lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | EEF1AKMT3,ARHGEF17,RPAP2,FERMT2,PMM2,VAMP1,TINF2,MTMR6,AMER1,MFSD8,ATL2,IFT22,WDFY1,MAST3,ELOVL5,LIG3,LIFR,TTC31,MAPK8IP3,MID1,VAMP4,FKBP7,SLF2,GNAQ,TMEM184C,HLTF,PIGG,SENP1,GPATCH11,RNF6,TOB2,HMGCS1,TFIP11,RETSAT,TULP3,WDR5B,ST7L,FAM200A,LYRM2,TRIM5,FAM133B,IBTK,TBK1,TTC14,CCNK,BRPF1,SH3GLB1,AFAP1,PPP1R3E,PPIP5K2,MAP3K5,SPTY2D1,PRLR,XPO7,C20orf194,ZFP1,ZIK1,NCBP1,PPIL4,FAM219B,RGP1,GCNT4,TRIM68,AP3M2,SMC2,PSIP1,RIC8B,UBE3A,TMEM30A,THOC2,ITGA7,ACAD10,MFSD1,TMEM231,ZBTB6,EDEM1,OTULIN,C3orf18,GTF3C2,ERG,OFD1,ATP9A,PEAK1,SEPTIN10,FRK,FCHSD1,UBE2H,MSH6,PURA,EPDR1,CD164,MIS12,HGS,ANO8,ZNHIT6,HERPUD2,CTDSP2,SEC63,NEMP1,CBX7,EML4,MIS18BP1,HEG1,MDN1,KLF10,TSPAN2,PTPRB,DIPK1A,BANK1,PAIP1,FAM53C,RAB5A,SRPK1,SEC23A,FKBP15,KCTD2,MBD4,LPCAT3,SECISBP2L,ORC2,SSBP2,VGLL4,PHC3,AIDA,ETFDH,ZZEF1,DHX35,INTS2,VPS26C,TFEC,SUPT20H,UBXN2A,CBX6,SLU7,ERGIC2,FAM83B,SMARCAD1,GIGYF1,KHNYN,ZNF14,CCDC69,MBTPS1,DZIP1,CDK17,IPO9,ATG2B,NIPAL3,USP38,HUWE1,B3GALNT1,AP4E1,DDX42,WDR35,TIFA,MKRN2,SEC24C,ACACA,CRYBG3,DNAJC24,PGGT1B,MAF,RALBP1,CYFIP2,MECOM,GCA,RCHY1,SELENOI,PITPNM2,TRIM14,SWT1,BRD3,TMEM64,PLPP1,RASSF6,RWDD4,ARHGEF9,AMMECR1L,TMEM168,IPO8,SLC26A11,FANCL,NCOA4,BRAP,KLF7,PHKA2,CPT1A,WBP4,UBE2G2,PAWR,MIB1,CEP68,OSMR,NEMP2,KLHL15,RAP2C,VEZF1,EPB41L4A,MLLT6,PRPF40A,ELAPOR2,GSK3B,ROBO1,ATG2A,ZNF70,MOB3B,INO80D,MYO10,ICE2,SMCR8,MAGI2,AGK,ENOPH1,TMED4,CRLF3,DDX24,TCAIM,PHF7,KPNA1,TESK2,STK3,MAPK9,DNAJB4,ZXDA,DNAJC21,ZDHHC7,TNFAIP3,RETREG1,RIOX2,SCO1,TRNT1,PPP2CB,CHD8,KIAA1143,DENND6A,RALGPS2,UBR4,TMED8,IFT74,RELL1,OSBPL8,NPR2,CBLB,XPO4,AP1AR,S1PR1,RASSF9,UFL1,OCRL,PKD1,RIMKLB,TAF8,LCP2,NUDT12,SLC25A46,PPM1L,SEMA4D,DCTN1,IGSF3,UTP14C,ARNT2,BBS2,RFWD3,TBC1D9,CEP83,DDX20,CPLANE2,NCOR2,LIN54,PLPBP,BMP2K,LRIG2,FBXO8,SEC23IP,ABCB7,SNAP23,OSBP,HIP1,LMOD1,CHRNB1,IKBKB,DTWD2,SLC38A1,BTC,BPNT2,DGKE,SRBD1,ANKRD49,CYB561D1,PAXBP1,IFNLR1,CCDC125,TMOD2,C6orf120,KCTD18,ACACB,PLEKHA8,RNF20,RSRP1,CPNE8,EYA3,ERO1B,RNF31,GOLGA4,FLCN,SMIM14,NVL,ATG14,GOLGB1,FOXP1,PWWP2A,DNAJC18,MOB4,MYNN,TSPYL5,RPE,SLC39A10,TMCO3,LINS1,CACNA1H,SNX5,LDAH,ANKEF1,SCAF11,CCDC25,EPHA4,ADAM9,PUS10,TMEM167A,CNOT7,DIABLO,DAB2,CCNG1,ZNF862,PDS5B,EDEM3,ALAD,SEC24A,DCTN4,SNX6,UEVLD,LDB2,KIF3A,GGT7,OAS3,TYW1,SCAMP1,ANKMY2,MAN1A1,CSTF2,EXOC7,NCK1,DISP1,MIDEAS,EPB41L2,TPCN1,ABLIM1,UNC119B,PIK3CA,LMLN,ATAD2B,SAR1B,ALG11,NMNAT3,PREPL,PI4KA,ESCO1,MTMR10,PBX2,FGD6,KLF12,CCDC85C,ANGEL2,ATPSCKMT,NIPAL2,NAA15,POC5,MYSM1,ITPRID2,METTL15,TTC23,LACTB2,UIMC1,ELF1,C12orf29,UVRAG,RNF44,SMARCE1,RBBP8,TNS1,CEP95,TMTC1,TTI1,KIAA0895,KLHL13,HELZ2,DDX52,SLC11A2,USP4,TASP1,IFT88,CYP1B1,ATF6B,NDNF,STARD7,CRY1,MEF2C,GTF3C3,AFTPH,ZMAT3,DHFR2,XPC,PRR14L,FAM160A1,FYTTD1,TOM1L2,BARD1,ERLIN2,RPS6KB1,SLC25A35,PTK2,SH3BGRL,N6AMT1,KANSL3,ITGA1,CDON,RGL1,SH2B3,DENND4A,INIP,CCDC32,DDX60,STAT3,MPHOSPH9,POM121C,HECTD1,TGFBR3,INTS7,SP100,EDC4,TMEM209,AGO2,ARL13B,RBM26,SVIL,CAMK2G,SPATA6,TMEM19,EFNB2,SYNE1,MLH3,KANSL1L,KIF21A,ATP8A1,ENTPD4,ENPP5,SMC5,GNA12,HEATR5B,ZNF106,LNX1,VPS53,SNW1,LCOR,PABIR2,CENPC,XRRA1,HERC2,OTUD6B,TDG,IQCK,WBP11,ABHD13,TUBGCP4,RCAN2,RUBCN,AOC3,PARN,PRRC2A,MEGF8,C2orf49,POLK,PLEKHA3,DCAF12,ABCF2,WRN,PRDM2,SPATA5,LCA5,VEZT,BAZ1B,USP12,MOB3C,TMEM87A,METTL6,AAK1,DSG2,THSD1,MOSMO,BAG4,ENOX2,FCF1,SP110,SPCS3,TAPBPL,SASH1,ZNF8,APP,AMMECR1,SUSD1,BMPR2,SNRNP200,SALL1,ERCC3,POLG,PLSCR4,FAM13B,ST8SIA4,UBA7,DYM,BRCA1,ARFGEF1,MTMR9,SPOPL,RAPGEF5,WDR6,DSE,BRD8,GNS,ERCC8,UBE2V1,GSPT1,ITGA6,USP16,MAPKAPK5,ZMYND8,TMEM47,SLC9A7,XYLT1,IARS1,BROX,IP6K1,ENTPD7,FBXL14,APPL1,RBM33,PAG1,NOL8,ADAL,MYLK,RAB11FIP1,COL4A5,PRMT5,FPGT,DSTYK,COL4A4,KLHL26,MPHOSPH10,LEPROT,SAMD4B,PLEKHA7,SCFD1,POLR1B,PDZD8,EXOC5,CAMK2D,CLEC16A,ERICH1,CSDE1,RFX5,ZNF2,GXYLT1,LNPK,POLR1F,ADGRF5,PLA2G12A,KANK1,ADCY9,MAP3K2,CSNK1A1,RPRD1A,GPATCH2,RPTOR,TOR1AIP1,LAMA5,NHLRC2,CAAP1,TTC28,ZFYVE26,VTI1B,MS4A7,BECN1,ANKRD42,LARP7,NIPSNAP3B,KMT2E,CWC22,AKT2,C2orf42,SIAE,MORC3,ITPR2,CPXM2,FAS,PEX1,CHORDC1,S100PBP,TRIM3,SLC30A1,KLHL18,AGPAT5,KAT6B,FBXW8,SUPT16H,PDSS2,CHTF8,RNF216,TRIM23,TSC2,DTWD1,TECPR2,SLC25A30,PCNT,N4BP1,YEATS2,USP54,RB1,TRAF3,IKZF4,TEFM,TCEANC2,NF1,LTBP2,C2orf69,SPIN4,ANKRD50,THAP9,ACTR3B,CDC42EP3,THAP6,FBXL5,DHX15,YWHAG,LSAMP,SEC62,GNB5,KIF20B,TRAM2,KLHL12,AQR,SENP7,RAB27A,GALNT11,STK24,PTPN21,TAF2,SEC16A,GOLGA3,TRPM7,PODXL,PXK,GTF2E1,MCM8,ACLY,TRAPPC8,TSPAN12,KCTD20,ARSK,ERV3-1,MKLN1,WDR41,GAB3,PHIP,TMED7,DOCK9,CLMN,LRRC58,DUS4L,PPP1R21,CRIM1,RBM18,JAK1,CCDC93,DLAT,LMBR1,SNX1,PATL1,GANAB,SIPA1L3,TM2D1,UBAP2,FAM98A,RASSF8,HP1BP3,GIMAP6,VPS13C,NLRC3,CBR4,ITPR1,WASHC5,USO1,DIDO1,CYREN,LTBP1,FKTN,ALG9,CEP41,SLC36A4,EMCN,PUM2,DHX36,MKRN1,BCDIN3D,IMPA1,TTLL3,TUBE1,RCSD1,YAP1,ARMC1,NEDD1,STOM,MEOX2,C2CD2,ARHGEF6,KATNIP,RCBTB2,RNF185,SPG11,WASL,FBXO42,LIMA1,CLIP1,PDCD7,NCAPD3,ITFG1,INTS4,F13A1,PIP5K1A,MAPK1,STK38,FRMD4B,TSPYL4,ELMO2,KRCC1,MGAT4A,PUM1,USP22,SLC12A2,NUP155,SPIRE1,RB1CC1,IL6ST,CILK1,FASTKD3,ZRANB1,DYNC2LI1,PDE5A,CAPZA1,ILDR1,DCP1A,CITED2,TANK,DMTF1,MKNK1,ZDHHC23,CCDC47,RFC1,FAM193A,NSD3,DCK,MAP4,MAK16,JADE3,USP36,SNIP1,SF3B3,VPS39,STAT2,PLEKHM1,GMFB,WWC2,KAT6A,AFF4,LZTFL1,ARAP2,STRBP,RNF38,LBR,NMNAT1,KPNB1,DEK,ZBED5,HUS1,INPP5B,SETD2,SIN3A,UFSP2,CREB1,MEX3C,SUDS3,MAP2K5,DPY19L4,CAPN7,ADAM10,CNN3,KIF16B,SMAD2,CLCN6,ADD3,IGIP,RFX2,PDE7A,SKAP2,VPS35L,MYEF2,RNF19A,RAPGEF6,FIG4,SEMA3G,ASH1L,CLIC2,POLG2,NFIA,PTAR1,CEP152,MGRN1,KATNAL1,CPSF7,JAK3,ZNFX1,FLI1,TRIM37,PARP4,CTC1,AGBL5,CCDC149,ARMC10,INTS6,FBXO38,TBCCD1,IFT172,SYPL1,HCFC2,ADAMTS9,DUSP3,SLC4A1AP,PRKACB,ARSG,RSRC2,SNX27,GLG1,NAPB,USP46,NAA25,ERMP1,PNMA8A,RHPN2,PRMT9,SLC30A4,ARL15,PRKX,TM9SF1,SLC35A3,CEP104,DOCK10,ITPKB,ZBED8,CRK,CHD9,HACD2,RALGAPA2,SVEP1,LMAN1,BACH1,PIGN,IQCG,HPS5,AUTS2,SBNO1,SYTL2,F2R,POGLUT1,TECPR1,CMKLR1,POGLUT3,MLLT3,TYW3,ABHD16A,TOP1,ARHGEF40,NXF1,RNF111,C6orf62,PAN2,LRTOMT,MOAP1,STAG2,PKN2,ARSD,SLC30A6,SLC12A4,SNX4,STYX,METTL4,FITM2,DCLRE1A,GRK3,TAGAP,MTMR4,HIPK2,ATXN2L,SDE2,ANKRD13C,RINT1,AHDC1,LAMB2,GOLGA2,INSR,STXBP5,ATF7IP,TMEM248,PIP4K2A,SHOC2,MEF2D,NARS2,CES2,ETV1,MOSPD2,RSPH3,HERC4,ATG4C,CD2AP,CASTOR3,RAB28,HSPG2,TMEM254,TRMO,PDCD10,MBOAT1,LUZP1,DENND5A,COPB1,DPY19L3,SMG7,MED7,TNFAIP1,DCLRE1B,LCORL,CSNK1G1,CEP57,LRRC8C,ENPP4,METAP1,ARIH2,SAMD9,LNX2,AZI2,HAUS2,AMD1,NEXN,RTL6,EIF2AK3,KIAA1191,BRCC3,SMAP1,DDX55,EIF4G2,WLS,SPART,SHQ1,RASSF2,GBP3,OTUD7B,UBA6,ACAP2,ASAH1,SWAP70,IFNGR1,ADD1,SESN3,TNKS2,LANCL1,ODR4,IFT140,NDC1,OXR1,AP5M1,OPA1,BCOR,NCKAP1L,DENND11,AMIGO1,LYSMD3,TRAF1,PRKCQ,APBB2,FNBP1,MSH3,NEK9,EP400,UACA,NATD1,ANKDD1A,GLT8D1,DDX6,TRMT2B,NOMO2,C17orf75,PGM2,SESTD1,LCMT2,EVC,TMED10,SLX4,SP2,MECP2,NBPF11,ATP6V1C1,RPRD1B,SEC31A,QRSL1,AASS,ACIN1,TMEM182,TNRC18,THAP1,CD3G,TTC8,TAOK2,USP6NL,MBLAC2,ITK,IGFBP5,GOLT1B,GTF2A1,CEP85,ZNF28,NUBPL,BOD1L1,FAXDC2,RAB33B,LTBP4,CHD2,GCLC,RASGRF2,PEX13,NBPF15,METTL14,ERN1,OTUD4,AHNAK,RABL3,PRKD1,TMPO,QKI,HERC1,TRIM22,POLR1A,STAT6,BCL2L11,TNFRSF10A,TRIM4,ZSWIM8,HERC3,GDAP2,RAD50,TXNDC5,PITPNA,LAPTM4A,PIGK,SBF2,LPP,LRBA,SLC31A1,ELP1,TXNRD3,RRN3,STARD8,MGAT2,TTC12,SYNGAP1,XAF1,CHD1,ZHX2,FBXO30,HECTD4,GTPBP8