | ENSG00000103047 | ENST00000561856 | -0.2459 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PNPO | ENSG00000108439 | ENST00000584061 | -0.1233 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PNPO | ENSG00000108439 | ENST00000583245 | -0.1497 | 13 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PNPO | ENSG00000108439 | ENST00000434554 | -0.1901 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PNPO | ENSG00000108439 | ENST00000225573 | -0.1183 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PNPO | ENSG00000108439 | ENST00000585320 | -0.1386 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PNPO | ENSG00000108439 | ENST00000582171 | -0.1360 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | GGCX | ENSG00000115486 | ENST00000233838 | -0.1672 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | GGCX | ENSG00000115486 | ENST00000430215 | -0.1664 | 23 | - | CHOL | | ENSG00000228109 | ENST00000446695 | GGCX | ENSG00000115486 | ENST00000423570 | -0.1398 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | GGCX | ENSG00000115486 | ENST00000421496 | -0.1231 | 16 | - | CHOL | | ENSG00000228109 | ENST00000446695 | GGCX | ENSG00000115486 | ENST00000428479 | -0.1025 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000409158 | -0.1339 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000403716 | -0.1339 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000439822 | -0.1339 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000234115 | -0.1339 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000628582 | -0.1489 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000438882 | -0.1489 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000404460 | -0.2284 | 3 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000409612 | -0.1339 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PLEKHB2 | ENSG00000115762 | ENST00000409279 | -0.1339 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PCYOX1 | ENSG00000116005 | ENST00000433351 | -0.1173 | 1 | - | CHOL,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | PCYOX1 | ENSG00000116005 | ENST00000264441 | -0.1229 | 1 | - | CHOL,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | DNAJC16 | ENSG00000116138 | ENST00000375838 | -0.1445 | 9 | - | CHOL | | ENSG00000228109 | ENST00000446695 | DNAJC16 | ENSG00000116138 | ENST00000375847 | -0.1664 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | DNAJC16 | ENSG00000116138 | ENST00000475133 | -0.1664 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | DNAJC16 | ENSG00000116138 | ENST00000375849 | -0.1159 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | DNAJC16 | ENSG00000116138 | ENST00000616884 | -0.1664 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | DNAJC16 | ENSG00000116138 | ENST00000472665 | -0.1233 | 9 | - | CHOL | | ENSG00000228109 | ENST00000446695 | SLC35D1 | ENSG00000116704 | ENST00000235345 | -0.1466 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | ACADM | ENSG00000117054 | ENST00000526129 | -0.1004 | 1 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | ACADM | ENSG00000117054 | ENST00000526196 | -0.1004 | 1 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | ACADM | ENSG00000117054 | ENST00000532509 | -0.1004 | 1 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | ACADM | ENSG00000117054 | ENST00000534334 | -0.1478 | 1 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | ACADM | ENSG00000117054 | ENST00000420607 | -0.8200 | 15 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | ACADM | ENSG00000117054 | ENST00000528016 | -0.1530 | 1 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | TMED5 | ENSG00000117500 | ENST00000370282 | -0.1033 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | TMED5 | ENSG00000117500 | ENST00000370290 | -0.1076 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | SLC31A1 | ENSG00000136868 | ENST00000374212 | -0.1367 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | IQGAP2 | ENSG00000145703 | ENST00000274364 | -0.1106 | 1 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IQGAP2 | ENSG00000145703 | ENST00000379730 | -0.1106 | 1 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IQGAP2 | ENSG00000145703 | ENST00000396234 | -0.1939 | 1 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IQGAP2 | ENSG00000145703 | ENST00000502745 | -0.1939 | 1 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | SEC24D | ENSG00000150961 | ENST00000514561 | -0.1177 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | PINK1 | ENSG00000158828 | ENST00000321556 | -0.1455 | 1 | - | GBM,CHOL,HNSC,ESCA,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | ACOX1 | ENSG00000161533 | ENST00000293217 | -0.1320 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000446695 | ACOX1 | ENSG00000161533 | ENST00000587927 | -0.1186 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000437064 | CHP1 | ENSG00000187446 | ENST00000560397 | -1.4100 | 58 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000437064 | PNPO | ENSG00000108439 | ENST00000583245 | -0.1092 | 1 | - | CHOL | | ENSG00000228109 | ENST00000437064 | SEC24A | ENSG00000113615 | ENST00000398844 | -0.1134 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000437064 | GGCX | ENSG00000115486 | ENST00000430215 | -1.0886 | 1 | - | CHOL | | ENSG00000228109 | ENST00000437064 | PLEKHB2 | ENSG00000115762 | ENST00000409158 | -0.1610 | 1 | - | GBM | | ENSG00000228109 | ENST00000437064 | PLEKHB2 | ENSG00000115762 | ENST00000409612 | -0.1610 | 1 | - | GBM | | ENSG00000228109 | ENST00000437064 | PLEKHB2 | ENSG00000115762 | ENST00000409279 | -0.1610 | 1 | - | GBM | | ENSG00000228109 | ENST00000437064 | PCYOX1 | ENSG00000116005 | ENST00000264441 | -0.1429 | 1 | - | CHOL,LUSC,LUAD | | ENSG00000228109 | ENST00000437064 | ACADM | ENSG00000117054 | ENST00000420607 | -0.1977 | 91 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000414354 | FYCO1 | ENSG00000163820 | ENST00000438446 | -0.1841 | 1 | - | BLCA,HNSC,ESCA,PCPG,LUSC,STAD,READ | | ENSG00000228109 | ENST00000414354 | CHP1 | ENSG00000187446 | ENST00000560784 | -0.1072 | 8 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000414354 | CHP1 | ENSG00000187446 | ENST00000560397 | -1.0300 | 92 | - | GBM,CHOL,COAD,READ | | ENSG00000228109 | ENST00000414354 | RETSAT | ENSG00000042445 | ENST00000438611 | -0.1222 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000414354 | PNPO | ENSG00000108439 | ENST00000585320 | -0.1208 | 1 | - | CHOL | | ENSG00000228109 | ENST00000414354 | GGCX | ENSG00000115486 | ENST00000430215 | -0.2126 | 52 | - | CHOL | | ENSG00000228109 | ENST00000414354 | GGCX | ENSG00000115486 | ENST00000428479 | -0.1018 | 21 | - | CHOL | | ENSG00000228109 | ENST00000414354 | PLEKHB2 | ENSG00000115762 | ENST00000404460 | -0.1798 | 1 | - | GBM | | ENSG00000228109 | ENST00000414354 | DNAJC16 | ENSG00000116138 | ENST00000472665 | -0.1121 | 1 | - | CHOL | | ENSG00000228109 | ENST00000414354 | ACADM | ENSG00000117054 | ENST00000420607 | -0.1977 | 251 | - | CHOL,KIRC,HNSC,PCPG,COAD,STAD,READ | | ENSG00000228109 | ENST00000414354 | IQGAP2 | ENSG00000145703 | ENST00000513534 | -0.1237 | 44 | - | CHOL,KIRC,HNSC,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000424769 | MAIP1 | ENSG00000162972 | ENST00000435773 | -0.2883 | 49 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000424769 | UGP2 | ENSG00000169764 | ENST00000483108 | -0.1013 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000424769 | BTD | ENSG00000169814 | ENST00000449107 | -0.2698 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000424769 | SDHD | ENSG00000204370 | ENST00000528182 | -0.1325 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000424769 | SDHD | ENSG00000204370 | ENST00000528048 | -0.1325 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000424769 | SDHD | ENSG00000204370 | ENST00000534010 | -0.1245 | 149 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000424769 | SELENOP | ENSG00000250722 | ENST00000507920 | -0.1019 | 1 | - | CHOL,BLCA,LUSC,LUAD,COAD,STAD,READ | | ENSG00000228109 | ENST00000424769 | LPCAT3 | ENSG00000111684 | ENST00000261407 | -0.1246 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000424769 | ATP6V1B2 | ENSG00000147416 | ENST00000521442 | -0.1602 | 1 | - | GBM | | ENSG00000228109 | ENST00000424769 | PDLIM5 | ENSG00000163110 | ENST00000512274 | -0.1242 | 56 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000415244 | ACOX1 | ENSG00000161533 | ENST00000301608 | -0.1009 | 427 | - | CHOL,COAD | | ENSG00000228109 | ENST00000415244 | ACOX1 | ENSG00000161533 | ENST00000589301 | -0.1245 | 100 | - | CHOL,COAD | | ENSG00000228109 | ENST00000415244 | MAIP1 | ENSG00000162972 | ENST00000435773 | -0.1805 | 172 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000415244 | UGP2 | ENSG00000169764 | ENST00000497510 | -0.1009 | 184 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000415244 | UGP2 | ENSG00000169764 | ENST00000467400 | -0.1061 | 184 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000415244 | UGP2 | ENSG00000169764 | ENST00000487640 | -0.1075 | 115 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000415244 | BTD | ENSG00000169814 | ENST00000417015 | -0.1535 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000415244 | BTD | ENSG00000169814 | ENST00000449107 | -0.1937 | 635 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000415244 | BTD | ENSG00000169814 | ENST00000437172 | -0.1446 | 95 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000415244 | SELENOP | ENSG00000250722 | ENST00000507920 | -0.1072 | 344 | - | CHOL,BLCA,LUSC,LUAD,COAD,STAD,READ | | ENSG00000228109 | ENST00000415244 | MIER3 | ENSG00000155545 | ENST00000451637 | -0.1235 | 301 | - | COAD,READ | | ENSG00000228109 | ENST00000415244 | ATP6V1B2 | ENSG00000147416 | ENST00000519667 | -0.1127 | 450 | - | GBM | | ENSG00000228109 | ENST00000415244 | PDLIM5 | ENSG00000163110 | ENST00000512274 | -0.1056 | 501 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | ACOX1 | ENSG00000161533 | ENST00000572047 | -0.1466 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000446695 | ACOX1 | ENSG00000161533 | ENST00000573078 | -0.1652 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000446695 | ACOX1 | ENSG00000161533 | ENST00000301608 | -0.1411 | 17 | - | CHOL,COAD | | ENSG00000228109 | ENST00000446695 | ACOX1 | ENSG00000161533 | ENST00000576743 | -0.1274 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000446695 | MAIP1 | ENSG00000162972 | ENST00000435773 | -0.1979 | 59 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | SMIM14 | ENSG00000163683 | ENST00000295958 | -0.1313 | 1 | - | CHOL,ESCA,PCPG,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | CLPX | ENSG00000166855 | ENST00000300107 | -0.1341 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000466642 | -0.1159 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000497510 | -0.1541 | 95 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000483108 | -0.1032 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000467400 | -0.1541 | 89 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000496334 | -0.1159 | 20 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000467999 | -0.1159 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | UGP2 | ENSG00000169764 | ENST00000487042 | -0.1159 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | BTD | ENSG00000169814 | ENST00000417015 | -0.1688 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000446695 | BTD | ENSG00000169814 | ENST00000449107 | -0.2155 | 49 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000446695 | BTD | ENSG00000169814 | ENST00000303498 | -0.1047 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000446695 | BTD | ENSG00000169814 | ENST00000437172 | -0.1047 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000446695 | BTD | ENSG00000169814 | ENST00000383778 | -0.1047 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000446695 | USP38 | ENSG00000170185 | ENST00000511739 | -0.1330 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | MTM1 | ENSG00000171100 | ENST00000370396 | -0.1089 | 1 | - | CHOL,PCPG,COAD,READ | | ENSG00000228109 | ENST00000446695 | SRPRA | ENSG00000182934 | ENST00000332118 | -0.1201 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | SRPRA | ENSG00000182934 | ENST00000532259 | -0.1201 | 1 | - | CHOL | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000375549 | -0.1050 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000528182 | -0.1176 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000528048 | -0.1176 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000528021 | -0.1156 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000526592 | -0.1149 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000525291 | -0.1050 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000530923 | -0.1149 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | SDHD | ENSG00000204370 | ENST00000534010 | -0.1203 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | CD302 | ENSG00000241399 | ENST00000429078 | -0.1075 | 1 | - | CHOL,ESCA,LUSC,LUAD,COAD,READ | | ENSG00000228109 | ENST00000446695 | CD302 | ENSG00000241399 | ENST00000553424 | -0.1075 | 1 | - | CHOL,ESCA,LUSC,LUAD,COAD,READ | | ENSG00000228109 | ENST00000446695 | CD302 | ENSG00000241399 | ENST00000259053 | -0.1075 | 1 | - | CHOL,ESCA,LUSC,LUAD,COAD,READ | | ENSG00000228109 | ENST00000446695 | SELENOP | ENSG00000250722 | ENST00000507920 | -0.1193 | 34 | - | CHOL,BLCA,LUSC,LUAD,COAD,STAD,READ | | ENSG00000228109 | ENST00000446695 | LPCAT3 | ENSG00000111684 | ENST00000535479 | -0.1522 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000446695 | LPCAT3 | ENSG00000111684 | ENST00000261407 | -0.1136 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000446695 | LPCAT3 | ENSG00000111684 | ENST00000540090 | -0.1192 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000446695 | LPCAT3 | ENSG00000111684 | ENST00000543794 | -0.1211 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000446695 | LPCAT3 | ENSG00000111684 | ENST00000536797 | -0.1252 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000446695 | LPCAT3 | ENSG00000111684 | ENST00000538910 | -0.1423 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000446695 | TRAK2 | ENSG00000115993 | ENST00000332624 | -0.1154 | 1 | - | BLCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | TRAK2 | ENSG00000115993 | ENST00000620184 | -0.1007 | 1 | - | BLCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | TRAK2 | ENSG00000115993 | ENST00000430254 | -0.1708 | 1 | - | BLCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | SPPL2A | ENSG00000138600 | ENST00000261854 | -0.1665 | 1 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000446695 | MIER3 | ENSG00000155545 | ENST00000381226 | -0.1901 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | MIER3 | ENSG00000155545 | ENST00000381213 | -0.1901 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | MIER3 | ENSG00000155545 | ENST00000381199 | -0.1901 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | MIER3 | ENSG00000155545 | ENST00000452157 | -0.1423 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | MIER3 | ENSG00000155545 | ENST00000409421 | -0.1901 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | MIER3 | ENSG00000155545 | ENST00000451637 | -0.1885 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | CTSS | ENSG00000163131 | ENST00000368985 | -0.1457 | 1 | - | LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | CTSS | ENSG00000163131 | ENST00000448301 | -0.1124 | 1 | - | LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | CTSS | ENSG00000163131 | ENST00000483930 | -0.1340 | 1 | - | LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | CCNYL1 | ENSG00000163249 | ENST00000295414 | -0.1330 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | CCNYL1 | ENSG00000163249 | ENST00000339882 | -0.1330 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | CCNYL1 | ENSG00000163249 | ENST00000420321 | -0.1267 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000446695 | FNIP2 | ENSG00000052795 | ENST00000264433 | -0.1031 | 1 | - | GBM,CHOL,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | ACO2 | ENSG00000100412 | ENST00000216254 | -0.1333 | 1 | - | GBM,HNSC,ESCA,COAD,STAD | | ENSG00000228109 | ENST00000446695 | ACO2 | ENSG00000100412 | ENST00000396512 | -0.1333 | 1 | - | GBM,HNSC,ESCA,COAD,STAD | | ENSG00000228109 | ENST00000446695 | ATP6V1B2 | ENSG00000147416 | ENST00000276390 | -0.1148 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | ATP6V1B2 | ENSG00000147416 | ENST00000519667 | -0.1232 | 43 | - | GBM | | ENSG00000228109 | ENST00000446695 | ATP6V1B2 | ENSG00000147416 | ENST00000521442 | -0.1391 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000437932 | -0.1405 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000615540 | -0.1405 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000359265 | -0.1013 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000317968 | -0.1405 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000512274 | -0.1001 | 95 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000503974 | -0.1035 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000437064 | ACOX1 | ENSG00000161533 | ENST00000301608 | -0.1165 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000437064 | ACOX1 | ENSG00000161533 | ENST00000589301 | -0.1067 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000437064 | MAIP1 | ENSG00000162972 | ENST00000295079 | -0.1065 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000437064 | MAIP1 | ENSG00000162972 | ENST00000392290 | -0.1065 | 1 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000437064 | MAIP1 | ENSG00000162972 | ENST00000435773 | -0.1540 | 175 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000437064 | UGP2 | ENSG00000169764 | ENST00000496334 | -0.1172 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000437064 | BTD | ENSG00000169814 | ENST00000449107 | -0.2013 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000437064 | SDHD | ENSG00000204370 | ENST00000534010 | -0.1245 | 97 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000437064 | SELENOP | ENSG00000250722 | ENST00000507920 | -0.1035 | 135 | - | CHOL,BLCA,LUSC,LUAD,COAD,STAD,READ | | ENSG00000228109 | ENST00000437064 | TRAK2 | ENSG00000115993 | ENST00000332624 | -0.1431 | 1 | - | BLCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000437064 | CTSS | ENSG00000163131 | ENST00000483930 | -0.1012 | 1 | - | LUSC,COAD,READ | | ENSG00000228109 | ENST00000437064 | CCNYL1 | ENSG00000163249 | ENST00000420822 | -0.1054 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000437064 | ATP6V1B2 | ENSG00000147416 | ENST00000521442 | -0.1265 | 1 | - | GBM | | ENSG00000228109 | ENST00000437064 | PDLIM5 | ENSG00000163110 | ENST00000512274 | -0.1444 | 123 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000414354 | ACOX1 | ENSG00000161533 | ENST00000589301 | -0.1698 | 1 | - | CHOL,COAD | | ENSG00000228109 | ENST00000414354 | MAIP1 | ENSG00000162972 | ENST00000435773 | -0.3243 | 175 | - | CHOL,PCPG | | ENSG00000228109 | ENST00000414354 | UGP2 | ENSG00000169764 | ENST00000497510 | -0.1243 