lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | ZNF30,CEBPZ,ILF3,GLI4,RBM39,EDC3,CIRBP,LCORL,MZF1,MYSM1,NRF1,ZNF2,THOC3,ARID2,ZXDC,DMTF1,RBM27,ZNF77,ZNF24,DDX52,RBM28,TBP,GABPA,IKZF5,DDX55,ZFP14,ZFP69,SAFB2,THOC1,LARP7,TRNT1,PNPT1,CNOT4,CHTOP,CWC22,DDX59,LSM5,ZZZ3,ZBED5,PA2G4,CENPT,TTC14,RC3H1,ZNF44,EEA1,ZNF91,RBMX,RBM4B,U2AF2,CSTF3,TIGD6,RBM26,ZFP1,RBM14,PURB,GZF1,NCBP1,MYEF2,PPIL4,CWC15,PUS7L,NFYC,DHX34,NFIA,FUBP1,RBM15,ZFP30,SP4,PPIE,CPSF7,LIN54,THAP9,THOC2,DDX6,TSN,ZBTB6,ZNF34,THAP6,DDX17,E2F5,E2F6,ZNF57,ZNF22,FMR1,LCOR,MECP2,PHF1,PURA,THYN1,ZNF41,ZC3H8,TTF1,HSF4,RBM41,THAP1,DZIP3,ILF2,ZFP62,PAIP1,ZNF28,RBM3,CELF1,PSPC1,KRR1,TMF1,ELK4,TIGD1,NFXL1,MBD4,AGO3,UNC50,SRSF2,DDX31,MYNN,TFCP2,ZMAT1,ZNF83,SKIL,NSUN6,ZNF3,ZFX,ZNF8,ZNF18,NKRF,NR2C2,EIF2D,PCGF2,GPBP1,C1D,RBM6,HMGN3,CPSF3,ZNF14,RBM48,PUM2,XPO1,RBM5,BRCA1,ELF2,SMAD4,NFYB,ZNF43,ZNF17,ZNF84,DDX42,BAZ2B,TRA2A,RBM8A,SMYD3,SRSF7,HINFP,XPA,NR2C1,THOC7,TFDP2,ZFP3,PCGF6,SFPQ,RBM33,NOL8,ATF2,ZNF10 | | Exist in public source | | NA |
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | E2F5,DDX17,MZF1,ZNF18,E4F1,RBM5,CNOT6,MECP2,DDX52,ATF1,U2AF2,MBNL1,ZMAT1,THOC2,DKC1,ZNF14,ZNF85,TFCP2,ZNF57,ZFP1,FUBP1,LCOR,MYNN,DDX46,ZNF3,CPSF7,PSPC1,RC3H1,NSUN6,ZNF83,RBM6,PPIL4,TERF1,RBM3,EIF2D,AGO4,SMYD3,BRCA1,NFIA,CELF1,ARID2,LIN54,CIRBP,LCORL,SRP14 | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
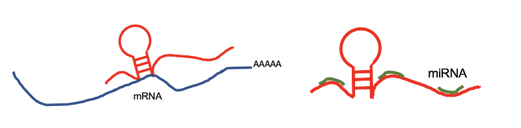
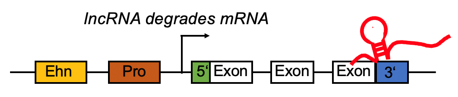
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | MAZ,RNH1,CIC | | Exist in public source | | NA |
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation |
| -lncRNA and exonskipping events are negatively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000225470 | ENST00000602546 | exon_skip_5818 | chr1:45568484-45568684 | -17.28 | -0.1529 | AKR1A1 | ENST00000351829,ENST00000372070 | Frame-shift | | ENSG00000225470 | ENST00000538519 | exon_skip_66185 | chr11:124626472-124626609 | -15.00 | -0.1562 | TBRG1 | ENST00000441174 | Frame-shift | | ENSG00000225470 | ENST00000538519 | exon_skip_382458 | chr3:38131595-38131638 | -9.55 | -0.3537 | ACAA1 | ENST00000333167 | Frame-shift | | ENSG00000225470 | ENST00000538519 | exon_skip_353131 | chr20:58669290-58669453 | -19.69 | -0.1304 | STX16 | ENST00000371141 | Frame-shift | | ENSG00000225470 | ENST00000538519 | exon_skip_470226 | chr7:103307595-103307708 | -9.24 | -0.2149 | PMPCB | ENST00000249269 | Frame-shift | | ENSG00000225470 | ENST00000538519 | exon_skip_380242 | chr3:186784961-186785101 | -9.14 | -0.1632 | EIF4A2 | ENST00000323963 | Frame-shift | | ENSG00000225470 | ENST00000538519 | exon_skip_319223 | chr19:43526200-43526349 | -19.27 | -0.1386 | ETHE1 | ENST00000292147 | Frame-shift |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
 |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000225470 | hsa-mir-22 | ENST00000602938 | chrX:73944559-73944645 | 177.00 | -73.99 | 71 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602938 | chrX:73944737-73944820 | 169.00 | -75.10 | 80 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602938 | chrX:73944479-73944571 | 152.00 | -73.28 | 89 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602938 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602294 | chrX:73944559-73944645 | 177.00 | -73.99 | 71 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602294 | chrX:73944479-73944571 | 152.00 | -73.28 | 89 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602294 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602920 | chrX:73944559-73944645 | 177.00 | -73.99 | 71 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602920 | chrX:73944479-73944571 | 152.00 | -73.28 | 89 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602920 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602737 | chrX:73944656-73944739 | 180.00 | -77.77 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602737 | chrX:73944770-73944856 | 162.00 | -75.36 | 78 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602737 | chrX:73944430-73944521 | 161.00 | -83.11 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73945818-73945905 | 186.00 | -79.54 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73944559-73944645 | 177.00 | -73.99 | 71 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73944734-73944824 | 162.00 | -73.34 | 84 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73945528-73945620 | 162.00 | -66.64 | 83 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73944986-73945069 | 157.00 | -78.06 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73945620-73945707 | 157.00 | -79.12 | 87 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73945294-73945375 | 156.00 | -69.13 | 83 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73944479-73944571 | 152.00 | -73.28 | 89 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73945081-73945179 | 152.00 | -83.87 | 97 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73944873-73944951 | 151.00 | -67.10 | 52 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602772 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73944538-73944621 | 180.00 | -77.77 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73944991-73945074 | 169.00 | -75.10 | 80 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73945133-73945208 | 168.00 | -81.98 | 82 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73944678-73944756 | 166.00 | -86.54 | 83 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73945320-73945401 | 165.00 | -68.78 | 82 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73946030-73946125 | 161.00 | -66.36 | 93 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73945216-73945303 | 157.00 | -76.90 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73945538-73945617 | 156.00 | -68.70 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73944863-73944951 | 155.00 | -64.75 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73945430-73945509 | 152.00 | -61.44 | 83 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000414209 | chrX:73946370-73946460 | 147.00 | -67.46 | 83 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73944615-73944698 | 169.00 | -75.10 | 80 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73944479-73944559 | 168.00 | -70.50 | 81 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73944757-73944832 | 168.00 | -81.98 | 82 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73944944-73945025 | 165.00 | -68.78 | 82 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73944840-73944927 | 157.00 | -76.90 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73945054-73945133 | 152.00 | -61.44 | 83 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602895 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000453317 | chrX:73944553-73944649 | 154.00 | -76.92 | 92 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000453317 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000453317 | chrX:73944479-73944556 | 147.00 | -69.44 | 80 