,ZEB2,HARS2,GNPTAB,EP400P1,HPS4,MCPH1,NBPF1,USP45,UBE3B,CARD8,TAF6,GOPC,EVI5,NCOA2,LRIF1,MED17,DNMT1,HECTD2,BCL6,PLCG1,RCBTB1,PRKAB2,RALGAPA1,LATS2,TMX3,ZBTB4,KATNAL2,EVC2,MDM1,ANK3,CA5B,MED13L,FCHO2,POLH,SMG6,OLFML1,NEK4,OSTM1,SPIN3,TBRG1,RUNX1,ITSN1,FAT1,PPP1R12A,POFUT1,HEYL,RNPC3,C14orf28,IDE,TRPS1,MNT,PCDH18,DOP1A,BTBD19,KIAA2026,SYNPO2,KMT2C,CCSAP,STAU2,SCAF4,YTHDF3,CCNT2,SNX14,RRM1,ORC3,HIF1A,PBXIP1,CLMP,NRAS,SAV1,RNF123,ZNF91,ZFP91,TMCO4,TRAPPC11,ARMCX3,OPHN1,ATG16L1,MARCHF1,ATP13A3,RBM25,PLD1,KLHL9,SMARCC1,TBC1D4,TTC30A,UBR5,RESF1,TRAM1L1,ETNK1,SERTAD2,LRIG3,ARL6IP5,CAVIN2,E2F6,PSKH1,ZC3H13,PANK3,SAP30L,NRP1,ZNF717,HEATR6,SLC15A4,HEXIM2,MAP3K7,AAGAB,NEPRO,LIG4,SRPK2,ANKRD27,IRF2,MFSD11,USP37,GASK1B,NMT1,USP19,PNRC1,SGSM2,TTC7A,SMAD3,VMAC,TGS1,ARHGEF26,TMEM161B,KRR1,SDCCAG8,EPM2A,WBP1L,VAMP3,HLCS,TBC1D2B,TPP1,FAM149B1,FBXO32,DENND1B,AMBRA1,ADAM17,PHLPP2,UBL3,PPP4R3A,ABI3BP,TMTC3,RFX3,UBA3,TMEM245,STAT1,CRCP,TMEM68,CAD,VWA8,C3orf38,IDH3A,USP48,FASTKD5,CYLD,TNS3,WWP2,ZCCHC10,PARP8,HELZ,MINDY1,SEL1L,LRRK1,ELF2,NDUFA5,SRRD,TMEM65,ZNF43,PTPRK,CDC14B,PDE3A,CACUL1,GLT8D2,ARL5B,TNS2,PMS2,RECQL,LZIC,TMX4,TPP2,PDCD4,FZD6,HDAC4,NR2C1,AP1S2,ABHD17B,ADAT1,SOS1,PARP16,AVL9,MED1,ZDHHC21,SEC22C,GGA3,CEP112,DNAAF2,ATG5,AP3M1,CPEB2,PPP2R1B,TRIM66,KIF5B,KIAA0100,TNPO1,NID2,MSL2,C3,SLFN5,POT1,KLHL2,STAG1,EIF5,TMEM267,COPG2,ITFG2,RNF4,KAT7,GNA13,YIPF5,AASDH,GLI3,RABGAP1L,LHFPL6,TRANK1,MAML2,SLC25A43,PHYKPL,SF3B1,POFUT2,ANKIB1,MXRA7,ZFP69,SETD7,LDLRAD4,PIGM,ATP11B,GOLGA1,PLAA,MINDY2,CUL7,MTPN,USP24,SOS2,IMMP2L,POGZ,BRWD3,AKAP10,RBMX,WDSUB1,MAP3K12,ORC4,FUBP1,NMD3,KLF11,TADA2B,SMPD4,TRIM33,NID1,NBAS,ACVR1,KIDINS220,DCAF1,DDX17,DGLUCY,C12orf66,MAP3K13,UBTD2,FMR1,SPRING1,CHMP2B,MYD88,ZDHHC2,MBP,DHX33,BBS12,EPB41,CTDSPL2,GNPDA2,SDR39U1,LIN52,SMC6,CDAN1,EXOC6B,GPALPP1,RHEB,UBFD1,WDR31,SORBS1,DLG1,ABCC5,AP3S2,ANO6,ARHGAP1,CERT1,CRYZL1,SLC22A5,LRP12,STK36,EPS8,GOLIM4,GRB2,TM9SF3,SS18,ALPK1,ZFX,VPS35,RMND5A,CBWD2,RABGAP1,STEEP1,PER3,SEPTIN7,SH3BP5,CPEB3,CEP162,DST,TRIP12,RBMS1,ZFP28,ZNF17,SRCAP,CRBN,RNF169,POLD3,ANTXR2,ZNF84,CCP110,BICRAL,WDR26,HACD4,HOOK1,SLFN11,BSCL2,ZDHHC6,PRPF40B,PCMTD1,ADAP2,ARHGAP21,TMEM35B,NGLY1,VPS37A,ASB8,KBTBD2,SLC16A14,USP13,IFIT5,ATF2,TRMT10A,C6orf89,REPS1,SEC22B,PEX2,COPS2,HELQ,CPLANE1,ERI2,LARS1,SLC25A32,ABCB1,OXNAD1,CSNK1G3,SH3D19,CCR5,ATP6V1A,BRD1,SPATA13,ACTN1,TRIQK,RBM7,IPP,NDST1,MFAP1,PPID,PPP1R3B,CHM,MMRN2,SUGP2,SLC16A4,TTC39B,ARL2BP,SEMA3C,PHF10,RAB6A,NUP50,FBXL17,UBE4A,GDAP1,ZBED4,WDCP,ANKH,PLRG1,SLC25A40,ZFP14,MYO1D,EGLN1,SNX12,TUBGCP6,SMARCC2,KCTD10,MYO9A,ACAD8,ITGB1,ALDH5A1,DENND1C,TVP23B,GNG10,PBX3,SGCB,ACTR6,UBA2,ATP8B1,KLHL42,KNTC1,RANBP6,SAMD12,SMIM15,SLC33A1,EXOC8,C19orf44,ATR,NIPA1,SARAF,STRN3,TMEM106A,ARCN1,HSPA13,SCYL3,OGT,CCDC146,GZF1,TNIK,BLZF1,CNST,FBXO5,CLOCK,TUT4,DNAJC16,RBM15,ANAPC10,MOCS3,BBIP1,SEPTIN6,BACE1,CLK1,ATXN7,AKT1,IGF1R,MCMBP,ST13,RFX7,ACADM,SLC17A5,MAP2K4,CD84,VTI1A,SUN2,MBD5,RAF1,KPNA4,PRKAR2A,RNF11,ZADH2,MAP3K14,GON4L,TRAM1,CDC27,CERS6,LPCAT2,CASK,BIN2,NARS1,WWTR1,CHIC1,ZNF45,YTHDC1,COQ7,RHOBTB1,CADPS2,TMEM185B,DCTN5,PPP6C,TFCP2,GTF2H1,MLH1,SMC1A,CCDC191,TAF5L,TNKS,NR2C2,SHE,SIX5,FBLN5,BMF,PPP4R2,TIMM10B,TAF1A,BICD1,TENT5C,RMI1,SCRN3,CYRIA,LRP10,FAM114A2,PLS1,PFKM,ZMPSTE24,ARHGAP29,NCAPG2,PIK3C2B,GLS,MAN2A1,MBTD1,SEPSECS,IFT122,FBXO21,MAN1A2,MANEA,RAB3IP,CEP192,ARHGEF11,ADGRL4,KAT14,VPS13A,NACC2,EPB41L3,SF3A1,ECPAS,DACH1,ZFP3,ARHGEF37,JKAMP,SH3PXD2A,LONRF1,RNF214,MAPK1IP1L,SDHD,HIBADH,ENTPD5,PIK3R4,FAM172A,THADA,SNX33,ILF3,GFPT1,CAND1,LENG8,PER2,SMURF2,FBXL3,NICN1,INPPL1,NEK11,FGD4,PRICKLE2,FAM168B,PPWD1,RC3H2,DARS2,TNKS1BP1,SLC30A7,SASS6,CLASP1,NHLRC3,ASTE1,ADGRA3,COG8,MNS1,CASP2,KRIT1,DCAF7,COG5,SERINC1,PROSER1,MAML3,VPS50,SDC3,WAPL,ZZZ3,UGGT2,ZMYM5,C2CD3,NOTCH4,CSNK1D,PARP12,PDCD6IP,CROT,MBNL3,SETX,TOP3A,TMEM116,TRIM13,ARHGEF38,WIPI1,LRP1,CD93,GRPEL2,OSBPL1A,PHC1,LRRFIP2,RAI1,SACS,MTUS1,PABIR1,MTRR,ZCCHC9,PUS7L,SLC39A9,SLC25A24,MEIS1,FZD5,DCTD,NUDCD3,CTBS,CD47,GGNBP2,ERCC6L2,RTF1,PPP1R13B,CABLES2,RASEF,FBXO3,ATP11C,ZBTB2,NEMF,RTTN,REV3L,IWS1,AFDN,PEX26,ZFYVE1,AGTPBP1,CEP70,MAU2,ENOSF1,DZIP3,SIDT1,PXMP4,GANC,WDR7,COA1,FNBP4,TNXB,PPM1K,TOX4,TMF1,SSH2,MLLT10,BHLHB9,PEX11A,DENND5B,TSTD2,AHR,MED23,UNC50,CLEC2D,PDLIM3,BAG5,MTIF2,SPTAN1,ZNF720,NCKAP1,MIER3,SYTL3,ILK,GIMAP8,BTBD7,MED29,PDHX,GPBP1,SCARF1,ARHGAP32,LSG1,EHMT1,TTBK2,PSD3,CASKIN2,ZFAND6,TP53BP1,CDK19,TMLHE,ZFHX3,ARID4A,ADPRH,PIGF,SMAD4,AASDHPPT,TTC37,VCPIP1,CCDC80,WASF2,TENT5A,TGOLN2,PPP3CA,CLIP4,PLPP3,RBBP9,RHOBTB3,WDR27,MPDZ,FAM104A,SPAG1,SFXN3,BBS7,BSDC1,MATN2,TFDP2,MFAP4,PNISR,TMEM63A,SEPTIN8,SMARCA1,ARHGAP42,GALC,ZNF10,PCYOX1,FBXO9,FAM126B,ABCB10,FAM234B,NPNT,AFG3L2,CEP120,CDC42BPA,B3GALNT2,OSBPL9,TGFBRAP1,ROCK1,WASHC2C,SHANK2,INTS12,IQCE,FAM214A,N4BP2,CEP250,ARID2,NUP214,STAM2,TTC5,MALT1,SPARCL1,PLEKHH1,PICALM,DBR1,SEMA5A,LHFPL2,NFKB1,HEATR1,WDR3,ZFAND4,PBRM1,TK2,RUNDC1,URB2,TTYH2,PHKB,TCAF1,TENT4A,PPP4R3B,OBI1,NECAP1,CPNE3,USP10,MTMR12,CXCL12,CCDC50,PHF8,SSH1,KLF6,COX18,FYCO1,ZFAND3,EEA1,LPIN1,PHKA1,UBE2W,PHTF2,CHD6,BMPR1A,CLDND1,DFFA,PCSK7,C5orf22,PTPN3,KIF27,FBXW7,GLE1,SLC7A6OS,ZFP30,SPRTN,POLI,AGFG1,PRPF39,ITSN2,DLC1,ZFAND1,TRIM26,SRRM2,PJA2,PIK3IP1,PARP3,IDS,ACOX1,SLC4A7,CMTR2,ADGRA2,GORAB,UBR3,COL14A1,ANGPT1,CEP44,DBT,PROSER3,SLIT3,PECAM1,GMCL1,PTPN23,SLC37A3,KPNA3,KRAS,SLC9A9,ANKRA2,SOGA1,CELF1,FAM126A,CDC40,CCDC121,EPG5,PEX3,CANX,PMS1,QRICH1,FAM160A2,ERAP1,CYP7B1,ZNF83,FAM161B,STAU1,EDA2R,DPH1,UHRF1BP1L,CDS2,ASPH,RUSF1,C4orf36,COPB2,RBBP5,SZT2,USP32,MYO18A,PRKDC,FECH,GIN1,MARK2,FASTKD2,STK4,ARHGAP31,BCL2L13,URGCP,XRN2,CCAR1,SACM1L,TMEM131,AREL1,MTREX,ORAI2,FBXO36,SLC10A7,EAF1,KLHL8,NANP,MFAP3,GTF2H3,TLN1,IFIH1,YPEL5,STX2,WBP1,ERMAP,CDYL,HCFC1,RAD17,PDCL,LRCH4,NUP133,USP51,PELI2,CDC42BPB,BET1,RYBP,CS,PROM1,MOB1A,ADAMTS10,RBM39,HMGCR,GPATCH8,F8,PGS1,ARMCX6,MDC1,VSTM4,TEK,C7orf31,GNG12,FAM120B,SCRN1,CKAP5,FBN1,BLOC1S6,EIF3C,INTS3,TWSG1,PDS5A,AZIN1,AP4S1,PHLDB1,EFCAB14,MPP7,ACBD5,ATRX,FEZ2,ABCA2,RBM28,C2CD5,ENPP2,GLIS3,ZBTB5,PTEN,FLNA,BCLAF1,GAB2,PIK3CB,RASAL2,PAN3,DCHS1,FOXO1,MOXD1,RNF168,RRP1B,DDX59,ISCA1,ZC3H4,NUMB,SHTN1,BCAP29,RGS5,ITPRIPL2,TIMP2,C1GALT1C1,CALCOCO1,ANKRD52,METTL25,DNAJB9,DPP8,APAF1,MMGT1,WDR55,TOGARAM1,FAM217B,B4GALT4,LPGAT1,FAM76B,IRS1,LRRFIP1,PDIA4,EOGT,NOTCH1,H2AZ2,RERG,ADNP,CAVIN1,ESYT2,FBXO33,MBD1,LMO7,ARID1B,ARRDC3,ERC1,GID4,SPPL2A,PRDM4,TCF20,ITIH5,SPTLC2,FLII,MRNIP,POLR2B,ABL2,RSBN1L,RAD52,CEP43,URB1,M6PR,PDE4D,STRN,MARF1,INO80,CEP290,NEK1,CERS2,RAD21,WSB1,TRAK2,CPSF2,ITGAL,RSPRY1,ALS2,NSD2,TRIM52,NCBP3,ATL3,DEPDC5,SALL2,BAHCC1,BRPF3,CUL4A,MTX3,SAP130,SPATA20,YOD1,MCM3AP,STRADA,NPTN,NAF1,CDC14A,TM9SF2,RANBP9,TJP1,LMBRD2,KANK2,ARNT,CREBBP,DOCK4,CHURC1,FSTL1,TFRC,HTT,GOLPH3L,TRIM35,CHML,FAM3C,VPS11,DNAJC11,SFMBT1,FYB1,IKZF2,C19orf71,TRMT61B,COLEC12,SDAD1,CAB39L,CCPG1,ARHGEF12,ADCY3,TUBD1,UTP15,LONP2,CDCA7L,FRYL,KIF13A,USP31,NGRN,DENND4B,NSUN3,TEP1,NOTCH2,MAN2C1,TAB3,SUCLA2,TBC1D5,SFMBT2,WHAMM,LIN7C,TET2,RABL2B,CFAP97,RAB23,ADH5,TSPYL2,SPAG9,GART,SAMHD1,RNF141,CMTM4,XRN1,TTF2,GLYR1,BRD7,AP3B1,ANAPC1,WDR89,TAB2,SCFD2,DIAPH1,DHX38,CCDC88A,RPS6KA3,N4BP2L2,RAB30,FBXL4,TXNDC16,L3HYPDH,RNF34,UNKL,NEDD4,WWP1,STIM1,GPCPD1,ZNF24,ASCC3,SIDT2,SIPA1L1,CELSR1,KMT2B,IRAK4,GSTCD,NBN,SERINC5,RHOT1,TP53INP1,RAB19,KLHL28,KDM5B,NOP9,RSF1,ABITRAM,THAP2,VIRMA,PIAS3,CLK4,SUMF2,GIMAP7,COMMD2,AMN1,ZUP1,USP30,ARL1,RPAP1,RNF40,PPP1R3D,GPRIN3,GPRASP2,KDM5A,SDHC,SERAC1,SLC40A1,FBXO34,BAHD1,ARHGAP19,ANKS1A,PJA1,NOD1,ARIH1,LRIG1,DIPK2B,CARNMT1,PPTC7,ARID4B,YTHDC2,FEM1C,MEIS2,DDB1,FAN1,TOP2B,RPAP3,SELENOP,IKBKG,DHX58,UBLCP1,HS2ST1,RBBP4,TMEM39A,CEP19,FOXJ3,TBC1D9B,SLC35A5,BMI1,BNIP3L,S1PR3,MIER1,NCOA6,TTC21B,NPAT,FANCF,AP1G1,ANKRD11,RRM2B,NBR1,ARID1A,RAB3GAP2,PRKAA2,UTP23,SETD5,ZFP62,FOXF1,SOCS4,CNOT8,BRAF,PLEKHA6,ACSL4,FRMD4A,BAZ1A,NKAPD1,OSBPL3,PTPN14,GUF1,SLC35F5,CUL9,PRRC1,LAMA4,DPYD,SSR1,PTPN13,CHRDL1,CAMSAP1,LRRC37B,DHX29,RSRC1,TAOK3,TMBIM6,ANKRD28,FIGNL1,DCN,FUT10,TMEM127,R3HDM1,KCTD12,TLR1,SMURF1,BRD2,LSM11,DPYSL2,COG7,CCDC186,BTBD9,OPN3,ZFC3H1,MIOS,CLCN3,NFYB,SP3,DNAJC3,ARHGEF7,CHSY1,HIPK1,TMEM50B,PIP4P2,SIK3,GALK2,PIK3C2A,PAFAH1B1,ICMT,CUL1,SVIP,PALB2,TXNIP,EIF2AK2,ABCE1,TSR1,FMNL3,SLF1,ITGB8,MARCHF6,RFFL,CUL3,LTN1,PRDM5,PLCH1,ATM,UBN1,SNX19,UBC,MED14,ARMCX2,NUP54,SYTL4,IQGAP1,CDK13,PRUNE2,NOA1,TRAF6,ZC3H10,BBOF1,FEM1B,CABIN1,SLFN12,COBL,RELCH,RNF14,RSBN1,ATG12,NR1D2,PDZRN3,PIBF1,AHI1,NCSTN,LAMTOR3,RORA,CNOT6L,KDM5C,ATP8B2,GPC6,PGAP1,MOB1B,USP53,FAM111A,PARP11,ARHGAP18,GCFC2,AKNA,UBR1,PHF12,AFF1,RPA1,ACTR8,RAPGEF1,KIF13B,PDP2,SNRNP48,RC3H1,UBE2J1,NBPF3,GALNT7,ZNF44,SLC12A6,SMG1,TAF7,MTURN,B4GALT5,KANSL2,XIAP,ELP4,BCLAF3,CD34,TMEM43,SLC35B4,MFSD9,LPAR1,DNAJB14,CHUK,PPM1B,MTOR,MCM9,IFT80,GMPS,PHTF1,ABRAXAS1,PELI1,ZCCHC14,ZGRF1,MYO5C,ATP11A,CDK5RAP2,NFIC,RMND5B,UBN2,FMN1,KIAA0586,RBM47,ABHD18,SLC5A3,SMCHD1,FILIP1L,PRR12,POLR2A,CREBRF,FOXN3,ABHD15,CIITA,ANGEL1,SMYD4,KLF3,CYP4V2,PRKD3,ZNF41,ZC3H8,METTL16,LATS1,ARHGAP17,SLC35D1,AKT3,TRAPPC6B,WDR82,RAB2B,KNOP1,SLC38A9,PGBD1,APPL2,EPS15,HEATR5A,SGTB,JADE2,TGDS,DCAF6,KMT2D,SYNRG,CYB5R4,FRMD6,PDGFD,PNPO,USP40,G2E3,TRAPPC13,GPR180,PTPN22,MCFD2,TRIM44,TUBGCP3,PIGO,CBLL1,ZFP36L1,ARHGAP6,KIAA0319L,TMEM218,ARMH3,BCL9L,ANKRD26,PSMD5,GSAP,GGA2,BTBD10,DOCK8,MYH9,NADK,C18orf32,MRE11,RBM6,TRMT13,AKTIP,ARHGAP12,COBLL1,NT5C2,SORL1,AGPS,ARPP19,VPS54,CCDC142,LYRM7,PAXIP1,FLT1,SYNE2,CBX5,MYH11,KIAA1549,RYK,EPM2AIP1,NAA35,TMEM144,FAM168A,MAML1,GTF3C4,MPPE1,ANKFY1,BORCS5,DIXDC1,TPR,PIGB,NOC3L,CEPT1,DDX23,DMXL2,NUCKS1,FAM13A,CUL4B,DET1,CLCN7,TBC1D15,ESYT1,TNPO3,INVS,EPRS1,EXO5,BCL2,SESN1,CEP63,MYO9B,EIF3A,GRSF1,STARD13,WDR11,TEX2,FKBP5,EXOC2,KDM3B,ADPGK,RBM23,CEP57L1,AKAP8,UBTF,SEC24B,VPS13B,FOSL2,TAF4,WDTC1,TBP,GABPA,SLX4IP,MANBA,DPY19L1,IRF2BPL,RNF145,GATAD1,TET3,TTC30B,BNIP2,PDPK1,PPM1A,INCA1,ATMIN,TOB1,GFM2,UBXN7,LRCH3,CNOT4,SON,DHX57,ALG12,RLIM,TYK2,ZMIZ1,CHST14,TDP1,MYO5B,PTGFRN,PTCD3,MAGI1,EFEMP1,NDE1,NFRKB,TMEM192,RAD1,BRWD1,CNOT1,NIN,DDI2,CHD3,BORCS7,RBMS3,ABCC4,NAPEPLD,PDK2,RO60,TBCK,SLC4A2,ACBD3,SEPTIN2,BBS10,ARHGEF3,SP4,CSGALNACT1,RBM43,MBIP,NOTCH3,ARL5A,CDC23,HDHD2,KIAA0753,FAM117A,COMMD10,OSBPL11,NDOR1,SGSH,VPS41,SETD3,TBL2,ZHX3,DDHD1,MAP4K5,SBDS,PHAX,TMEM128,DLG5,CWC27,CIAO1,RNGTT,FBXW11,KDR,C1RL,CCNL1,TLK1,USP28,TMEM263,ASB7,UBE2G1,ZDHHC20,NCOR1,SCMH1,VAV3,MADD,RBM45,C5orf15,NFXL1,RPS6KA6,NFIB,RNFT1,DDX31,PBLD,ADCY6,DCUN1D1,NUP98,SPOP,WDR36,PRUNE1,GAS7,RANBP10,DDX58,CLEC7A,TMUB2,RMDN1,RPS6KC1,KANSL1,ZNF3,TBC1D20,ZMYM3,DNMT3A,HAUS6,SYNJ1,TBC1D8,LRRK2,CLINT1,GOLPH3,PCYT1A,FAM161A,JCAD,KDM2B,C12orf4,DDX5,G3BP2,STX17,SEC24D,PPP2R5E,C8orf37,LIPT1,CWF19L2,KITLG,DENND2B,LYST,EMC1,PNN,MICAL3,PDGFC,LUC7L2,IRF2BP2,GAPVD1,ARHGAP5,BBS4,CASP8,SNX13,ALKBH1,IQSEC1,KMT2A,REV1,SH2D3C,NEURL4,PREX1,KDM1B,NRIP2,RAB5B,FAM135A,SEC14L1,ETV3,LAMP2,CAMKK2,CEBPZ,HACE1,GLB1,GABPB2,KLF9,PIGA,ATXN7L1,TMEM167B,MAPKBP1,C21orf91,TAF9B,TRIM25,ATN1,TBL1XR1,MYOF,NRF1,VASH1,SCP2,WASHC4,ZXDC,PTGIS,NSRP1,SPTLC1,FNDC3A,RARB,IL17RA,PIGL,ZC3H7A,BAZ2A,GEMIN5,RPS6KA2,SSPN,DCAF8,BRI3BP,ABCA5,PAK2,NKIRAS1,SNX29,MAP3K4,LIPA,CEP350,CXorf38,PTCD2,BCL2L2,ATP2B1,SUPT7L,PSME4,EXTL2,SNX16,ARL6,CTSO,OSGEPL1,CHPF2,SLC41A2,RAB11FIP2,ITM2A,VAMP2,USP7,EXOG,SYT11,KCTD9,NCOA5,SCAI,MYO19,PAFAH2,DCAF5,ANKRD40,MAFB,PRIMPOL,ABHD2,GCN1,COPA,UBQLN1,TENT4B,TIRAP,RAB22A,WDR20,SCAPER,LRP6,ELMOD3,DYNC2I1,ITGA2,LIAS,ASXL2,EIF2AK1,GCC1,PNPLA8,MAP4K3,DUSP16,CARMIL1,TC2N,PFAS,VPS52,ELK4,CNNM3,TWF1,ASAP1,USF3,LMBRD1,FBXO11,LPIN2,BBS9,ZMAT1,IRAG1,CHL1,CSGALNACT2,ZW10,CMTM6,RERE,SGPP1,TEX10,APPBP2,POC1B,CNOT2,SHROOM4,ABCA1,C1orf109,EDNRB,RBM48,GEM,PIKFYVE,LGR4,RBM5,RAB3GAP1,ULK2,NKTR,PLA2R1,IPO5,LEPR,MTF1,FARP2,SLC30A9,CEP170,SLC35B3,ETS1,SCYL2,WDR33,PTPRA,NRIP1,SLAMF6,C17orf80,TRA2B,PIAS2,TSC1,DMD,TMEM185A,DCAF11,LRRC40,USPL1,CCDC112,PRPF4B,LACC1 | | Exist in public source | | NA |
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | TRIM33,ENDOD1,KDELC2,PRUNE,MKRN2,FRMD4B,CASC4,SCPEP1,HELZ2,ABCA5,CD93,COX18,UTP20,ANKRD40,DNAJB4,IST1,UBA7,BRD7,EP300,PLEKHH2,FECH,HOOK1,AP5M1,ATG4C,CAAP1,TCAF1,TRIM5,DHX33,WDR31,MEF2D,EVI5,SLC40A1,VAMP2,C2CD5,RAPGEF1,IL10RA,CANX,TMCO4,PIGK,GRAMD3,XRRA1,JAZF1,EXOC5,WNK1,IFT81,ATP2A2,FAM63A,PIK3CG,KCTD3,MED1,PIGV,FAM126A,TERF1,TJP1,PMPCB,LIG4,PCYT1A,CCDC88A,TNKS2,GK5,ZDHHC2,SSPN,XAF1,ATL3,ENTPD1,KIAA1671,TUBGCP4,ROR1,APP,HACD4,ARL13B,SRSF8,SMPD4,NPEPPS,IRF2BPL,SLK,LCA5,ERAP1,MMRN1,PEX11A,SS18,MBNL1,AVL9,DIEXF,NMNAT3,ZNF14,TPCN1,WIPF2,SHROOM3,GGCX,ITPRIPL2,AGO4,FAM133B,INPP4A,BBS7,RIMKLB,SSH2,ZC3H11A,MFN1,SUN2,PIGF,PIK3R4,C17orf75,COG6,XIAP,AKAP12,TNFRSF10A,PHLDB1,RASSF8,WSB1,MRPL42,ARMC1,ICE2,WDR35,LPIN2,TOR1AIP1,CSDE1,TRIM25,PATL1,JAK1,LDB2,PCBD2,PRRC2B,NFIA,ADAMTS9,DDX1,HACD3,RAB23,AMBRA1,ACLY,ZMAT1,ARHGEF10,PIK3CB,ALPK1,GORAB,TESK2,MYO1D,CERS6,SLC39A10,RABL3,PGAP1,STOM,KANSL1L,IQCE,RCHY1,MYOF,ZNF83,KPNA6,EIF4G2,SUPT7L,SPPL2A,ERMAP,SEC31A,PCMTD1,YOD1,TUBGCP3,TMEM131,LIPA,B4GALT5,TOPBP1,YEATS2,EYA3,ANKRD26,NFX1,N4BP2,ADAM10,MAVS,ITPR3,ELAC1,TAF1A,WAPL,NEU3,UFSP2,LRRC40,MFSD8,TGFBRAP1,UBR5,STK36,LACTB2,ASAH1,ELMO2,SLIT3,RBMS2,SH3BP5,TUBD1,CPSF7,WDR43,SENP5,SLX4,RBM6,ABI3BP,SLC9A6,ABHD16A,CC2D2A,SORBS2,ZYG11B,CD3G,ATXN1L,FAM220A,BRCC3,RAB6A,FAM73A,IFIH1,IPO8,MORC4,PDIA4,EPRS,STK38,UHRF2,ST5,TLR3,FBXO33,CEP290,FAM114A2,H6PD,NCBP3,YME1L1,PAK2,BCL9,EXTL2,PMS2,ENOSF1,SMURF2,PTGER4,PTPN12,PTPRB,USP28,CLASP1,MIS18BP1,VTA1,COPA,TEP1,OSBPL3,SLU7,CNOT7,AKAP1,CDC23,NAA35,SCARA3,OSBPL11,EAF1,TBRG1,BRPF3,CHD8,SH3BGRL,PRKAA1,TTC30B,FOXJ2,GRAMD1C,UBTF,GNA13,CLTC,NUP98,LRP1,SYDE1,MECP2,SMARCA5,MYLK,RANBP6,FAM120C,TIFA,ANKRD6,PTPRF,KIF16B,RNF219,GTF3C2,HAUS6,USP32,CTTNBP2NL,DDX52,UBXN7,GCLC,NUS1,MAFK,NLK,CTSO,RHOBTB3,MCCC2,MAPK8IP3,TBC1D9B,CCDC66,RASAL2,ERCC4,STAT6,MYO5B,SKIV2L2,EXD2,KLHL8,FYB,SPICE1,PDE4D,DNAJC16,TMED5,DHX15,SLC38A9,CDK13,BRD3,MED21,TINF2,CFLAR,SRP72,CARD8,ERCC3,CTDSP2,UGGT2,NUDT12,SGSH,KIDINS220,RP2,SPTLC1,MYLIP,KANK2,BRWD1,REST,HMGXB3,GRB2,ABCB10,TTC23,KIAA0513,MAK16,GALNT11,EP400NL,SLC12A4,IMPA1,B3GALNT1,CDS1,MCTP2,ATP2C1,COA1,RUFY1,INSR,GIMAP8,EXOC6B,MINA,SLC4A7,PLEKHA5,ELMSAN1,SEC23IP,ARMCX5,SDR39U1,ACBD3,PFAS,PAPLN,MLLT10,TLK1,FNDC3B,UBE2J1,SETX,KIAA0319L,FBLN5,THAP6,UBA2,GTF2I,WWP2,ZDHHC