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000414354 | UGP2 | ENSG00000169764 | ENST00000483108 | -0.1001 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000414354 | UGP2 | ENSG00000169764 | ENST00000467400 | -0.1298 | 1 | - | CHOL,HNSC,PCPG,COAD,READ | | ENSG00000228109 | ENST00000414354 | BTD | ENSG00000169814 | ENST00000417015 | -0.1435 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000414354 | BTD | ENSG00000169814 | ENST00000449107 | -0.1650 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000414354 | BTD | ENSG00000169814 | ENST00000303498 | -0.1221 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000414354 | BTD | ENSG00000169814 | ENST00000437172 | -0.1221 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000414354 | BTD | ENSG00000169814 | ENST00000383778 | -0.1221 | 1 | - | CHOL,ESCA,LUSC,COAD,STAD | | ENSG00000228109 | ENST00000414354 | SDHD | ENSG00000204370 | ENST00000534010 | -0.1130 | 13 | - | CHOL,COAD,READ | | ENSG00000228109 | ENST00000414354 | LPCAT3 | ENSG00000111684 | ENST00000536797 | -0.1097 | 1 | - | CHOL,LUAD,COAD | | ENSG00000228109 | ENST00000414354 | MIER3 | ENSG00000155545 | ENST00000451637 | -0.1298 | 85 | - | COAD,READ | | ENSG00000228109 | ENST00000414354 | CCNYL1 | ENSG00000163249 | ENST00000420321 | -0.3082 | 1 | - | COAD,READ | | ENSG00000228109 | ENST00000414354 | PDLIM5 | ENSG00000163110 | ENST00000512274 | -0.1304 | 6 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000424769 | PRKAR1A | ENSG00000108946 | ENST00000588188 | -0.1099 | 1 | - | GBM | | ENSG00000228109 | ENST00000415244 | PDLIM5 | ENSG00000163110 | ENST00000509357 | -0.1118 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000415244 | PDLIM5 | ENSG00000163110 | ENST00000506632 | -0.1121 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000415244 | PRKAR1A | ENSG00000108946 | ENST00000588188 | -0.1136 | 128 | - | GBM | | ENSG00000228109 | ENST00000415244 | MYO18A | ENSG00000196535 | ENST00000533112 | -0.1003 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000415244 | MYO18A | ENSG00000196535 | ENST00000531253 | -0.1003 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000415244 | MYO18A | ENSG00000196535 | ENST00000527372 | -0.1003 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000415244 | MYO18A | ENSG00000196535 | ENST00000628822 | -0.1003 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000415244 | MYO18A | ENSG00000196535 | ENST00000588791 | -0.1486 | 112 | - | GBM,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000627587 | -0.1461 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000542407 | -0.1405 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000514743 | -0.1405 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000509357 | -0.1461 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PDLIM5 | ENSG00000163110 | ENST00000506632 | -0.1123 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000446695 | PGM2 | ENSG00000169299 | ENST00000381967 | -0.1275 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | PGM2 | ENSG00000169299 | ENST00000515668 | -0.1099 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | PGM2 | ENSG00000169299 | ENST00000505986 | -0.1099 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | PGM2 | ENSG00000169299 | ENST00000512556 | -0.1099 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | PGM2 | ENSG00000169299 | ENST00000510084 | -0.1142 | 1 | - | PCPG | | ENSG00000228109 | ENST00000446695 | ATE1 | ENSG00000107669 | ENST00000224652 | -0.1084 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | ATE1 | ENSG00000107669 | ENST00000369040 | -0.1084 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | ATE1 | ENSG00000107669 | ENST00000543447 | -0.1084 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | ATE1 | ENSG00000107669 | ENST00000540606 | -0.1084 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | ATE1 | ENSG00000107669 | ENST00000369043 | -0.1084 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PRKAR1A | ENSG00000108946 | ENST00000392710 | -0.1178 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PRKAR1A | ENSG00000108946 | ENST00000586397 | -0.1696 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PRKAR1A | ENSG00000108946 | ENST00000588188 | -0.2569 | 135 | - | GBM | | ENSG00000228109 | ENST00000446695 | PRKAR1A | ENSG00000108946 | ENST00000592800 | -0.1005 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | PRKAR1A | ENSG00000108946 | ENST00000586541 | -0.1178 | 1 | - | GBM | | ENSG00000228109 | ENST00000446695 | TGFBR2 | ENSG00000163513 | ENST00000295754 | -0.1081 | 1 | - | BLCA,ESCA,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | TGFBR2 | ENSG00000163513 | ENST00000359013 | -0.1081 | 1 | - | BLCA,ESCA,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | MYO18A | ENSG00000196535 | ENST00000533112 | -0.1272 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000446695 | MYO18A | ENSG00000196535 | ENST00000531253 | -0.1272 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000446695 | MYO18A | ENSG00000196535 | ENST00000527372 | -0.1272 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000446695 | MYO18A | ENSG00000196535 | ENST00000628822 | -0.1272 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000446695 | MYO18A | ENSG00000196535 | ENST00000530254 | -0.1948 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000446695 | MYO18A | ENSG00000196535 | ENST00000588791 | -0.1731 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000437064 | PDLIM5 | ENSG00000163110 | ENST00000509357 | -0.1006 | 1 | - | CHOL,BLCA,HNSC | | ENSG00000228109 | ENST00000437064 | PGM2 | ENSG00000169299 | ENST00000510084 | -0.1228 | 1 | - | PCPG | | ENSG00000228109 | ENST00000437064 | MYO18A | ENSG00000196535 | ENST00000530254 | -0.1034 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000414354 | PRKAR1A | ENSG00000108946 | ENST00000588188 | -0.2979 | 1 | - | GBM | | ENSG00000228109 | ENST00000414354 | MYO18A | ENSG00000196535 | ENST00000588791 | -0.1576 | 1 | - | GBM,HNSC | | ENSG00000228109 | ENST00000424769 | IL6ST | ENSG00000134352 | ENST00000502326 | -0.2167 | 180 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000415244 | IL6ST | ENSG00000134352 | ENST00000502326 | -0.9487 | 359 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | SPTBN1 | ENSG00000115306 | ENST00000356805 | -0.1349 | 1 | - | GBM,PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | SPTBN1 | ENSG00000115306 | ENST00000615901 | -0.1349 | 1 | - | GBM,PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | SPTBN1 | ENSG00000115306 | ENST00000333896 | -0.1220 | 1 | - | GBM,PCPG,LUSC,LUAD | | ENSG00000228109 | ENST00000446695 | IL6ST | ENSG00000134352 | ENST00000381298 | -0.1061 | 1 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IL6ST | ENSG00000134352 | ENST00000336909 | -0.1061 | 1 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IL6ST | ENSG00000134352 | ENST00000381287 | -0.1024 | 1 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IL6ST | ENSG00000134352 | ENST00000503773 | -0.1024 | 1 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000446695 | IL6ST | ENSG00000134352 | ENST00000502326 | -0.1147 | 59 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000437064 | IL6ST | ENSG00000134352 | ENST00000502326 | -0.2167 | 128 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ | | ENSG00000228109 | ENST00000414354 | IL6ST | ENSG00000134352 | ENST00000502326 | -0.9487 | 143 | - | CHOL,BLCA,HNSC,ESCA,LUSC,COAD,READ |
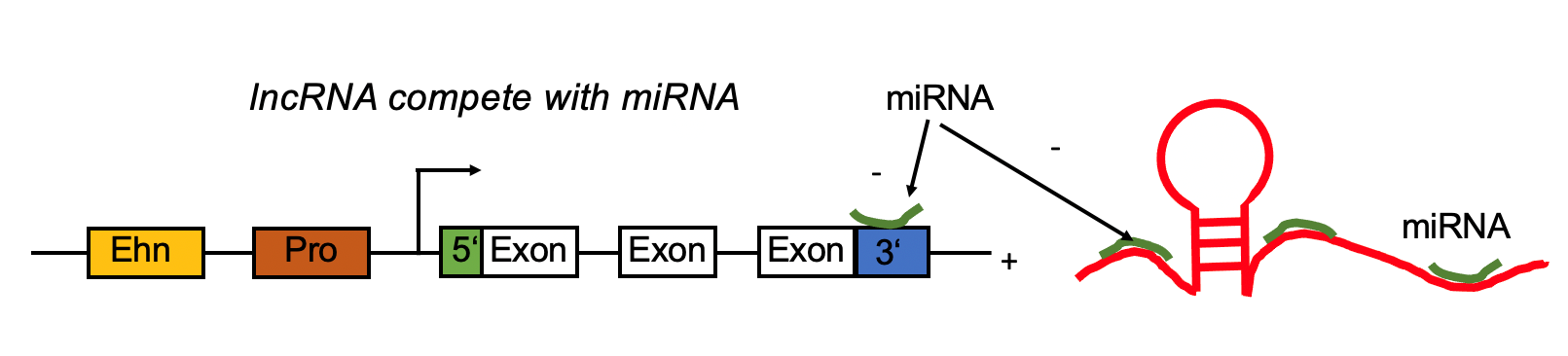
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,WDR33 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,ZHX2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RXRB | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CRMP1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,C1orf35 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PHF2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,DDX42 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,KAT7 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,COIL | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,TRMT10B | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBM6 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,USP21 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,YTHDC1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,LCMT2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CBX1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PRR14 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,C2orf68 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PM20D2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RPRD2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,INO80D | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RING1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,KANSL1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,ZNF7 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,NAA16 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,FAM117B | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,SENP7 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,BOLA1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PARP2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,KAT2A | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,TSEN2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,ERCC3 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PHC1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,EXD3 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBM4B | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,COA5 