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602546 | chrX:73944508-73944594 | 177.00 | -73.99 | 71 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602546 | chrX:73944686-73944769 | 169.00 | -75.10 | 80 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602546 | chrX:73944828-73944903 | 168.00 | -81.98 | 82 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602546 | chrX:73945015-73945096 | 165.00 | -68.78 | 82 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602546 | chrX:73944911-73944998 | 157.00 | -76.90 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602546 | chrX:73944428-73944520 | 152.00 | -73.28 | 89 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000602985 | chrX:73944359-73944451 | 150.00 | -77.35 | 85 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000415215 | chrX:73944538-73944621 | 180.00 | -77.77 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000415215 | chrX:73944422-73944508 | 150.00 | -61.59 | 81 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000538519 | chrX:73944656-73944739 | 180.00 | -77.77 | 86 | NA | | ENSG00000225470 | hsa-mir-22 | ENST00000538519 | chrX:73944770-73944856 | 162.00 | -75.36 | 78 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name | | LncEditing_399571 | ENSG00000225470.5 | chrX | 73952148 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 | | LncEditing_400467 | ENSG00000225470.5 | chrX | 73956637 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 | | LncEditing_400491 | ENSG00000225470.5 | chrX | 73956852 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 | | LncEditing_400547 | ENSG00000225470.5 | chrX | 73956961 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 | | LncEditing_400643 | ENSG00000225470.5 | chrX | 73957388 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 | | LncEditing_400667 | ENSG00000225470.5 | chrX | 73957498 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 | | LncEditing_400675 | ENSG00000225470.5 | chrX | 73957510 | + | lincRNA | JPX | ENST00000602985.4 | lincRNA | JPX-008 |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
 |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change | | ENSG00000225470 | LncEditing_399571 | LAML | chrX_73952148_+ | 1.386423e+01 | 8.593942e+00 | 1.449327e+00 | 4.990299e+00 | 6.028603e-07 | 1.448507e-06 | UP | | ENSG00000225470 | LncEditing_400467 | LAML | chrX_73956637_+ | 1.102511e+01 | 8.993460e+00 | 3.403162e+00 | 2.142858e+00 | 3.212452e-02 | 3.300647e-02 | UP | | ENSG00000225470 | LncEditing_400491 | LAML | chrX_73956852_+ | 1.100567e+01 | 8.416191e+00 | 2.583926e+00 | 4.388599e+00 | 1.140834e-05 | 2.141230e-05 | UP | | ENSG00000225470 | LncEditing_400547 | LAML | chrX_73956961_+ | 1.222290e+01 | 8.790268e+00 | 2.102572e+00 | 2.861378e+00 | 4.218038e-03 | 4.942388e-03 | UP | | ENSG00000225470 | LncEditing_400643 | LAML | chrX_73957388_+ | 1.247558e+01 | 8.809445e+00 | 1.992096e+00 | 3.320252e+00 | 8.993616e-04 | 1.181608e-03 | UP | | ENSG00000225470 | LncEditing_400667 | LAML | chrX_73957498_+ | 1.158333e+01 | 8.612227e+00 | 2.338680e+00 | 3.703850e+00 | 2.123517e-04 | 3.140771e-04 | UP | | ENSG00000225470 | LncEditing_400675 | LAML | chrX_73957510_+ | 1.170542e+01 | 8.886112e+00 | 2.515391e+00 | 3.439803e+00 | 5.821376e-04 | 7.952876e-04 | UP |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
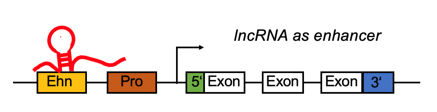
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation |
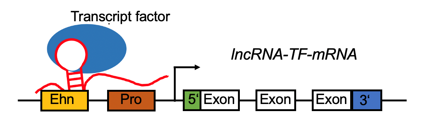
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000133740 | E2F5 | 5 | COAD,KIRC,KIRP,PAAD,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000133740 | E2F5 | 7 | COAD,KIRC,KIRP,LAML,PAAD,SKCM,THCA | | ENSG00000225470 | ENSG00000186416 | NKRF | ENSG00000133740 | E2F5 | 5 | COAD,KIRP,LAML,PAAD,THCA | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000133740 | E2F5 | 6 | COAD,KIRC,KIRP,LAML,PAAD,THCA | | ENSG00000225470 | ENSG00000117000 | RLF | ENSG00000100201 | DDX17 | 6 | COAD,KICH,KIRP,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000100201 | DDX17 | 16 | ACC,BRCA,CHOL,COAD,ESCA,HNSC,KICH,KIRC,KIRP,LAML,LGG,LIHC,LUAD,PAAD,SARC,SKCM | | ENSG00000225470 | ENSG00000036549 | ZZZ3 | ENSG00000100201 | DDX17 | 5 | COAD,ESCA,KICH,LAML,UCEC | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000100201 | DDX17 | 5 | HNSC,KIRC,KIRP,LIHC,PAAD | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000100201 | DDX17 | 8 | COAD,HNSC,KICH,LAML,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000100201 | DDX17 | 7 | BRCA,CHOL,HNSC,KIRC,KIRP,LIHC,SKCM | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000100201 | DDX17 | 10 | ACC,BRCA,CHOL,COAD,ESCA,LIHC,LUAD,PRAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000114126 | TFDP2 | ENSG00000100201 | DDX17 | 5 | COAD,HNSC,KICH,LAML,LIHC | | ENSG00000225470 | ENSG00000120784 | ZFP30 | ENSG00000100201 | DDX17 | 5 | COAD,KICH,KIRC,KIRP,LAML | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000100201 | DDX17 | 7 | COAD,KICH,KIRC,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000100201 | DDX17 | 11 | BRCA,ESCA,KICH,KIRC,KIRP,LAML,LUAD,PAAD,PRAD,SARC,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000100201 | DDX17 | 10 | COAD,HNSC,LAML,LGG,LIHC,LUAD,PAAD,SARC,SKCM,UCEC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000100201 | DDX17 | 5 | COAD,HNSC,LGG,LIHC,SKCM | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000100201 | DDX17 | 5 | CHOL,COAD,LGG,LUAD,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000100201 | DDX17 | 7 | HNSC,KIRC,KIRP,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000100201 | DDX17 | 9 | COAD,ESCA,HNSC,LAML,LGG,LIHC,LUAD,PAAD,SKCM | | ENSG00000225470 | ENSG00000166526 | ZNF3 | ENSG00000100201 | DDX17 | 5 | ACC,HNSC,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000100201 | DDX17 | 9 | BRCA,CHOL,COAD,KIRP,LAML,LUAD,PAAD,PRAD,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000100201 | DDX17 | 10 | BRCA,COAD,KICH,KIRC,KIRP,LIHC,LUAD,PRAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000100201 | DDX17 | 5 | KIRP,LAML,LIHC,PAAD,SKCM | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000100201 | DDX17 | 6 | BRCA,COAD,KICH,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000100201 | DDX17 | 6 | COAD,ESCA,KICH,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000100201 | DDX17 | 7 | COAD,KIRC,LAML,LUAD,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000100201 | DDX17 | 11 | BRCA,COAD,HNSC,KICH,KIRP,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000100201 | DDX17 | 15 | BRCA,CHOL,COAD,ESCA,HNSC,KIRC,KIRP,LAML,LGG,LIHC,LUAD,PAAD,PRAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198455 | ZXDB | ENSG00000100201 | DDX17 | 6 | ACC,COAD,ESCA,HNSC,LAML,SKCM | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000100201 | DDX17 | 8 | COAD,HNSC,KICH,KIRP,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000100201 | DDX17 | 12 | BRCA,CHOL,COAD,HNSC,KICH,LAML,LGG,LIHC,LUAD,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000100201 | DDX17 | 7 | COAD,ESCA,KICH,LUAD,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000100201 | DDX17 | 7 | CHOL,COAD,ESCA,LAML,LIHC,SARC,SKCM | | ENSG00000225470 | ENSG00000158711 | ELK4 | ENSG00000100201 | DDX17 | 6 | COAD,ESCA,LAML,LGG,LUAD,PRAD | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000100201 | DDX17 | 12 | BRCA,COAD,ESCA,HNSC,KICH,LAML,LGG,LIHC,LUAD,PAAD,PRAD,SKCM | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000100201 | DDX17 | 8 | BRCA,KIRC,KIRP,LAML,LGG,LUAD,PRAD,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000100201 | DDX17 | 15 | BRCA,CHOL,COAD,HNSC,KICH,KIRC,KIRP,LAML,LGG,LIHC,LUAD,PAAD,PRAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000100201 | DDX17 | 11 | BRCA,CHOL,KIRC,LAML,LGG,LUAD,PAAD,PRAD,SARC,SKCM,UCEC | | ENSG00000225470 | ENSG00000196233 | LCOR | ENSG00000100201 | DDX17 | 5 | BRCA,COAD,KIRC,LIHC,LUAD | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000099326 | MZF1 | 6 | HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000099326 | MZF1 | 6 | BRCA,KIRC,KIRP,LIHC,SKCM,THCA | | ENSG00000225470 | ENSG00000167967 | E4F1 | ENSG00000099326 | MZF1 | 5 | CHOL,HNSC,KIRC,PAAD,THCA | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000099326 | MZF1 | 6 | BLCA,KIRP,LIHC,PAAD,SKCM,UCS | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000099326 | MZF1 | 9 | BLCA,CHOL,HNSC,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000099326 | MZF1 | 5 | HNSC,LIHC,PAAD,SKCM,THCA | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000099326 | MZF1 | 8 | BLCA,CHOL,HNSC,KIRC,KIRP,LIHC,SKCM,THCA | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000099326 | MZF1 | 6 | BLCA,KIRC,PAAD,SKCM,THCA,UCS | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000154957 | ZNF18 | 5 | HNSC,PAAD,READ,THCA,UCEC | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000154957 | ZNF18 | 6 | HNSC,KIRC,PAAD,READ,THCA,UCEC | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000167967 | E4F1 | 5 | CHOL,HNSC,KIRC,PAAD,THCA | | ENSG00000225470 | ENSG00000117000 | RLF | ENSG00000003756 | RBM5 | 6 | COAD,KICH,KIRP,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000003756 | RBM5 | 9 | COAD,KICH,KIRC,LAML,LIHC,LUAD,PAAD,SARC,SKCM | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000003756 | RBM5 | 6 | KIRC,KIRP,LIHC,PAAD,PRAD,THCA | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000003756 | RBM5 | 8 | BLCA,BRCA,KIRC,KIRP,LIHC,PAAD,SKCM,THCA | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000003756 | RBM5 | 7 | BRCA,COAD,LIHC,LUAD,PRAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000120784 | ZFP30 | ENSG00000003756 | RBM5 | 5 | COAD,KICH,KIRC,KIRP,LAML | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000003756 | RBM5 | 7 | COAD,KICH,KIRC,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000003756 | RBM5 | 9 | BRCA,KICH,KIRC,KIRP,LAML,LUAD,PAAD,SARC,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000003756 | RBM5 | 6 | COAD,LAML,LIHC,LUAD,PAAD,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000003756 | RBM5 | 9 | BLCA,COAD,KIRC,KIRP,LAML,LUAD,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000154957 | ZNF18 | ENSG00000003756 | RBM5 | 5 | BRCA,KIRC,PAAD,THCA,UCEC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000003756 | RBM5 | 5 | COAD,LAML,LIHC,PAAD,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000003756 | RBM5 | 7 | BRCA,COAD,KIRP,LUAD,PAAD,PRAD,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000003756 | RBM5 | 12 | BRCA,COAD,KICH,KIRC,KIRP,LAML,LIHC,LUAD,PRAD,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000003756 | RBM5 | 7 | BLCA,KIRP,LAML,LIHC,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000003756 | RBM5 | 5 | BRCA,COAD,KICH,LUAD,SKCM | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000003756 | RBM5 | 5 | COAD,KICH,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000003756 | RBM5 | 6 | COAD,KIRC,LAML,LUAD,PAAD,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000003756 | RBM5 | 6 | BRCA,COAD,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000003756 | RBM5 | 13 | BLCA,BRCA,COAD,KIRC,KIRP,LAML,LIHC,LUAD,PAAD,PRAD,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000003756 | RBM5 | 7 | COAD,KICH,LAML,LIHC,LUAD,PAAD,SKCM | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000003756 | RBM5 | 6 | COAD,KICH,LUAD,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000003756 | RBM5 | 5 | COAD,LAML,LIHC,SARC,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000003756 | RBM5 | 10 | BRCA,COAD,KICH,LAML,LIHC,LUAD,PAAD,PRAD,SKCM,THCA | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000003756 | RBM5 | 6 | BRCA,KIRC,KIRP,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000003756 | RBM5 | 14 | BLCA,BRCA,COAD,KICH,KIRC,KIRP,LAML,LIHC,LUAD,PAAD,PRAD,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000003756 | RBM5 | 11 | BLCA,BRCA,KIRC,LAML,LUAD,PAAD,PRAD,SARC,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000196233 | LCOR | ENSG00000003756 | RBM5 | 5 | BRCA,COAD,KIRC,LIHC,LUAD | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000113300 | CNOT6 | 6 | COAD,LAML,LGG,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000113300 | CNOT6 | 6 | COAD,LAML,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000113300 | CNOT6 | 6 | COAD,LAML,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000113300 | CNOT6 | 5 | COAD,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000113300 | CNOT6 | 7 | COAD,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000113300 | CNOT6 | 6 | COAD,LAML,LGG,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000113300 | CNOT6 | 5 | COAD,LAML,LGG,LUAD,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000113300 | CNOT6 | 6 | COAD,LAML,LGG,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000113300 | CNOT6 | 5 | COAD,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000113300 | CNOT6 | 7 | COAD,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000113300 | CNOT6 | 7 | COAD,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000113300 | CNOT6 | 5 | COAD,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000113300 | CNOT6 | 7 | COAD,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000113300 | CNOT6 | 6 | COAD,LAML,LGG,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000113300 | CNOT6 | 7 | COAD,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000169057 | MECP2 | 6 | COAD,ESCA,HNSC,LAML,PAAD,UCS | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000169057 | MECP2 | 5 | COAD,ESCA,HNSC,LAML,PAAD | | ENSG00000225470 | ENSG00000198455 | ZXDB | ENSG00000169057 | MECP2 | 5 | COAD,ESCA,HNSC,LAML,UCS | | ENSG00000225470 | ENSG00000186416 | NKRF | ENSG00000169057 | MECP2 | 5 | COAD,ESCA,HNSC,PAAD,UCS | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000278053 | DDX52 | 7 | COAD,DLBC,LAML,LGG,LIHC,SARC,SKCM | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LGG,LIHC,SKCM | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000278053 | DDX52 | 6 | COAD,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000278053 | DDX52 | 7 | COAD,DLBC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LGG,READ,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000278053 | DDX52 | 6 | COAD,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LGG,LIHC,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LGG,LIHC,SKCM | | ENSG00000225470 | ENSG00000198455 | ZXDB | ENSG00000278053 | DDX52 | 5 | COAD,DLBC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000278053 | DDX52 | 5 | COAD,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000278053 | DDX52 | 6 | COAD,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000278053 | DDX52 | 6 | COAD,LAML,LIHC,READ,SARC,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000278053 | DDX52 | 5 | COAD,LAML,LGG,LIHC,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000278053 | DDX52 | 6 | COAD,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000152601 | MBNL1 | 5 | COAD,KICH,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000152601 | MBNL1 | 5 | COAD,KICH,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000152601 | MBNL1 | 5 | COAD,KICH,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000152601 | MBNL1 | 5 | COAD,KICH,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000166432 | ZMAT1 | 5 | ACC,LUAD,PAAD,SARC,THYM | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000166432 | ZMAT1 | 6 | KIRC,KIRP,PAAD,PCPG,PRAD,THCA | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000166432 | ZMAT1 | 6 | BRCA,KIRC,KIRP,PAAD,THCA,UCS | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000166432 | ZMAT1 | 5 | BRCA,LAML,LUAD,PRAD,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000166432 | ZMAT1 | 9 | ACC,BRCA,KIRC,KIRP,LAML,LUAD,PAAD,PRAD,SARC | | ENSG00000225470 | ENSG00000154957 | ZNF18 | ENSG00000166432 | ZMAT1 | 5 | BRCA,KIRC,PAAD,THCA,UCEC | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000166432 | ZMAT1 | 6 | BRCA,KIRP,LAML,LUAD,PAAD,PRAD | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000166432 | ZMAT1 | 8 | BRCA,KIRC,KIRP,LUAD,PCPG,PRAD,THCA,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000166432 | ZMAT1 | 10 | BRCA,CESC,KIRC,KIRP,LAML,LUAD,PAAD,PRAD,THCA,UCEC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000166432 | ZMAT1 | 6 | BRCA,LAML,LUAD,PAAD,PRAD,THCA | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000166432 | ZMAT1 | 5 | BRCA,KIRC,KIRP,LAML,LUAD | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000166432 | ZMAT1 | 10 | BRCA,KIRC,KIRP,LAML,LUAD,PAAD,PCPG,PRAD,THCA,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000166432 | ZMAT1 | 12 | BRCA,CESC,KIRC,LAML,LUAD,PAAD,PRAD,SARC,THCA,THYM,UCEC,UCS | | ENSG00000225470 | ENSG00000117000 | RLF | ENSG00000125676 | THOC2 | 6 | COAD,KICH,KIRP,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000125676 | THOC2 | 13 | BRCA,COAD,HNSC,KICH,KIRC,KIRP,LAML,LGG,LIHC,LUAD,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000036549 | ZZZ3 | ENSG00000125676 | THOC2 | 6 | COAD,ESCA,KICH,LAML,READ,UCEC | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000125676 | THOC2 | 8 | COAD,HNSC,KICH,LAML,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000102189 | EEA1 | ENSG00000125676 | THOC2 | 5 | COAD,KICH,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000125676 | THOC2 | 7 | BRCA,COAD,LIHC,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000114126 | TFDP2 | ENSG00000125676 | THOC2 | 5 | COAD,KICH,LAML,LIHC,READ | | ENSG00000225470 | ENSG00000120784 | ZFP30 | ENSG00000125676 | THOC2 | 5 | COAD,KICH,KIRC,KIRP,LAML | | ENSG00000225470 | ENSG00000125482 | TTF1 | ENSG00000125676 | THOC2 | 7 | HNSC,KICH,LAML,LGG,LIHC,LUSC,READ | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000125676 | THOC2 | 7 | COAD,KICH,KIRC,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000125676 | THOC2 | 9 | BRCA,ESCA,KICH,KIRC,KIRP,LAML,LUAD,PAAD,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000125676 | THOC2 | 11 | COAD,ESCA,HNSC,LAML,LGG,LIHC,LUAD,PAAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000125676 | THOC2 | 9 | COAD,ESCA,HNSC,LAML,LGG,LIHC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000125676 | THOC2 | 6 | COAD,LAML,LGG,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000125676 | THOC2 | 10 | COAD,HNSC,KIRC,KIRP,LUAD,PAAD,READ,SKCM,STAD,UCEC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000125676 | THOC2 | 10 | COAD,ESCA,HNSC,LAML,LGG,LIHC,LUAD,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000166526 | ZNF3 | ENSG00000125676 | THOC2 | 7 | COAD,HNSC,LGG,LIHC,LUSC,READ,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000125676 | THOC2 | 8 | BRCA,COAD,KIRP,LAML,LUAD,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000125676 | THOC2 | 5 | BRCA,COAD,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000125676 | THOC2 | 6 | COAD,KIRP,LAML,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000125676 | THOC2 | 6 | BRCA,COAD,KICH,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000125676 | THOC2 | 6 | COAD,KICH,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000125676 | THOC2 | 7 | COAD,KIRC,LAML,LUAD,PAAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000125676 | THOC2 | 10 | BRCA,COAD,HNSC,KIRP,LAML,LGG,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000125676 | THOC2 | 16 | BRCA,CESC,COAD,ESCA,HNSC,KIRC,KIRP,LAML,LGG,LIHC,LUAD,PAAD,READ,SKCM,STAD,UCEC | | ENSG00000225470 | ENSG00000198455 | ZXDB | ENSG00000125676 | THOC2 | 8 | CESC,COAD,ESCA,HNSC,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000125676 | THOC2 | 9 | COAD,HNSC,KICH,KIRP,LGG,LIHC,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000025293 | PHF20 | ENSG00000125676 | THOC2 | 5 | COAD,LAML,LIHC,READ,STAD | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000125676 | THOC2 | 12 | BRCA,COAD,HNSC,KICH,LAML,LGG,LIHC,LUAD,PAAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000125676 | THOC2 | 7 | COAD,ESCA,KICH,PAAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000125676 | THOC2 | 6 | COAD,ESCA,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000135457 | TFCP2 | ENSG00000125676 | THOC2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000186416 | NKRF | ENSG00000125676 | THOC2 | 13 | CESC,COAD,ESCA,HNSC,KIRP,LAML,LUAD,LUSC,PAAD,READ,SKCM,STAD,UCEC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000125676 | THOC2 | 11 | BRCA,COAD,ESCA,KICH,LAML,LGG,LIHC,LUAD,PAAD,SKCM,STAD | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000125676 | THOC2 | 7 | BRCA,KIRC,KIRP,LAML,LGG,LUAD,SKCM | | ENSG00000225470 | ENSG00000198538 | ZNF28 | ENSG00000125676 | THOC2 | 5 | COAD,ESCA,KICH,LAML,LIHC | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000125676 | THOC2 | 14 | BRCA,COAD,HNSC,KICH,KIRC,KIRP,LAML,LGG,LIHC,LUAD,PAAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000125676 | THOC2 | 7 | BRCA,KIRC,LAML,LGG,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196233 | LCOR | ENSG00000125676 | THOC2 | 5 | BRCA,COAD,KIRC,LIHC,LUAD | | ENSG00000225470 | ENSG00000125482 | TTF1 | ENSG00000130826 | DKC1 | 5 | ESCA,HNSC,LGG,LIHC,LUSC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000130826 | DKC1 | 5 | HNSC,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000130826 | DKC1 | 5 | DLBC,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000130826 | DKC1 | 5 | COAD,ESCA,HNSC,KIRP,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000130826 | DKC1 | 5 | COAD,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000166526 | ZNF3 | ENSG00000130826 | DKC1 | 6 | COAD,LGG,LIHC,LUSC,READ,SKCM | | ENSG00000225470 | ENSG00000135457 | TFCP2 | ENSG00000130826 | DKC1 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000186416 | NKRF | ENSG00000130826 | DKC1 | 7 | COAD,ESCA,HNSC,KIRP,LUSC,READ,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000105708 | ZNF14 | 6 | ACC,KICH,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000105708 | ZNF14 | 6 | COAD,HNSC,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000105708 | ZNF14 | 5 | ACC,COAD,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000114126 | TFDP2 | ENSG00000105708 | ZNF14 | 5 | COAD,HNSC,KICH,LAML,READ | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000105708 | ZNF14 | 5 | ACC,KICH,LAML,PAAD,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000105708 | ZNF14 | 6 | COAD,HNSC,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000105708 | ZNF14 | 6 | ACC,COAD,HNSC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000105708 | ZNF14 | 6 | COAD,HNSC,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000166526 | ZNF3 | ENSG00000105708 | ZNF14 | 5 | ACC,COAD,HNSC,LAML,READ | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000105708 | ZNF14 | 5 | ACC,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000105708 | ZNF14 | 5 | COAD,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000105708 | ZNF14 | 6 | COAD,HNSC,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000105708 | ZNF14 | 5 | COAD,HNSC,KICH,READ,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000105708 | ZNF14 | 6 | COAD,KICH,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000105708 | ZNF14 | 5 | COAD,KICH,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000105708 | ZNF14 | 7 | COAD,HNSC,KICH,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000105708 | ZNF14 | 6 | COAD,KICH,LAML,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000105750 | ZNF85 | 6 | KIRC,LAML,LGG,LUAD,PRAD,UCEC | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000105750 | ZNF85 | 5 | KIRC,LAML,LGG,LUAD,PRAD | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000105750 | ZNF85 | 6 | KIRC,LAML,LGG,LUAD,PRAD,UCEC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000166526 | ZNF3 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000135457 | TFCP2 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000117000 | RLF | ENSG00000162613 | FUBP1 | 5 | COAD,KICH,KIRP,LIHC,LUAD | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000162613 | FUBP1 | 8 | COAD,ESCA,KICH,KIRC,KIRP,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000120784 | ZFP30 | ENSG00000162613 | FUBP1 | 5 | COAD,KICH,KIRC,KIRP,LAML | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000162613 | FUBP1 | 5 | COAD,KICH,KIRC,LAML,LUAD | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000162613 | FUBP1 | 5 | KICH,KIRC,KIRP,LAML,LUAD | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,KICH,LAML,LIHC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000162613 | FUBP1 | 6 | COAD,ESCA,KIRP,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000162613 | FUBP1 | 7 | COAD,ESCA,KIRC,KIRP,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,KICH,KIRP,LIHC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000162613 | FUBP1 | 5 | COAD,KICH,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000186416 | NKRF | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,KIRP,LAML,LUAD | | ENSG00000225470 | ENSG00000198538 | ZNF28 | ENSG00000162613 | FUBP1 | 5 | COAD,ESCA,KICH,LAML,LIHC | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000162613 | FUBP1 | 8 | COAD,ESCA,KICH,KIRC,KIRP,LAML,LIHC,LUAD | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000196233 | LCOR | 6 | BRCA,COAD,KIRC,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000196233 | LCOR | 5 | BRCA,COAD,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000196233 | LCOR | 5 | BRCA,COAD,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000196233 | LCOR | 6 | BRCA,COAD,KIRC,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000196233 | LCOR | 5 | BRCA,COAD,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000196233 | LCOR | 5 | BRCA,COAD,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000196233 | LCOR | 5 | BRCA,KIRC,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000085274 | MYNN | 7 | COAD,HNSC,KICH,LAML,LGG,LIHC,SKCM | | ENSG00000225470 | ENSG00000036549 | ZZZ3 | ENSG00000085274 | MYNN | 5 | COAD,KICH,LAML,READ,UCEC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000085274 | MYNN | 6 | COAD,LAML,LIHC,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000114126 | TFDP2 | ENSG00000085274 | MYNN | 5 | COAD,HNSC,KICH,LAML,LIHC | | ENSG00000225470 | ENSG00000125482 | TTF1 | ENSG00000085274 | MYNN | 6 | HNSC,KICH,LAML,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000085274 | MYNN | 5 | COAD,KICH,LAML,SKCM,UCEC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000085274 | MYNN | 7 | COAD,HNSC,LAML,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000085274 | MYNN | 7 | COAD,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000085274 | MYNN | 5 | COAD,LAML,LGG,READ,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000085274 | MYNN | 7 | COAD,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000085274 | MYNN | 5 | COAD,KICH,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000085274 | MYNN | 6 | COAD,KICH,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000085274 | MYNN | 7 | COAD,HNSC,LAML,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000085274 | MYNN | 7 | COAD,HNSC,LAML,LGG,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000085274 | MYNN | 8 | COAD,HNSC,KICH,LGG,LIHC,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000085274 | MYNN | 9 | COAD,HNSC,KICH,LAML,LGG,LIHC,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000085274 | MYNN | 5 | COAD,KICH,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000105708 | ZNF14 | ENSG00000085274 | MYNN | 6 | COAD,HNSC,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000085274 | MYNN | 5 | COAD,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000085274 | MYNN | 8 | COAD,HNSC,KICH,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000085274 | MYNN | 9 | COAD,HNSC,KICH,LAML,LGG,LIHC,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000145833 | DDX46 | 5 | COAD,KICH,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000125482 | TTF1 | ENSG00000145833 | DDX46 | 5 | ESCA,KICH,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000145833 | DDX46 | 5 | COAD,ESCA,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000145833 | DDX46 | 5 | COAD,ESCA,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000145833 | DDX46 | 5 | COAD,ESCA,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000145833 | DDX46 | 6 | COAD,ESCA,KICH,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000145833 | DDX46 | 5 | COAD,KICH,LAML,LGG,LIHC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000145833 | DDX46 | 5 | COAD,ESCA,KICH,LGG,LIHC | | ENSG00000225470 | ENSG00000198538 | ZNF28 | ENSG00000145833 | DDX46 | 5 | COAD,ESCA,KICH,LAML,LIHC | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000145833 | DDX46 | 5 | COAD,ESCA,KICH,LGG,LIHC | | ENSG00000225470 | ENSG00000114126 | TFDP2 | ENSG00000166526 | ZNF3 | 5 | COAD,HNSC,LAML,LIHC,READ | | ENSG00000225470 | ENSG00000125482 | TTF1 | ENSG00000166526 | ZNF3 | 6 | HNSC,LAML,LGG,LIHC,LUSC,READ | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000166526 | ZNF3 | 7 | COAD,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000166526 | ZNF3 | 9 | ACC,COAD,ESCA,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000166526 | ZNF3 | 6 | BLCA,COAD,HNSC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000166526 | ZNF3 | 8 | COAD,HNSC,LAML,LGG,LIHC,LUSC,READ,SKCM | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000166526 | ZNF3 | 5 | BLCA,COAD,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000166526 | ZNF3 | 8 | COAD,ESCA,HNSC,LAML,LGG,LIHC,MESO,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000166526 | ZNF3 | 9 | BLCA,COAD,ESCA,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000166526 | ZNF3 | 6 | COAD,HNSC,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000166526 | ZNF3 | 7 | COAD,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000105708 | ZNF14 | ENSG00000166526 | ZNF3 | 5 | ACC,COAD,HNSC,LAML,READ | | ENSG00000225470 | ENSG00000135457 | TFCP2 | ENSG00000166526 | ZNF3 | 5 | COAD,HNSC,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000166526 | ZNF3 | 6 | COAD,HNSC,LAML,LGG,LIHC,READ | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000166526 | ZNF3 | 8 | BLCA,COAD,HNSC,LAML,LGG,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000149532 | CPSF7 | 6 | HNSC,KIRP,LIHC,PAAD,SARC,THCA | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000149532 | CPSF7 | 6 | HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000149532 | CPSF7 | 6 | HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000149532 | CPSF7 | 6 | HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000149532 | CPSF7 | 6 | HNSC,KIRC,KIRP,LIHC,PAAD,THCA | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000135870 | RC3H1 | 12 | ACC,BRCA,CHOL,COAD,DLBC,ESCA,KIRC,LAML,LIHC,LUAD,PCPG,SKCM | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000135870 | RC3H1 | 5 | COAD,LAML,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000135870 | RC3H1 | 10 | ACC,BRCA,CHOL,COAD,ESCA,LAML,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000135870 | RC3H1 | 6 | COAD,KIRC,LAML,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000135870 | RC3H1 | 5 | BRCA,KIRC,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000135870 | RC3H1 | 7 | COAD,ESCA,LAML,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000135870 | RC3H1 | 8 | ACC,COAD,DLBC,ESCA,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000135870 | RC3H1 | 5 | CHOL,COAD,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000135870 | RC3H1 | 6 | BLCA,KIRC,LUAD,PCPG,SKCM,UCEC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000135870 | RC3H1 | 6 | COAD,ESCA,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000135870 | RC3H1 | 6 | BRCA,CHOL,COAD,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000135870 | RC3H1 | 6 | BRCA,COAD,LIHC,LUAD,PCPG,SKCM | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000135870 | RC3H1 | 5 | BRCA,COAD,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000135870 | RC3H1 | 5 | COAD,ESCA,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000135870 | RC3H1 | 5 | COAD,KIRC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000135870 | RC3H1 | 6 | BRCA,COAD,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000135870 | RC3H1 | 9 | BLCA,BRCA,CHOL,COAD,KIRC,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198455 | ZXDB | ENSG00000135870 | RC3H1 | 6 | COAD,DLBC,ESCA,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000135870 | RC3H1 | 8 | BRCA,CHOL,COAD,LAML,LIHC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000135870 | RC3H1 | 5 | COAD,ESCA,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000135870 | RC3H1 | 6 | CHOL,COAD,ESCA,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000158711 | ELK4 | ENSG00000135870 | RC3H1 | 5 | COAD,DLBC,ESCA,LAML,LUAD | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000135870 | RC3H1 | 6 | BRCA,COAD,ESCA,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000135870 | RC3H1 | 5 | BRCA,KIRC,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000135870 | RC3H1 | 11 | BRCA,CHOL,COAD,DLBC,KIRC,LAML,LIHC,LUAD,PCPG,SKCM,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000135870 | RC3H1 | 6 | BRCA,CHOL,KIRC,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196233 | LCOR | ENSG00000135870 | RC3H1 | 5 | BRCA,COAD,KIRC,LIHC,LUAD | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000241058 | NSUN6 | 5 | COAD,KIRP,LAML,PAAD,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000241058 | NSUN6 | 6 | COAD,LUAD,PAAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000241058 | NSUN6 | 10 | BRCA,COAD,HNSC,LAML,LUAD,PAAD,PRAD,READ,SKCM,THCA | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000241058 | NSUN6 | 10 | BRCA,KIRC,LAML,LUAD,PAAD,PRAD,SKCM,THCA,THYM,UCEC | | ENSG00000225470 | ENSG00000117000 | RLF | ENSG00000167766 | ZNF83 | 5 | COAD,KICH,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000167766 | ZNF83 | 8 | BRCA,COAD,KICH,LIHC,LUAD,MESO,READ,SKCM | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000167766 | ZNF83 | 6 | KIRC,KIRP,LIHC,PCPG,PRAD,THCA | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000167766 | ZNF83 | 5 | COAD,KICH,LAML,LIHC,SKCM | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000167766 | ZNF83 | 6 | BRCA,KIRC,KIRP,LIHC,SKCM,THCA | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000167766 | ZNF83 | 9 | BRCA,COAD,LAML,LIHC,LUAD,PRAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000120784 | ZFP30 | ENSG00000167766 | ZNF83 | 5 | COAD,KICH,KIRP,LAML,READ | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000167766 | ZNF83 | 7 | BRCA,KICH,KIRC,KIRP,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000167766 | ZNF83 | 5 | COAD,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000167766 | ZNF83 | 5 | COAD,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000167766 | ZNF83 | 7 | KIRC,KIRP,LUAD,PCPG,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000167766 | ZNF83 | 5 | COAD,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000167766 | ZNF83 | 7 | BRCA,COAD,KIRP,LAML,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000167766 | ZNF83 | 6 | BRCA,KIRP,LAML,LIHC,SKCM,UCEC | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000167766 | ZNF83 | 5 | BRCA,COAD,KICH,LUAD,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000167766 | ZNF83 | 7 | BRCA,COAD,KIRP,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000167766 | ZNF83 | 13 | BRCA,COAD,KIRC,KIRP,LAML,LIHC,LUAD,PRAD,READ,SKCM,STAD,THCA,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000167766 | ZNF83 | 7 | BRCA,COAD,KICH,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000167766 | ZNF83 | 6 | COAD,KICH,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000105708 | ZNF14 | ENSG00000167766 | ZNF83 | 5 | COAD,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000167766 | ZNF83 | 11 | BRCA,COAD,KICH,LAML,LIHC,LUAD,PRAD,READ,SKCM,STAD,THCA | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000167766 | ZNF83 | 6 | BRCA,KIRC,KIRP,LAML,LUAD,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000167766 | ZNF83 | 14 | BRCA,COAD,KICH,KIRC,KIRP,LAML,LIHC,LUAD,PCPG,PRAD,READ,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000167766 | ZNF83 | 8 | BRCA,KIRC,LAML,LUAD,PRAD,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000004534 | RBM6 | 6 | LAML,LIHC,READ,SARC,SKCM,STAD | | ENSG00000225470 | ENSG00000065029 | ZNF76 | ENSG00000004534 | RBM6 | 8 | HNSC,KIRC,KIRP,LIHC,PAAD,PCPG,PRAD,THCA | | ENSG00000225470 | ENSG00000099326 | MZF1 | ENSG00000004534 | RBM6 | 9 | BRCA,HNSC,KIRC,KIRP,LIHC,PAAD,SKCM,THCA,UCS | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000004534 | RBM6 | 8 | BRCA,KIRC,KIRP,LAML,LUAD,PAAD,SARC,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000004534 | RBM6 | 7 | COAD,LAML,LIHC,LUAD,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000004534 | RBM6 | 11 | COAD,HNSC,KIRC,KIRP,LUAD,PAAD,PCPG,READ,SKCM,STAD,UCEC | | ENSG00000225470 | ENSG00000154957 | ZNF18 | ENSG00000004534 | RBM6 | 6 | HNSC,KIRC,PAAD,READ,THCA,UCEC | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000004534 | RBM6 | 6 | COAD,LAML,LIHC,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000166526 | ZNF3 | ENSG00000004534 | RBM6 | 6 | COAD,HNSC,LAML,LIHC,READ,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000004534 | RBM6 | 6 | COAD,KIRP,LUAD,PAAD,READ,SKCM | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000004534 | RBM6 | 14 | BRCA,COAD,KIRC,KIRP,LAML,LIHC,LUAD,PCPG,PRAD,READ,SKCM,STAD,THCA,UCEC | | ENSG00000225470 | ENSG00000168661 | ZNF30 | ENSG00000004534 | RBM6 | 5 | KIRP,LAML,LIHC,PAAD,SKCM | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000004534 | RBM6 | 6 | COAD,KIRC,LAML,LUAD,PAAD,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000004534 | RBM6 | 6 | COAD,HNSC,LAML,LIHC,LUAD,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000004534 | RBM6 | 15 | BRCA,COAD,HNSC,KIRC,KIRP,LAML,LIHC,LUAD,PAAD,PRAD,READ,SKCM,STAD,THCA,UCEC | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000004534 | RBM6 | 5 | HNSC,LIHC,READ,SKCM,STAD | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000004534 | RBM6 | 6 | COAD,HNSC,LAML,LIHC,PAAD,SKCM | | ENSG00000225470 | ENSG00000102878 | HSF4 | ENSG00000004534 | RBM6 | 5 | HNSC,PAAD,PCPG,THCA,UCEC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000004534 | RBM6 | 12 | BRCA,COAD,HNSC,LAML,LIHC,LUAD,PAAD,PRAD,READ,SKCM,STAD,THCA | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000004534 | RBM6 | 15 | BRCA,COAD,HNSC,KIRC,KIRP,LAML,LIHC,LUAD,PAAD,PCPG,PRAD,READ,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000004534 | RBM6 | 11 | BRCA,KIRC,LAML,LUAD,PAAD,PRAD,SARC,SKCM,THCA,UCEC,UCS | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,KIRP,LAML,SKCM | | ENSG00000225470 | ENSG00000036549 | ZZZ3 | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,LAML,READ,UCEC | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000131013 | PPIL4 | 6 | COAD,KICH,LAML,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000131013 | PPIL4 | 6 | COAD,LAML,PRAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000120784 | ZFP30 | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,KIRP,LAML,READ | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,LAML,SKCM,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000131013 | PPIL4 | 5 | KICH,KIRP,LAML,PRAD,SKCM | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000131013 | PPIL4 | 5 | COAD,LAML,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000131013 | PPIL4 | 6 | COAD,KIRP,LAML,PRAD,READ,SKCM | | ENSG00000225470 | ENSG00000186130 | ZBTB6 | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000131013 | PPIL4 | 5 | COAD,KIRP,LAML,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000131013 | PPIL4 | 5 | COAD,LAML,PRAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000131013 | PPIL4 | 6 | COAD,KICH,KIRP,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000131013 | PPIL4 | 6 | COAD,KICH,LAML,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000105708 | ZNF14 | ENSG00000131013 | PPIL4 | 5 | COAD,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000131013 | PPIL4 | 6 | COAD,KICH,LAML,PRAD,READ,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000131013 | PPIL4 | 6 | COAD,KICH,PRAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000147601 | TERF1 | 8 | COAD,HNSC,KICH,KIRP,LAML,READ,SARC,SKCM | | ENSG00000225470 | ENSG00000036549 | ZZZ3 | ENSG00000147601 | TERF1 | 5 | COAD,ESCA,KICH,LAML,READ | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000147601 | TERF1 | 6 | COAD,HNSC,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000125482 | TTF1 | ENSG00000147601 | TERF1 | 5 | ESCA,HNSC,KICH,LAML,READ | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000147601 | TERF1 | 6 | COAD,HNSC,LAML,READ,SARC,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000147601 | TERF1 | 5 | COAD,HNSC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000147789 | ZNF7 | ENSG00000147601 | TERF1 | 7 | COAD,ESCA,HNSC,KIRP,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000147601 | TERF1 | 5 | COAD,HNSC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000147601 | TERF1 | 5 | COAD,KIRP,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000147601 | TERF1 | 6 | COAD,ESCA,HNSC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000198455 | ZXDB | ENSG00000147601 | TERF1 | 5 | COAD,HNSC,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000147601 | TERF1 | 6 | COAD,HNSC,KICH,LAML,READ,SKCM | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000134698 | AGO4 | 5 | BLCA,COAD,HNSC,KIRC,LUAD | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000134698 | AGO4 | 5 | BLCA,COAD,HNSC,KIRC,LUAD | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000162599 | NFIA | 5 | BRCA,COAD,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000162599 | NFIA | 5 | BRCA,COAD,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000162599 | NFIA | 6 | BRCA,COAD,LUAD,SKCM,THCA,UCEC | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000149187 | CELF1 | 8 | BRCA,COAD,ESCA,KIRC,KIRP,LIHC,LUAD,SARC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000149187 | CELF1 | 5 | BRCA,COAD,ESCA,LIHC,LUAD | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000149187 | CELF1 | 6 | BRCA,COAD,KIRC,KIRP,LIHC,LUAD | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000149187 | CELF1 | 6 | BRCA,COAD,ESCA,KIRP,LIHC,LUAD | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000149187 | CELF1 | 7 | BRCA,COAD,ESCA,KIRC,KIRP,LIHC,LUAD | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000149187 | CELF1 | 7 | BRCA,COAD,ESCA,KIRC,KIRP,LIHC,LUAD | | ENSG00000225470 | ENSG00000196233 | LCOR | ENSG00000149187 | CELF1 | 5 | BRCA,COAD,KIRC,LIHC,LUAD | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000189079 | ARID2 | 9 | BRCA,COAD,KIRC,LAML,LGG,LIHC,LUAD,READ,SARC | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000189079 | ARID2 | 5 | COAD,LAML,LGG,LIHC,UCEC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000189079 | ARID2 | 7 | BRCA,COAD,LAML,LIHC,LUAD,READ,UCEC | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000189079 | ARID2 | 5 | COAD,KIRC,LAML,LUAD,UCEC | | ENSG00000225470 | ENSG00000142065 | ZFP14 | ENSG00000189079 | ARID2 | 5 | BRCA,KIRC,LAML,LUAD,SARC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000189079 | ARID2 | 8 | COAD,LAML,LGG,LIHC,LUAD,READ,SARC,UCEC | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000189079 | ARID2 | 6 | COAD,LAML,LGG,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000147124 | ZNF41 | ENSG00000189079 | ARID2 | 5 | COAD,LAML,LGG,LUAD,READ | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000189079 | ARID2 | 6 | COAD,LAML,LGG,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000167232 | ZNF91 | ENSG00000189079 | ARID2 | 5 | BRCA,COAD,LAML,LUAD,READ | | ENSG00000225470 | ENSG00000167766 | ZNF83 | ENSG00000189079 | ARID2 | 5 | BRCA,COAD,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000186272 | ZNF17 | ENSG00000189079 | ARID2 | 5 | COAD,KIRC,LAML,LUAD,UCEC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000189079 | ARID2 | 7 | BRCA,COAD,LAML,LGG,LIHC,LUAD,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000189079 | ARID2 | 8 | BRCA,COAD,KIRC,LGG,LIHC,LUAD,READ,UCEC | | ENSG00000225470 | ENSG00000025156 | HSF2 | ENSG00000189079 | ARID2 | 5 | COAD,LGG,LIHC,READ,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000189079 | ARID2 | 8 | BRCA,COAD,LAML,LGG,LIHC,LUAD,READ,UCEC | | ENSG00000225470 | ENSG00000115966 | ATF2 | ENSG00000189079 | ARID2 | 5 | COAD,LAML,LIHC,READ,SARC | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000189079 | ARID2 | 6 | BRCA,COAD,LGG,LIHC,LUAD,READ | | ENSG00000225470 | ENSG00000198521 | ZNF43 | ENSG00000189079 | ARID2 | 5 | BRCA,KIRC,LAML,LGG,LUAD | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000189079 | ARID2 | 8 | BRCA,COAD,KIRC,LGG,LIHC,LUAD,READ,UCEC | | ENSG00000225470 | ENSG00000256223 | ZNF10 | ENSG00000189079 | ARID2 | 6 | BRCA,KIRC,LGG,LUAD,SARC,UCEC | | ENSG00000225470 | ENSG00000196233 | LCOR | ENSG00000189079 | ARID2 | 5 | BRCA,COAD,KIRC,LIHC,LUAD | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000189308 | LIN54 | 5 | COAD,HNSC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000146757 | ZNF92 | ENSG00000189308 | LIN54 | 5 | COAD,HNSC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000164631 | ZNF12 | ENSG00000189308 | LIN54 | 5 | COAD,HNSC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000189308 | LIN54 | 5 | COAD,HNSC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000197857 | ZNF44 | ENSG00000189308 | LIN54 | 5 | COAD,HNSC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000189308 | LIN54 | 5 | COAD,HNSC,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000005889 | ZFX | ENSG00000178177 | LCORL | 5 | COAD,KICH,LUAD,READ,SKCM | | ENSG00000225470 | ENSG00000085274 | MYNN | ENSG00000178177 | LCORL | 5 | COAD,KICH,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000109381 | ELF2 | ENSG00000178177 | LCORL | 5 | COAD,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000126804 | ZBTB1 | ENSG00000178177 | LCORL | 5 | COAD,KICH,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000146587 | RBAK | ENSG00000178177 | LCORL | 5 | COAD,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000185129 | PURA | ENSG00000178177 | LCORL | 5 | COAD,KICH,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000196670 | ZFP62 | ENSG00000178177 | LCORL | 5 | COAD,KICH,LUAD,SKCM,UCEC | | ENSG00000225470 | ENSG00000198040 | ZNF84 | ENSG00000178177 | LCORL | 5 | COAD,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000062194 | GPBP1 | ENSG00000178177 | LCORL | 6 | COAD,KICH,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000095574 | IKZF5 | ENSG00000178177 | LCORL | 5 | COAD,LUAD,READ,SKCM,UCEC | | ENSG00000225470 | ENSG00000236287 | ZBED5 | ENSG00000178177 | LCORL | 6 | COAD,KICH,LUAD,READ,SKCM,UCEC |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
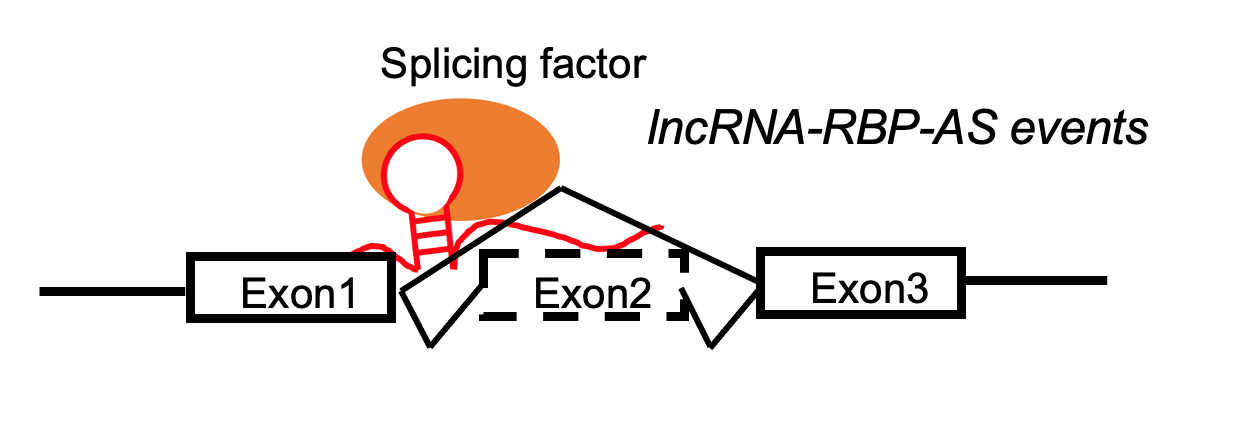
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type | | ENSG00000225470 | ENSG00000004534 | RBM6 | exon_skip_382458 | ENSG00000060971:chr3:38131595-38131638 | ACAA1 | ENST00000333167 | Frame-shift | STAD,UVM,SKCM,READ,PCPG,PAAD,THCA | | ENSG00000225470 | ENSG00000111364 | DDX55 | exon_skip_382458 | ENSG00000060971:chr3:38131595-38131638 | ACAA1 | ENST00000333167 | Frame-shift | STAD,UVM,SKCM,READ,PAAD,THCA | | ENSG00000225470 | ENSG00000139746 | RBM26 | exon_skip_382458 | ENSG00000060971:chr3:38131595-38131638 | ACAA1 | ENST00000333167 | Frame-shift | STAD,UVM,SKCM,READ,PAAD,THCA | | ENSG00000225470 | ENSG00000124193 | SRSF6 | exon_skip_382458 | ENSG00000060971:chr3:38131595-38131638 | ACAA1 | ENST00000333167 | Frame-shift | UVM,SKCM,READ,PAAD,THCA | | ENSG00000225470 | ENSG00000079134 | THOC1 | exon_skip_382458 | ENSG00000060971:chr3:38131595-38131638 | ACAA1 | ENST00000333167 | Frame-shift | STAD,UVM,SKCM,READ,THYM,PCPG,PAAD,THCA |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation |
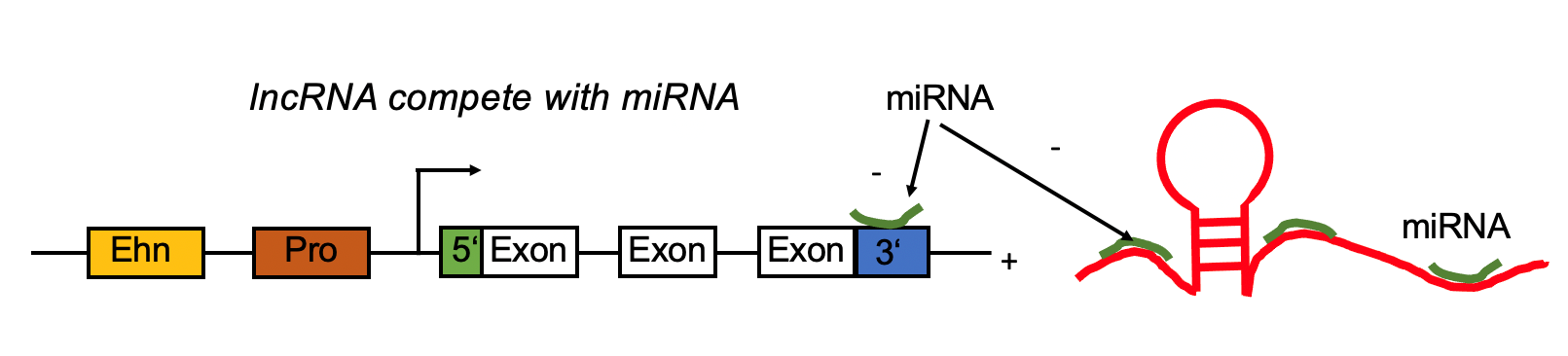
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000225470.5 | ENST00000602938.4 | 67 | 756 | 553 | | ENSG00000225470.5 | ENST00000602294.4 | 73 | 493 | 272 | | ENSG00000225470.5 | ENST00000602920.4 | 87 | 318 | 55 | | ENSG00000225470.5 | ENST00000602737.4 | 87 | 318 | 55 | | ENSG00000225470.5 | ENST00000602772.4 | 94 | 409 | 125 | | ENSG00000225470.5 | ENST00000414209.4 | 103 | 1209 | 898 | | ENSG00000225470.5 | ENST00000602895.4 | 111 | 1330 | 995 | | ENSG00000225470.5 | ENST00000453317.1 | 74 | 176 | 400 | | ENSG00000225470.5 | ENST00000602546.4 | 51 | 474 | 319 | | ENSG00000225470.5 | ENST00000602985.4 | 73 | 420 | 199 | | ENSG00000225470.5 | ENST00000415215.1 | 62 | 110 | 298 | | ENSG00000225470.5 | ENST00000538519.3 | 79 | 251 | 12 |
|

 Gene summary
Gene summary AS events and RNA A-to-I editing events of lncRNA in TCGA based on Genvode V22 structure.
AS events and RNA A-to-I editing events of lncRNA in TCGA based on Genvode V22 structure.
 Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).