4,CEPT1,MTIF2,SLC35A3,TNXB,HSPG2,ARL5A,USP42,ERC1,KDM5A,PAFAH1B1,CTR9,SEPP1,SETD7,KANSL2,INCENP,JAK2,UBC,P2RY10,TWSG1,GCN1,MSL2,RYK,NUMB,ARHGAP25,SH3D19,PJA2,PPP1R3B,MAT2B,DDX60,EXOC4,THSD4,CADPS2,INO80D,PHACTR4,SERINC5,NIPA2,USP6NL,RAB11FIP1,CSNK1A1,TSC22D2,CCDC121,SGTB,TRMT10A,NEXN,NOTCH3,BAZ1B,PTPRG,NMNAT1,LRCH3,ITPKB,VPS37A,RLIM,PRMT6,LACC1,ZMAT3,PLXDC2,KIAA0556,KIAA0586,DNAAF2,ERLIN2,SMARCC2,LEMD3,OSGEPL1,SETD2,ACACA,DDX58,ZFYVE9,SLC33A1,MTMR12,CNST,NXPE3,PPIL4,SRRM2,RNF168,GNG10,MED12,SENP2,SPATA25,UHRF1BP1L,HHAT,TCTN2,CRYZL1,PHKA1,RIF1,INSIG2,CHSY1,JAM2,MET,EIF2AK4,METTL21B,SRSF5,PLEKHA2,USP13,TLR4,PUS10,CEP162,SUZ12,NDUFA5,COQ7,LZIC,PDLIM5,PRLR,TAGAP,RAB27A,UTRN,EML1,NUPL2,PPIL6,MR1,PTPRK,CMPK1,C18orf25,FLII,ENTPD5,SLFN5,E2F5,ARHGAP19,GIGYF2,NOC3L,ZC3H12C,ABCB1,BAZ2A,DGCR8,LMBRD2,SMARCA2,FAN1,ORC3,ATP11B,CASP8AP2,ANAPC10,ZFP14,RINT1,DIP2B,TMEM30A,TIGD6,SNX27,LDOC1L,PRPF40B,CDK17,STAT3,PIK3IP1,CYFIP2,SLC30A7,SNX4,FOCAD,SLC36A4,HERC4,PLAA,KAT5,PTPN9,LTBP2,CNTNAP1,TLK2,PLEKHH1,GPATCH2L,MIOS,TM9SF4,MAML3,GPATCH11,MAN2B2,EPM2A,SRPK2,IKBKG,RBMS3,ARGLU1,SEC22B,FLI1,DCAF8,HNF4G,OGDH,SOCS5,GGA3,TJP2,NANP,LPCAT3,XPO4,DENND6A,C6orf89,FAM173B,PITPNA,CTC1,PRKCQ,ANGEL1,FER,NCK1,HERC2,CHD9,HSPA4L,IL17RA,DHX29,GPRIN3,SLC4A1AP,ICMT,FAM185A,NAPEPLD,DOCK8,SESN3,CPSF2,STAU1,HERC6,SAMD9L,ATG2A,E2F6,LARS,IFT74,PDLIM3,NBPF3,MPHOSPH9,PDGFD,ZFHX3,BDP1,USP33,ANGEL2,PRRG1,LIMD1,ITPR2,MLXIP,ELF1,CEP120,ARHGEF9,ZC3H4,CARD6,CLIC4,PDS5A,BCAS3,PAN3,DHX9,PI4KA,TMEM123,MDM4,GON4L,SUPT20H,SEC24B,WDR60,SMAD1,SLC23A2,SIDT1,SLAMF6,CRISPLD1,CNNM4,FRS2,BORCS7,MSN,PPM1A,SCLT1,FKBP9,ZNF720,TAF2,RNFT1,LDAH,GUCY1A3,DYNC1LI2,USP4,RSPRY1,BOC,ARHGAP24,TRIM14,ZC3H6,LCORL,C2orf42,FLCN,CDYL,SIN3B,RBBP8,POLD3,HEATR6,SLC35F5,S1PR1,ADGRL2,HLF,RFWD3,MERTK,ABHD18,PDP2,SPTY2D1,RAD17,PARP9,SLC25A24,PDZRN3,KIF27,CPQ,CCNL1,S100PBP,DDX31,SLC25A35,TMEM248,FKTN,N6AMT1,ZFP3,CASK,ZCCHC11,CDAN1,PEAK1,NEK4,DCAF7,ZCCHC10,SF3A1,SLC12A6,ZNF3,CCDC125,NBAS,BCL2,PPID,ITCH,ZNF70,RSBN1L,RIN2,RORA,SPRY1,LIN54,ERG,TBC1D4,LRIG3,SACM1L,KDSR,IFNAR2,ZUFSP,KDM1B,ITSN1,FOXO3,CDKAL1,SP110,MPHOSPH10,ITGA4,NEK7,CCDC82,MPDZ,TCP11L1,PAPD5,SCFD2,GEM,MAU2,DNMT1,SIKE1,ACBD5,PIK3C2B,DPH1,NRAS,MAML2,MAPK1IP1L,LMTK2,ZZEF1,TFCP2,LCP2,FASTKD5,HECA,MED28,NMD3,WDR91,FAM175B,VRK2,IPP,CCNT1,VPRBP,POC5,PGS1,DTX3L,IQCG,USP36,EFHC1,GATS,MAP2K5,FOSL2,CDK5RAP2,FITM2,POC1B,KIAA0355,DCTN1,LIMS1,AGO2,WDR55,PPP1R12B,TANGO6,ATF1,STAT2,FUBP1,TNFAIP3,ANKRD13C,TMEM19,FBXL7,TET2,CALCOCO2,UHRF1BP1,VKORC1L1,TMEM263,FAM69A,SSH1,LRRC57,BTBD7,CEP78,STRADA,MFAP3,LYST,FBXO42,PRMT9,MOB1B,EXO5,ARAP2,FAM111A,SLC41A2,YWHAB,SNIP1,GANC,PDP1,BTN2A2,PCNT,IARS2,CCSAP,CBFA2T2,SIN3A,DIRC2,SPRED2,CD34,METTL14,METTL6,ARSD,COMMD2,TRIM23,INTS12,GEMIN5,ZMIZ2,RERG,HAUS2,ACOX1,WDR48,CRKL,MBLAC2,ADAT1,JMJD1C,SPG20,KIAA1551,ACTR3B,TRIM69,USP9X,AGO1,FZD5,KIF2A,DBR1,FAM122B,TK2,SEPSECS,CTDSPL2,DDX46,ZFP30,PDCL,GFM2,SNX1,SLC26A11,ABHD13,HEATR5B,PCM1,ZBED6,UEVLD,MAN2C1,BCL2L13,SH3BGRL2,CEP135,ILDR1,KRCC1,PRRX1,PDE3A,COG7,DFFA,PLEKHG1,MAP3K1,CABIN1,ATAD5,ZBTB6,MCM3AP,KCTD2,TASP1,NCAPD3,TYW1,ZDHHC7,KLHL28,RAD52,CAB39,SMG1,NCOA6,CDK12,LONP2,RPAP1,TBP,DDX17,TNS1,ATP8A1,MFSD11,C19orf44,MOB1A,RAP2C,RAB3IP,BACH1,KLHL7,PPHLN1,NR2C2,PIK3C3,C12orf66,FAM179B,MCMBP,GOLPH3,LAP3,ALDH5A1,ZNF45,TRAPPC10,PPP2R1B,GIMAP6,ZEB2,ARFGEF2,AREL1,RALGAPA1,PALB2,SBDS,CNOT6,KDM4A,CDON,WDR20,TOM1L2,H2AFV,TMEM50B,RB1,CD109,PHF14,RHPN2,SYNPO2,ST8SIA4,MAST3,GCA,COIL,PEAR1,VPS13C,ORC2,SDAD1,KIF3B,BNIP2,CBL,SYTL2,RNF185,NICN1,PDXDC1,RAB3GAP1,KPNA5,MYNN,THAP9,C12orf49,TMEM87A,UPRT,PDE5A,BCL6,ANKRD52,PPTC7,PLEKHM1,DNAJC18,MLEC,ROBO1,TRMO,RALGPS1,NARS,GLG1,MOAP1,LAMA2,CAD,GIT2,RERE,NID2,PHTF1,SGPL1,C10orf76,TRIM22,IPO5,TMX3,CLPX,PSMD5,SPIN4,TSPAN2,NUDT9,GPC6 | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
| Only predicted by TDF | | APITD1,PSMB3,CUEDC2,MED16,DGCR14,KARS,FKBPL,PIH1D1,PRDX5,HINT2,JTB,ARPC5L,HCFC1R1,MAP2K2,GADD45GIP1,CAPNS1,C16orf91,G6PC3,VPS72,CLPP,MRPL52,ECHS1,NCLN,RBM42,CLTB,ADRM1,ATPIF1,BABAM1,RPS23,THOP1,ECSIT,NFKBIB,LONP1,SVBP,DTYMK,MCRS1,TIMM17B,BANF1,METTL1,ECI1,ANKRD39,PA2G4,POLRMT,PSMB5,APOE,AURKB,HM13,BCAT2,PSMB1,NDUFA1,SSR2,CCDC94,PSMA2,YIF1B,CCDC137,NDUFS5,RNF25,RRP7A,SGTA,TFPT,DAPK3,MRPL47,PDCD2L,CENPB,ATP6V1F,PEX16,CCDC167,PGLS,FAM110A,CENPM,MZT2B,COX6A1,TOMM22,PTTG1,TMEM134,ENO1,C14orf2,NAA10,RPL35,ACTL10,THAP4,HMOX2,PSMB6,AIP,THAP7,EMC8,CDC34,PRDX4,INO80C,FAM127B,THOC6,SCNM1,COA6,VPS25,WDR13,TIMM9,MZT2A,SGF29,ATIC,TK1,MRPL4,ENDOG,GPX4,DYNLRB1,GTF3C6,SNRPC,ATRAID,PRR7,SHFM1,VPS28,COX17,COX14,PTDSS2,COX8A,RNF126,MTCH2,UBE2T,RRP12,FAM195B,AVEN,C20orf24,UBA52,SH3GL1,CKS1B,TPRA1,PRR14,MFSD12,RABEPK,KIAA2013,PQBP1,MIIP,TIMM10,CORO1B,RTKN,LAGE3,MACROD1,UQCC2,SLC25A11,PMVK,NDUFA2,YDJC,TBL3,COMTD1,RABL6,RFC2,PFDN4,GNB2,CSTB,TMEM9,LIMD2,SPC24,C11orf24,GPAA1,ICT1,GFER,NDUFB1,VAMP8,COX7B,TMEM161A,LSM2,S100A10,UBALD2,ABHD12,NENF,SAT2,C8orf76,ZMYND19,COPS6,DGAT1,TXNL4A,CYC1,NSMCE1,THAP3,LAMTOR4,B9D2,PCYT2,HSCB,CWC15,NUDT1,SF3B4,GUK1,OXLD1,PMF1-BGLAP,PPM1G,LSM10,EBPL,ALDOA,DOHH,C7orf50,RPSA,NDUFA3,PSMA4,CITED4,SYNGR2,NAA38,TRMT112,USMG5,PES1,PMF1,RNF113A,SART1,EIF3G,TWF2,EIF3F,NDUFB11,NT5C,FAM234A,POLR2J,FKBP8,RPP25,UBXN1,NOSIP,UBXN6,NANS,PSMD4,NDUFAB1,PSMA5,C12orf45,SLC25A10,NMRAL1,SNU13,RAB25,BAX,PARK7,PSMB4,PPP2R1A,STOML2,ITPA,HDGF,OCEL1,NTMT1,SURF1,TSTA3,NDUFA8,CNPY3,PHB2,NNMT,GSTP1,ELOF1,ATP5D,TMEM160,MRPL24,AURKAIP1,PRR13,H2AFX,RPL23A,PPP1CA | | Exist in public source | | NA |
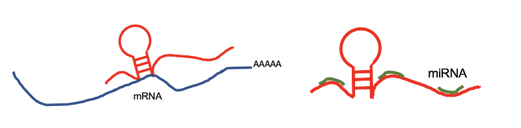
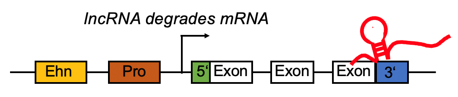
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | TECR,SH3GL1,DAPK3,ECH1,RNF181,SF3A2,CTDNEP1,PNKD,PAXX,PSMG1,FTL,PCYT2,IMP4,MRPL40,NACA,ZNHIT1,GSTP1,CIB1,EIPR1,ANAPC11,GNB1L,GALE,PIN1,FPGS,RPS19BP1,RASSF7,AUP1,RPL35A,METTL5,TPT1,ASPSCR1,UBL5,VAMP8,MVD,C4orf48,MIIP,MRPL43,MYG1,CNPY3,STX10,SLIRP,TSR3,LIMD2,POP5,LRFN4,PSMA4,MRPL48,GCAT,DHRS4L2,COPE,SCYL1,MRPS12,POLR2L,PPP1CA,GMPPA,SELENOO,NDUFB9,ADAT3,HSCB,PSMA7,MAD1L1,UBALD2,PSENEN,MRPL41,RNASEH2C,COMMD1,ATG101,SDF2L1,EMC8,HDGF,PPDPF,NDUFA12,EEF1B2,CHMP1A,NDUFV1,MRPL28,HMOX2,NDUFC1,MRPL37,RABGGTA,WDR74,PPIB,NMRAL1,LYPLA2,HRAS,FBXW5,GSTO1,RPL32,STUB1,PTMS,CDT1,ZBTB8OS,MZT2B,TXNDC17,EEF1E1,NDUFAF8,QPCTL,CLN6,RPSA,NDUFS6,MRPL4,SPAG7,NOSIP,SAC3D1,IPO4,NAA10,NHP2,SEC13,DBI,TBL3,FLYWCH2,HIKESHI,TRPT1,MRPL20,RRP1,PGLS,SRM,ATP5PF,FKBP1A,PCGF1,PEX16,NR1H2,FAM207A,SWI5,NDUFS7,TMEM258,NT5DC2,PQBP1,UBE2F,NFKBIL1,PSMB1,ARMC6,C19orf53,HDGFL2,COX7A2,UROS,TRAPPC2L,SLC39A3,PSMB6,PFDN6,NDUFC2,MRPL34,LAMTOR1,INTS11,PSMB3,BRI3,RPS29,RANGRF,MFSD12,FKBP2,NDUFB8,GFUS,PEMT,KXD1,DNPH1,GEMIN7,TMEM147,PRDX2,RPL37A,DPP7,REX1BD,PRDX4,FAAP20,RPL36A,ENDOG,PWWP2B,UXT,MRPL51,RPS24,AURKAIP1,RPL26L1,RFXANK,C6orf136,ALDOA,ATP5PO,BLOC1S1,SLC25A11,UROD,NAA38,MAD2L2,TALDO1,SIRT6,ARPC4,SLC25A39,FAM234A,SF3B5,RUVBL2,YPEL3,REEP4,POLR2H,LONP1,TMUB1,SERF2,NOP2,ATF4,TIMM17B,MRPS21,ATP5ME,PA2G4,MBD3,COX4I1,C12orf57,SDHAF2,PHPT1,ISY1,DCXR,RPL35,EIF2B3,KRTCAP2,QARS1,H2AJ,CORO1B,CLTB,DGAT1,NDUFA11,SLC3A2,LSR,ZDHHC16,NCAPH2,TMEM208,CNPY2,C