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RAD52 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PCF11 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,HEXIM2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,VEZF1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,NAB1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PAXBP1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,LUC7L3 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PAGR1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,ING5 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CLK2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,LUC7L2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,SENP6 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,KHDC4 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CEP95 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,EIF4ENIF1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,TIA1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,DPF2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RSBN1L | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,MSL1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RC3H1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBBP6 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,EDRF1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CTPS2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,SBK1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,TAF7 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,POLG2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,MLLT6 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,NXF1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,NVL | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,MYEF2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,MRPL38 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBMX | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PMS1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,UBE2G2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,FBXO46 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PHF12 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,MSL2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBM45 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBM5 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CXXC1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,COPS7B | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RCOR3 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,ANGEL2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,C14orf93 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RBM14 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,HINFP | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,UIMC1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,MKS1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PCIF1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,SF3B1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,FNBP4 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,SRSF1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,DGCR8 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,DET1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,POGZ | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,BRD8 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,TBP | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,RYK | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CTDSPL2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,BUD13 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,LYSMD1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,DCAF8 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,NSMCE4A | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,CASC3 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,N4BP2L2 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,PRPF39 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,WBP1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,NFRKB | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,ZBTB5 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,SMAD4 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-22,KATNBL1 | KICH,CHOL,LGG,SARC,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-378a,CRTC1 | PCPG,TGCT,CHOL,THCA,KIRP | | ENSG00000228109 | MELTF-AS1,hsa-mir-378a,PLXNB1 | PCPG,TGCT,CHOL,THCA,KIRP |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000228109.1 | ENST00000424769.1 | 130 | 389 | 0 | | ENSG00000228109.1 | ENST00000415244.1 | 94 | 0 | 284 | | ENSG00000228109.1 | ENST00000446695.1 | 105 | 1 | 318 | | ENSG00000228109.1 | ENST00000437064.1 | 57 | 170 | 0 | | ENSG00000228109.1 | ENST00000414354.1 | 44 | 98 | 232 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).