12orf45,PKN1,RABEPK,FKBP8,ABHD17A,RHOG,JPT1,RPS17,UBXN1,RPS11,TK1,C6orf226,PPIF,ATP5PD,CYC1,POLR1C,CCDC124,PPP1R7,BOLA3,NDUFB1,NFKBIE,GPAA1,TKT,FUOM,LAGE3,MRPL22,GPX1,RNF25,MPST,RBM42,SART1,C11orf24,WDR18,TMEM222,CDC123,COA8,RPS8,CUEDC2,GGCT,NDUFB4,EBPL,RPS21,MRPL47,GNL2,ZNHIT2,MRTO4,GUK1,NDUFAF2,PSMD4,NDUFA9,DTYMK,TIMM9,IDH3G,SURF2,RABAC1,AP2S1,RPP21,PUF60,BSG,NUDT14,PREB,UQCC3,RAC3,DOHH,MACROD1,OAZ1,NANS,STX8,TRMT112,ADCK5,TUBB4B,FBXL6,HINT2,NAT14,C7orf50,ARRDC1,BRMS1,MFSD10,GMFG,SNRPA,COX8A,OTUB1,NTMT1,COX5A,PPP1R35,EIF3K,AGPAT2,TAF10,RPS27A,LSM7,EPN1,SAT2,MRPS15,ATP5MF,PRR13,TMEM161A,DUSP23,SLC52A2,NFKBIB,GLRX3,METTL26,FDX2,BID,NDUFA6,RNF208,SEM1,SHARPIN,SLC66A2,VPS72,LYRM4,IDH3B,CDC37,METTL1,LAMTOR2,NUDT8,FBXL15,MLST8,PEX14,C8orf76,GSTZ1,CHID1,PRAF2,CDC34,XAB2,PHB2,VPS28,CCT7,MPDU1,PTOV1,BABAM1,SCNM1,CTU2,NDUFS5,PPIH,CHMP2A,NME1,PTTG1,RPL13A,VASP,FAM50A,AVEN,POLRMT,RPS26,THAP7,SSR2,EIF1,SLC66A1,PSMD13,MRPL57,TFPT,HSD11B1L,RAB4B,THAP3,RFC2,YIF1A,GPX4,USE1,EMG1,FAM174C,STXBP2,RABL6,MRPS34,ATP5MC2,ALG3,HAX1,ELOC,MICOS13,PFDN5,MZT2A,WDR83OS,WDR13,LCMT1,SIVA1,MRPL55,ROMO1,CYBA,NDUFB10,C8orf82,NELFE,ECE2,PKP3,GAPDH,NUDC,PARK7,COMTD1,ENSA,SNF8,NNMT,MVB12A,RAB25,DYNLL1,PSMC5,MYL6B,RFNG,TIMM13,RNH1,COX17,PLEKHJ1,ZFAND2B,HSD17B10,EIF6,PDCD5,AKR1A1,LMNA,CSNK2B,KIAA2013,MPND,COX6C,SGF29,ATP5F1EP2,MRPL54,PDCD2L,PPP2R1A,CUTA,JOSD2,ATP5MJ,STOML2,NDUFB11,DHCR7,POLR1H,VKORC1,FARSA,NDUFV2,TSSC4,KLHDC3,G6PC3,IRF3,PRR7,RPS6,NOP16,BANF1,CARS2,TPGS1,ORMDL2,NT5C,OXLD1,RPP38,NTHL1,POLR2I,MRPL52,PSMG3,RHOD,RHOC,RPL10,UQCC2,WDR46,MFSD3,RPL27,PSMB7,RPS6KB2,ATIC,RNF113A,COPS6,UBA52,RACK1,TARBP2,ALKBH7,MPV17L2,TXNL4A,TBC1D7,TMEM219,NDUFB7,PPIA,BUD31,PHKG2,ANKRD13D,CCS,GNB2,NDUFAF3,FKBPL,PRMT1,MSRB1,PUSL1,TP53I13,RPS10,MPG,MCRIP2,HAGH,STARD10,NDUFA13,CD320,CHCHD6,ATRAID,MIX23,APOE,SMIM26,THOC6,PET100,TMEM9,SMPD2,CORO7,INO80C,SURF1,COX5B,COX6A1,DDX49,CHCHD10,SELENOW,PAFAH1B3,SRA1,THAP4,TMEM160,DUS1L,BCAT2,ABHD14A,DRG1,UBXN6,ANTKMT,SIL1,ARHGDIA,POLR2J,TRABD,GAMT,RPL36,SCAND1,DDA1,ZNRD2,AAMDC,COX6B1,NDUFA7,FLAD1,UQCRC1,MYDGF,PSMC3,BOP1,GPS1,DDX39A,CHCHD5,NAPRT,THOP1,BAD,PMF1,APRT,ECSIT,C19orf48,PPP4C,SELENOH,IFT43,COX14,TEAD4,NARF,TEDC1,RECQL4,GTF3C6,ATP5MK,EIF3F,PHLDA3,DPM3,MED16,GALK1,SSBP4,GIPC1,EIF3I,C11orf98,UQCR11,MRPS9,HMBS,MCRIP1,DYNC2I2,SPC24,ERGIC3,UQCRQ,TXN2,TMEM256,BIRC5,PSME2,TMEM205,MRPL24,COX7B,NAA60,PRDX5,EIF3G,RPP40,RPL23A,B3GAT3,HSPE1,PMF1-BGLAP,ECI1,UBALD1,ENKD1,AURKB,HSF1,NDUFS3,SSBP1,PFDN2,COA4,PIH1D1,SLC39A4,SSR4,SCAMP3,LAMTOR4,NAXE,SIGIRR,LMAN2,CIAO2B,UBE2J2,GFER,C9orf16,MAF1,RBIS,ATP5MC1,CFL1,HOMER3,DCTN3,PELP1,PYCR3,NOC4L,MIF,CCDC85B,UBE2S,PSMC4,TMEM126A,POLR2E,TRIM28,ATP5IF1,MCRS1,SMIM29,KARS1,YIF1B,PPAN,PRR14,RPS23,ARL2,IFI27L2,NUDT1,RPL23,MRPS18B,ETFB,TMEM141,KLF16,ABHD12,HCFC1R1,OCEL1,COMMD4,NDUFA3,DPCD,B9D2,MTX1,TMEM134,NDUFA1,SMIM4,NUBP2,NSMCE1,TBCB,C19orf73,CDC20 | | Exist in public source | | NA |
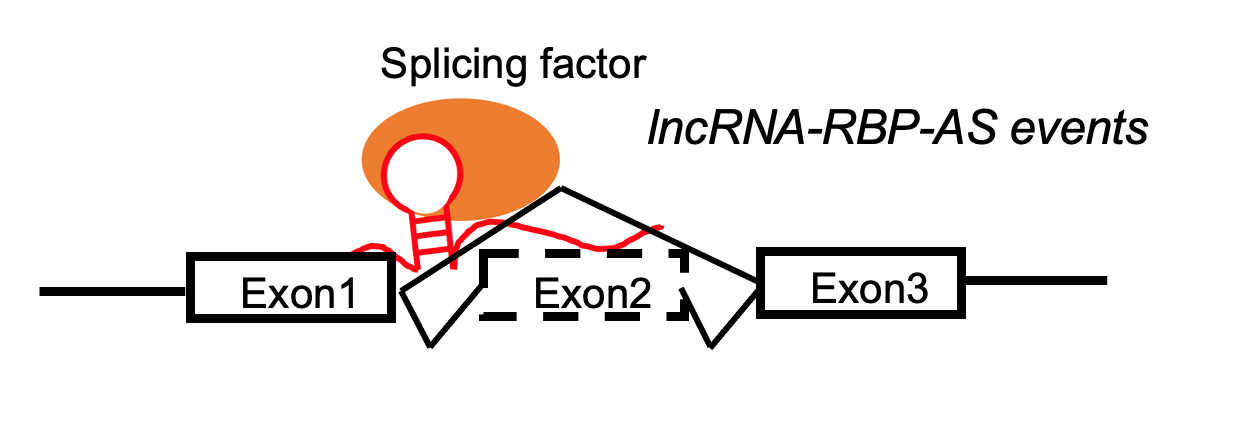
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000214106 | ENST00000449486 | exon_skip_89597 | chr12:6672349-6672532 | -25.31 | -0.1654 | ZNF384 | ENST00000361959,ENST00000396801 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_111672 | chr14:23089981-23090101 | -17.93 | -0.2214 | ACIN1 | ENST00000262710 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_140339 | chr16:635611-635774 | -19.72 | -0.1529 | METTL26 | ENST00000301686 | Frame-shift | | ENSG00000214106 | ENST00000449486 | exon_skip_150854 | chr17:32366664-32366757 | -13.82 | -0.1536 | ZNF207 | ENST00000321233 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_291980 | chr17:59119318-59119495 | -15.16 | -0.1046 | SKA2 | ENST00000330137 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_319500 | chr19:45413962-45414034 | -17.88 | -0.3031 | ERCC1 | ENST00000300853,ENST00000589165 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_322579 | chr19:55455635-55455845 | -27.66 | -0.1581 | ISOC2 | ENST00000425675 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_390412 | chr3:183833723-183833825 | -16.11 | -0.2404 | PARL | ENST00000317096 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_426982 | chr4:163515312-163515461 | -17.07 | -0.1552 | TMA16 | ENST00000358572 | Frame-shift | | ENSG00000214106 | ENST00000449486 | exon_skip_445390 | chr5:150403120-150403312 | -23.18 | -0.1302 | CD74 | ENST00000009530 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_390418 | chr3:183842297-183842447 | -13.69 | -0.1574 | PARL | ENST00000317096 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000214106 | ENST00000449486 | exon_skip_99856 | chr13:39042104-39042305 | -13.79 | -0.1661 | NHLRC3 | ENST00000379600 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_98384 | chr12:124327408-124327633 | -31.19 | -0.1386 | NCOR2 | ENST00000405201 | In-frame | | ENSG00000214106 | ENST00000449486 | exon_skip_482403 | chr8:27445793-27445919 | -14.08 | -0.1154 | PTK2B | ENST00000346049,ENST00000397501 | In-frame |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
 |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000214106 | hsa-mir-1307 | ENST00000411526 | chr7:154928641-154928802 | 243.00 | -137.10 | 161 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000411526 | chr7:154928843-154928990 | 212.00 | -121.31 | 157 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000411526 | chr7:154929509-154929653 | 208.00 | -121.16 | 152 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000411526 | chr7:154929234-154929378 | 191.00 | -106.45 | 146 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000411526 | chr7:154929022-154929174 | 175.00 | -105.30 | 148 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000411526 | chr7:154928500-154928638 | 164.00 | -110.02 | 143 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154928641-154928802 | 243.00 | -137.10 | 161 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154929908-154930050 | 229.00 | -126.30 | 155 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154928843-154928990 | 212.00 | -121.31 | 157 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154929597-154929744 | 199.00 | -120.09 | 148 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154929234-154929378 | 191.00 | -106.45 | 146 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154929022-154929174 | 175.00 | -105.30 | 148 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154928518-154928638 | 158.00 | -103.47 | 129 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154929423-154929569 | 157.00 | -106.60 | 155 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154930042-154930193 | 153.00 | -113.91 | 149 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000397551 | chr7:154929758-154929905 | 152.00 | -119.11 | 155 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154928641-154928802 | 243.00 | -137.10 | 161 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154930275-154930428 | 231.00 | -133.62 | 150 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154929960-154930108 | 216.00 | -129.35 | 145 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154929553-154929700 | 214.00 | -127.28 | 151 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154928843-154928990 | 212.00 | -121.31 | 157 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154930473-154930624 | 204.00 | -128.05 | 153 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154929734-154929895 | 197.00 | -127.21 | 161 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154929234-154929378 | 191.00 | -106.45 | 146 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154929022-154929174 | 175.00 | -105.30 | 148 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154929380-154929540 | 155.00 | -123.34 | 158 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154928529-154928638 | 151.00 | -101.39 | 113 | NA | | ENSG00000214106 | hsa-mir-1307 | ENST00000449486 | chr7:154930121-154930275 | 140.00 | -97.53 | 158 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154928947-154929046 | 186.00 | -52.16 | 82 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154928518-154928605 | 185.00 | -52.29 | 81 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154929222-154929317 | 171.00 | -53.72 | 63 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154928806-154928907 | 166.00 | -78.06 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154929362-154929453 | 155.00 | -53.68 | 57 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154928648-154928742 | 150.00 | -67.18 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154929063-154929154 | 142.00 | -53.44 | 84 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000411526 | chr7:154929299-154929383 | 141.00 | -53.97 | 52 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154928947-154929046 | 186.00 | -52.16 | 82 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154928518-154928605 | 185.00 | -52.29 | 81 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929222-154929317 | 171.00 | -53.72 | 63 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929803-154929888 | 169.00 | -57.44 | 78 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154928806-154928907 | 166.00 | -78.06 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929493-154929580 | 155.00 | -52.67 | 54 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929935-154930027 | 154.00 | -65.79 | 71 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929639-154929729 | 153.00 | -57.65 | 46 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154928648-154928742 | 150.00 | -67.18 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929063-154929154 | 142.00 | -53.44 | 84 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929355-154929433 | 142.00 | -50.72 | 54 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000397551 | chr7:154929299-154929383 | 141.00 | -53.97 | 52 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154928947-154929046 | 186.00 | -52.16 | 82 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154928529-154928605 | 183.00 | -50.28 | 79 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154930024-154930115 | 176.00 | -64.26 | 82 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929222-154929317 | 171.00 | -53.72 | 63 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154930534-154930620 | 167.00 | -63.24 | 57 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154928806-154928907 | 166.00 | -78.06 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929733-154929833 | 165.00 | -71.68 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929853-154929927 | 165.00 | -55.04 | 83 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154930306-154930395 | 160.00 | -65.08 | 86 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154930064-154930148 | 159.00 | -56.95 | 41 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929362-154929453 | 155.00 | -53.68 | 57 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154928648-154928742 | 150.00 | -67.18 | 93 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929513-154929598 | 149.00 | -53.84 | 59 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929561-154929649 | 144.00 | -63.25 | 51 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154930451-154930537 | 143.00 | -61.85 | 78 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929063-154929154 | 142.00 | -53.44 | 84 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929299-154929383 | 141.00 | -53.97 | 52 | NA | | ENSG00000214106 | hsa-mir-3613 | ENST00000449486 | chr7:154929940-154930020 | 140.00 | -58.78 | 48 | NA | | ENSG00000214106 | hsa-mir-18a | ENST00000411526 | chr7:154929451-154929533 | 153.00 | -53.81 | 77 | NA | | ENSG00000214106 | hsa-mir-18a | ENST00000449486 | chr7:154929451-154929533 | 153.00 | -53.81 | 77 | NA | | ENSG00000214106 | hsa-mir-18a | ENST00000449486 | chr7:154930553-154930628 | 142.00 | -51.09 | 74 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000411526 | chr7:154929660-154929738 | 185.00 | -52.42 | 70 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000411526 | chr7:154929429-154929501 | 165.00 | -55.70 | 71 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000411526 | chr7:154929146-154929217 | 153.00 | -56.02 | 61 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000397551 | chr7:154929146-154929217 | 153.00 | -56.02 | 61 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000449486 | chr7:154929960-154930027 | 172.00 | -55.08 | 69 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000449486 | chr7:154929429-154929501 | 165.00 | -55.70 | 71 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000449486 | chr7:154929146-154929217 | 153.00 | -56.02 | 61 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000449486 | chr7:154930406-154930475 | 146.00 | -60.94 | 71 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000449486 | chr7:154929863-154929931 | 144.00 | -52.07 | 71 | NA | | ENSG00000214106 | hsa-mir-32 | ENST00000449486 | chr7:154929509-154929586 | 141.00 | -52.79 | 77 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
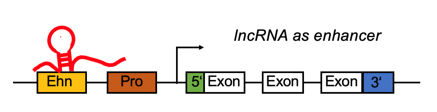
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000214106 | ENSG00000005469 | CROT | ACC,BLCA,BRCA,DLBC,GBM,KIRC,KIRP,LAML,LGG,OV,PAAD,READ,SARC,SKCM,TGCT,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000006756 | ARSD | ACC,BLCA,CESC,GBM,KIRP,LGG,LIHC,TGCT,THYM | GBM | | | ENSG00000214106 | ENSG00000073331 | ALPK1 | ACC,BLCA,BRCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,KICH,KIRP,LAML,PAAD,READ,SARC,SKCM,STAD,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000102699 | PARP4 | ACC,BLCA,CESC,CHOL,DLBC,GBM,KICH,KIRC,KIRP,LAML,LGG,LIHC,PAAD,TGCT,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000106049 | HIBADH | ACC,CESC,DLBC,GBM,KICH,KIRC,KIRP,LAML,SARC,TGCT,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000114473 | IQCG | ACC,GBM,KIRP,LGG,THCA | GBM | | | ENSG00000214106 | ENSG00000122783 | CYREN | ACC,CHOL,GBM,KIRP,LAML,LGG,LIHC,READ,STAD,THCA,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000128513 | POT1 | ACC,BLCA,BRCA,CESC,CHOL,DLBC,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,LIHC,PAAD,READ,SARC,SKCM,THCA,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000131269 | ABCB7 | ACC,BRCA,CHOL,GBM,KICH,KIRC,LIHC,PAAD,SKCM,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000132122 | SPATA6 | ACC,BLCA,BRCA,CESC,CHOL,COAD,ESCA,GBM,KIRC,KIRP,LGG,PAAD,READ,SARC,STAD,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000139192 | TAPBPL | ACC,CHOL,GBM,LGG,TGCT | GBM | | | ENSG00000214106 | ENSG00000139350 | NEDD1 | ACC,CHOL,COAD,DLBC,GBM,KICH,KIRC,KIRP,LAML,LGG,LIHC,PAAD,READ,SKCM,THCA,UCEC | GBM | | | ENSG00000214106 | ENSG00000146833 | TRIM4 | ACC,BLCA,BRCA,CESC,CHOL,DLBC,GBM,KICH,KIRC,KIRP,LAML,LGG,LIHC,SARC,SKCM,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000155189 | AGPAT5 | ACC,CHOL,DLBC,GBM,KIRC,KIRP,THYM | GBM | | | ENSG00000214106 | ENSG00000172037 | LAMB2 | ACC,ESCA,GBM,LGG,STAD,TGCT,UCS | GBM | | | ENSG00000214106 | ENSG00000177119 | ANO6 | ACC,BLCA,BRCA,CHOL,COAD,DLBC,ESCA,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,READ,STAD,TGCT,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000181826 | RELL1 | ACC,BRCA,CHOL,ESCA,GBM,KIRC,KIRP,LGG,PAAD,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000197324 | LRP10 | ACC,CHOL,DLBC,GBM,LIHC,TGCT,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000205084 | TMEM231 | ACC,GBM,LGG,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000256043 | CTSO | ACC,BLCA,BRCA,CESC,COAD,DLBC,ESCA,GBM,KICH,KIRC,KIRP,LAML,LIHC,PAAD,READ,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000064012 | CASP8 | BLCA,CESC,CHOL,COAD,DLBC,GBM,LIHC,PAAD,SARC,SKCM,THYM | GBM | | | ENSG00000214106 | ENSG00000103852 | TTC23 | BLCA,COAD,GBM,KIRP,LGG,READ,STAD,TGCT,THCA,UCEC | GBM | | | ENSG00000214106 | ENSG00000104365 | IKBKB | BLCA,CESC,CHOL,DLBC,GBM,KIRP,LAML,LGG,PAAD,TGCT,THCA,THYM | GBM | | | ENSG00000214106 | ENSG00000111684 | LPCAT3 | BLCA,CHOL,DLBC,GBM,KICH,SARC,TGCT,THCA,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000115825 | PRKD3 | BLCA,CESC,COAD,DLBC,GBM,KIRC,KIRP,LGG,LIHC,PAAD,READ,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000132256 | TRIM5 | BLCA,BRCA,CHOL,COAD,DLBC,GBM,KIRC,KIRP,LAML,LGG,LIHC,PAAD,SARC,SKCM,STAD,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000136560 | TANK | BLCA,BRCA,CHOL,COAD,DLBC,GBM,LGG,PAAD,READ,SKCM,TGCT,THYM | GBM | | | ENSG00000214106 | ENSG00000153790 | C7orf31 | BLCA,DLBC,GBM,KICH,KIRP,LAML,PAAD,READ,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000166801 | FAM111A | BLCA,BRCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,KICH,KIRC,KIRP,LAML,LGG,LIHC,PAAD,SKCM,TGCT,UCS | GBM | | | ENSG00000214106 | ENSG00000183323 | CCDC125 | BLCA,CESC,ESCA,GBM,KIRC,KIRP,LAML,LGG,UCS | GBM | | | ENSG00000214106 | ENSG00000198001 | IRAK4 | BLCA,CESC,CHOL,DLBC,GBM,KICH,KIRC,KIRP,LAML,LGG,LIHC,PAAD,SKCM,TGCT,THCA,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000102362 | SYTL4 | BRCA,COAD,GBM,THCA,UCEC | GBM | | | ENSG00000214106 | ENSG00000129675 | ARHGEF6 | BRCA,CESC,CHOL,COAD,DLBC,ESCA,GBM,KICH,LAML,PAAD,READ,SKCM,STAD,TGCT,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000137693 | YAP1 | BRCA,CHOL,DLBC,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000163482 | STK36 | BRCA,CESC,CHOL,DLBC,ESCA,GBM,KIRP,LGG,LIHC,SARC,THYM | GBM | | | ENSG00000214106 | ENSG00000168679 | SLC16A4 | BRCA,DLBC,GBM,LAML,LGG,SKCM,TGCT,THYM | GBM | | | ENSG00000214106 | ENSG00000172123 | SLFN12 | BRCA,CESC,CHOL,DLBC,ESCA,GBM,KIRC,KIRP,PAAD,SARC,SKCM,STAD,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000138587 | MNS1 | CESC,GBM,SKCM,TGCT,THCA,UCS | GBM | | | ENSG00000214106 | ENSG00000145365 | TIFA | CESC,CHOL,DLBC,GBM,KIRP,PAAD,SKCM,TGCT,THCA,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000059378 | PARP12 | CHOL,DLBC,GBM,LAML,LGG,SKCM | GBM | | | ENSG00000214106 | ENSG00000085719 | CPNE3 | CHOL,GBM,KICH,KIRC,PAAD,SARC,SKCM,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000101346 | POFUT1 | CHOL,DLBC,GBM,KIRC,KIRP,LGG,LIHC,TGCT,THCA,THYM | GBM | | | ENSG00000214106 | ENSG00000104450 | SPAG1 | CHOL,DLBC,GBM,LGG,THYM | GBM | | | ENSG00000214106 | ENSG00000115267 | IFIH1 | CHOL,GBM,KICH,KIRC,LGG,LIHC,PAAD,SKCM | GBM | | | ENSG00000214106 | ENSG00000140575 | IQGAP1 | CHOL,COAD,DLBC,GBM,KICH,KIRC,KIRP,LGG,LIHC,PAAD,THYM,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000162542 | TMCO4 | CHOL,GBM,LGG,LIHC,TGCT | GBM | | | ENSG00000214106 | ENSG00000168743 | NPNT | CHOL,GBM,KICH,LGG,TGCT,THYM | GBM | | | ENSG00000214106 | ENSG00000173905 | GOLIM4 | CHOL,DLBC,GBM,KICH,KIRC,KIRP,LIHC,PAAD,READ,TGCT,THCA,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000178202 | POGLUT3 | CHOL,COAD,DLBC,GBM,KICH,KIRC,LGG,LIHC,PAAD,STAD,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000213694 | S1PR3 | CHOL,DLBC,GBM,LGG,PAAD,STAD,TGCT | GBM | | | ENSG00000214106 | ENSG00000114790 | ARHGEF26 | COAD,GBM,LGG,READ,STAD,TGCT,THCA,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000114978 | MOB1A | COAD,DLBC,GBM,KIRC,KIRP,LGG,LIHC,PAAD,SKCM,TGCT,THCA,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000128791 | TWSG1 | COAD,DLBC,GBM,KICH,LGG,PAAD,READ,STAD,TGCT,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000132274 | TRIM22 | COAD,DLBC,GBM,LAML,LGG,PAAD,SKCM,STAD,THYM,UCEC | GBM | | | ENSG00000214106 | ENSG00000139178 | C1RL | COAD,DLBC,ESCA,GBM,LGG,SARC,SKCM,STAD,TGCT | GBM | | | ENSG00000214106 | ENSG00000089006 | SNX5 | DLBC,GBM,LGG,PAAD,THYM | GBM | | | ENSG00000214106 | ENSG00000106785 | TRIM14 | DLBC,GBM,LGG,LIHC,OV,PAAD,SKCM | GBM | | | ENSG00000214106 | ENSG00000121578 | B4GALT4 | DLBC,GBM,KIRP,LGG,READ,THYM,UCS | GBM | | | ENSG00000214106 | ENSG00000172936 | MYD88 | DLBC,GBM,KIRP,LGG,TGCT,THYM | GBM | | | ENSG00000214106 | ENSG00000173852 | DPY19L1 | DLBC,GBM,KICH,KIRP,LGG,LIHC,SARC,TGCT,UCEC,UCS | GBM | | | ENSG00000214106 | ENSG00000182179 | UBA7 | ESCA,GBM,OV,PAAD,SKCM,STAD,TGCT | GBM | | | ENSG00000214106 | ENSG00000117226 | GBP3 | GBM,LAML,LGG,LIHC,SKCM,TGCT | GBM | | | ENSG00000214106 | ENSG00000124067 | SLC12A4 | GBM,KIRP,LIHC,STAD,TGCT,THYM | GBM | | | ENSG00000214106 | ENSG00000131941 | RHPN2 | GBM,KICH,KIRP,LGG,LIHC,THCA | GBM | | | ENSG00000214106 | ENSG00000197763 | TXNRD3 | GBM,KICH,KIRC,KIRP,LAML,LIHC,TGCT,THCA,UCS | GBM | |
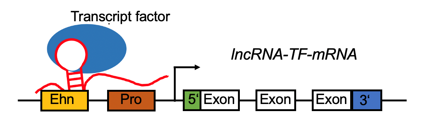
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000214106 | ENST00000411526 | FBXL15 | ENSG00000107872 | ENST00000224862 | -0.2215 | 583 | - | GBM | | ENSG00000214106 | ENST00000411526 | FBXL15 | ENSG00000107872 | ENST00000369956 | -0.2215 | 583 | - | GBM | | ENSG00000214106 | ENST00000397551 | FBXL15 | ENSG00000107872 | ENST00000224862 | -0.2215 | 564 | - | GBM | | ENSG00000214106 | ENST00000397551 | FBXL15 | ENSG00000107872 | ENST00000369956 | -0.2215 | 564 | - | GBM | | ENSG00000214106 | ENST00000449486 | FBXL15 | ENSG00000107872 | ENST00000224862 | -0.2215 | 553 | - | GBM | | ENSG00000214106 | ENST00000449486 | FBXL15 | ENSG00000107872 | ENST00000369956 | -0.2215 | 553 | - | GBM | | ENSG00000214106 | ENST00000411526 | UBALD1 | ENSG00000153443 | ENST00000587615 | -0.1268 | 196 | - | GBM | | ENSG00000214106 | ENST00000411526 | UBALD1 | ENSG00000153443 | ENST00000587649 | -0.1056 | 262 | - | GBM | | ENSG00000214106 | ENST00000397551 | UBALD1 | ENSG00000153443 | ENST00000587615 | -0.1268 | 177 | - | GBM | | ENSG00000214106 | ENST00000397551 | UBALD1 | ENSG00000153443 | ENST00000587649 | -0.1056 | 243 | - | GBM | | ENSG00000214106 | ENST00000449486 | UBALD1 | ENSG00000153443 | ENST00000587615 | -0.1268 | 166 | - | GBM | | ENSG00000214106 | ENST00000449486 | UBALD1 | ENSG00000153443 | ENST00000587649 | -0.1126 | 1 | - | GBM | | ENSG00000214106 | ENST00000411526 | PWWP2B | ENSG00000171813 | ENST00000631148 | -0.1002 | 341 | - | GBM | | ENSG00000214106 | ENST00000397551 | PWWP2B | ENSG00000171813 | ENST00000631148 | -0.1002 | 322 | - | GBM | | ENSG00000214106 | ENST00000449486 | PWWP2B | ENSG00000171813 | ENST00000631148 | -0.1002 | 311 | - | GBM | | ENSG00000214106 | ENST00000411526 | UROS | ENSG00000188690 | ENST00000368797 | -0.1021 | 168 | - | GBM | | ENSG00000214106 | ENST00000411526 | UROS | ENSG00000188690 | ENST00000368786 | -0.1262 | 283 | - | GBM | | ENSG00000214106 | ENST00000397551 | UROS | ENSG00000188690 | ENST00000368797 | -0.1021 | 149 | - | GBM | | ENSG00000214106 | ENST00000397551 | UROS | ENSG00000188690 | ENST00000368786 | -0.1262 | 264 | - | GBM | | ENSG00000214106 | ENST00000449486 | UROS | ENSG00000188690 | ENST00000622016 | -0.1159 | 1367 | - | GBM | | ENSG00000214106 | ENST00000449486 | UROS | ENSG00000188690 | ENST00000368797 | -0.1097 | 969 | - | GBM | | ENSG00000214106 | ENST00000449486 | UROS | ENSG00000188690 | ENST00000368786 | -0.1091 | 1859 | - | GBM | | ENSG00000214106 | ENST00000449486 | UROS | ENSG00000188690 | ENST00000462490 | -0.1028 | 1399 | - | GBM | | ENSG00000214106 | ENST00000449486 | UROS | ENSG00000188690 | ENST00000368774 | -0.2320 | 1986 | - | GBM | | ENSG00000214106 | ENST00000411526 | NDUFB8 | ENSG00000166136 | ENST00000299166 | -0.1425 | 585 | - | GBM | | ENSG00000214106 | ENST00000411526 | NDUFB8 | ENSG00000166136 | ENST00000370322 | -0.1311 | 588 | - | GBM | | ENSG00000214106 | ENST00000411526 | NDUFB8 | ENSG00000166136 | ENST00000370320 | -0.1108 | 541 | - | GBM | | ENSG00000214106 | ENST00000397551 | NDUFB8 | ENSG00000166136 | ENST00000299166 | -0.1425 | 566 | - | GBM | | ENSG00000214106 | ENST00000397551 | NDUFB8 | ENSG00000166136 | ENST00000370322 | -0.1311 | 569 | - | GBM | | ENSG00000214106 | ENST00000397551 | NDUFB8 | ENSG00000166136 | ENST00000370320 | -0.1108 | 522 | - | GBM | | ENSG00000214106 | ENST00000449486 | NDUFB8 | ENSG00000166136 | ENST00000299166 | -0.1425 | 555 | - | GBM | | ENSG00000214106 | ENST00000449486 | NDUFB8 | ENSG00000166136 | ENST00000370322 | -0.1311 | 558 | - | GBM | | ENSG00000214106 | ENST00000449486 | NDUFB8 | ENSG00000166136 | ENST00000370320 | -0.1388 | 1889 | - | GBM |
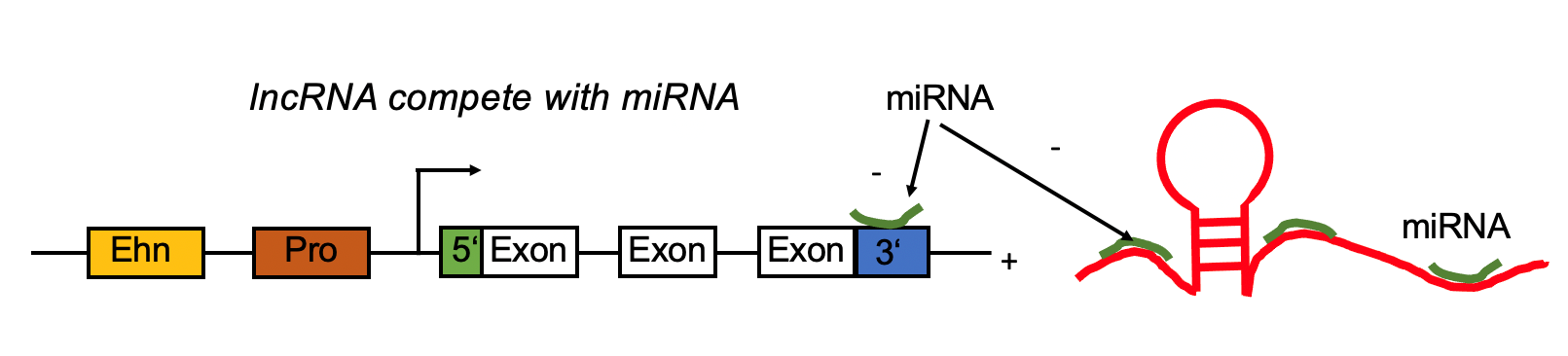
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,ASB8 | KICH,TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CRIM1 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,TRAK2 | BRCA,PRAD,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,RSPH3 | TGCT,BRCA,PRAD,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,SNX9 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,AHNAK | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,RBM43 | TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,TRIP11 | KICH,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CRY2 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,MINDY2 | KICH,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,FAM114A1 | TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,UEVLD | TGCT,BRCA,PRAD,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CBX7 | TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,SMIM14 | TGCT,BRCA,ACC,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,TMEM59 | TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,TOR1AIP1 | TGCT,BRCA,PRAD,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,AFF4 | BRCA,PRAD,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,ARL1 | KICH,TGCT,BRCA,PRAD,ACC | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CLIP1 | KICH,ACC,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,DIXDC1 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,ZBTB4 | TGCT,BRCA,PRAD,ACC,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CREBRF | BRCA,PRAD,ACC,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,EFCAB14 | TGCT,BRCA,PRAD,READ,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,LEPROT | TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,SERINC1 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,PJA2 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,HCFC2 | KICH,TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,SEL1L | KICH,TGCT,BRCA,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,MR1 | KICH,TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,SOS2 | KICH,TGCT,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,FYCO1 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,GOLM2 | KICH,TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,EVC | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CPEB4 | KICH,TGCT,BRCA,PRAD,ACC,READ | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,NFIC | TGCT,BRCA,ACC,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CROT | KICH,TGCT,BRCA,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,ANKMY2 | KICH,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,TGOLN2 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,CTSO | TGCT,BRCA,PRAD,READ,UCS | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,NDFIP1 | TGCT,BRCA,PRAD,UCS,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-18a,ARHGEF12 | TGCT,BRCA,PRAD,READ,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-3613,AHNAK | UCEC,BRCA,READ,GBM,KIRP | | ENSG00000214106 | PAXIP1-AS2,hsa-mir-1307,CBX7 | UCEC,KICH,BRCA,ACC,UCS |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000214106.6 | ENST00000411526.4 | 89 | 397 | 663 | | ENSG00000214106.6 | ENST00000397551.5 | 131 | 464 | 69 | | ENSG00000214106.6 | ENST00000449486.1 | 131 | 445 | 50 |
|

 Gene summary
Gene summary AS events and RNA A-to-I editing events of lncRNA in TCGA based on Genvode V22 structure.
AS events and RNA A-to-I editing events of lncRNA in TCGA based on Genvode V22 structure.
 Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).