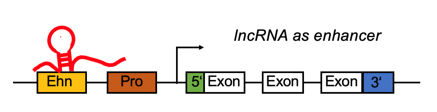
lncRNA targets the enhancer region and positively regulates the gene expression. lncRNA targets the enhancer region and positively regulates the gene expression. |
| Only predicted by TDF | | RAB23,ARHGEF17,FERMT2,FKBP8,KCNMB1,TCEA2,PLIN4,CTF1,IFT22,TMEM234,ACTN1,CIRBP,PLLP,DYNC2LI1,RNF166,DAPK3,WDR83OS,KLF2,CHRDL2,FKBP7,FOXF2,ACD,MIEF2,ISCU,CENPB,SLC25A12,GNAZ,EDIL3,LIMS2,HIRIP3,PELI3,TERF2IP,SGCB,RAB34,TRIP10,CCDC8,NIBAN1,SVBP,CLEC14A,TAZ,TACR2,USE1,CYB5D2,DIPK2B,CDC42EP5,CA11,VKORC1,MEIS2,RARA,RING1,SLC22A17,CHKB,CBY1,ITGA7,RAB3IL1,C3orf18,PDZD4,SERTAD4,SGCA,CCDC9B,FRZB,LRRN4CL,PAMR1,S1PR3,FEZ1,LOXL1,RBM42,MYO1C,FGFR1,CBX7,FOXF1,TSPAN2,COPZ2,ITPKB,SH3RF3,WWTR1,PPP1R14A,C22orf39,FUZ,TCN2,SSBP2,LAMA4,CHRDL1,TES,FAM229B,PLEKHA4,PCOLCE,ARHGEF40,FZD7,FBLN5,MRC2,KCTD12,MOAP1,CCDC69,SLC12A4,DZIP1,CZIB,CPXM1,THBS4,TSPAN4,EHD2,IFT122,HSPB7,CCDC24,GPX8,AHDC1,CCDC12,LAMB2,TXNIP,ARHGEF37,MEF2D,MCRIP1,CRLF1,PHYHD1,CLU,ABHD8,A2M,TRIR,DNALI1,ARMCX2,MRPL55,PRUNE2,SNRNP25,CRIP2,PPP1R12C,PPOX,BBOF1,PRICKLE2,PDZRN3,NEXN,C19orf53,DNAJB5,GUK1,KAZALD1,ATP8B2,ANG,DNAJB4,LAYN,APBB1,JPH2,ZNF25,GNPTG,SDC3,C11orf68,ADD1,EPN2,SSC5D,PIP5K1C,TMEM8B,TMEM220,POPDC2,NPR2,CACNA1C,HTRA1,DACT3,EFEMP2,MTURN,PKD1,OSBPL1A,NME4,CRYAB,TMEM43,LPAR1,SHISA4,MEIS1,HAND2,MRGPRF,NELFE,NFIC,EXD3,GLT8D1,P4HTM,ABTB1,STX8,EVA1B,VIM,FILIP1L,LMOD1,SV2A,SYCE1L,LMF1,MCAM,PPP1R3F,AKT3,THOC6,ST3GAL3,ZNF48,DENND6B,ACACB,FAXDC2,IL11RA,LTBP4,TLE2,PDGFD,COL16A1,OLFML3,PDLIM3,DNAJC18,LRP3,VPS4A,TUBB6,ACTA2,AHNAK,HACD1,CACNA1H,ILK,CD2BP2,TNFAIP8L3,PTOV1,PYURF,SGCE,TMEM35A,SLC10A3,KHDC1,KCNMA1,SFRP1,C1R,TEAD3,ORAI3,OSBPL5,PDLIM7,CNPY4,CUEDC2,CCDC80,LDB2,MYH11,DENND2A,DSTN,PER1,SHF,BMERB1,S100A13,DIXDC1,CROCC,DISP1,MFAP4,B3GNT9,ARHGEF25,TRIOBP,BAG2,CLDN5,ENKD1,KCNJ8,ZDHHC1,BLOC1S1,IFFO1,MORN2,BCL2,CLEC3B,ZBTB4,CCDC107,CTBP1,MEIS3,MDM1,COL6A2,IGFBP7,CDPF1,SBSPON,TTC23,OLFML1,NAT14,P3H3,TNS1,SPARCL1,C1orf35,ROR2,PIGC,DMPK,C14orf28,C1orf21,ZER1,GABBR1,USP11,H2AJ,SGSM3,TP53I13,WDR45,CHST14,CNRIP1,FYCO1,TGFBR3,PBXIP1,CLMP,NREP,ARMC5,ST6GALNAC6,ACBD4,RBPMS,OLFML2A,CAMK2G,SVIL,RBM43,RAMP2,CD151,SEMA6C,SH3BGR,DLC1,ZNF34,MYL6,KLHDC8B,CAVIN2,PHF1,TMEM59L,CETN2,CCN1,MFAP2,ANKS3,NOXA1,UCK1,SMDT1,AOC3,PIGQ,S100B,LMNA,COX4I1,SCMH1,C16orf89,NAB2,FBXO32,RASL12,GAS1,PPM1M,SPOP,CIAO2B,MFGE8,HSPB6,AAMDC,MYL6B,FAM234A,JCAD,CSPG4,PLSCR4,DTX3,REEP1,TLE5,KITLG,DENND2B,LUC7L,ARHGAP31,COA5,PDGFC,PDE3A,ECSCR,GLT8D2,C14orf93,TNS2,SMARCD3,TMEM47,TAGLN,CSRP2,HDAC4,TINAGL1,NDRG2,CRELD1,NRIP2,LTBP3,RAI2,BDH2,DPT,B9D2,NOP53,IRF2BP1,HEBP2,SMIM19,ANGPTL2,MAOB,SLC25A27,MRPS26,ACKR1,PCSK1N,VSTM4,CYS1,RRNAD1,CD99L2,TGFB1I1,CCDC92,PTGIS,PMP22,RBPMS2,RP11-523H20.5,REEP2,LHFPL6,RILPL1,PPIC,MYL9,OSCP1,RPS6KA2,SSPN,BCL7C,CALHM2,FHL3,GREM2,TSPAN18,TMEM9,KCNH2,FLNA,MXRA7,IFT27,DCHS1,PMF1,POLR2J,LRRC75B,HOPX,PPP1R3C,NT5C3B,RGS5,DLG4,TIMP2,C1orf216,CPXM2,CTSO,EMP3,SHC2,CALCOCO1,CFAP410,IGFBP4,VAMP2,TUSC1,CTSF,OAZ2,MXRA8,CCDC102A,SYT11,RERG,SCARF2,CAVIN1,IL17RD,LCAT,UBL5,PSD,ITIH5,GJC1,CDC42EP3,GPC4,CYGB,TTLL1,HAAO,ZYX,THYN1,RRAS,PTPN21,PRAF2,EID1,SORBS1,FAM228B,SLC25A23,PTMS,PCBP4,MTFR1L,VEGFB,CRYZL1,MOSPD3,MOB2,COMMD6,DPYSL3,ITPR1,LZTS2,TMA7,RABGAP1,FLOT1,KANK2,GAS6,PER3,EMCN,GEM,CLIP3,BOLA1,YPEL3,PLEKHO1,COLEC12,PACSIN3,GNG7,ANTXR2,TUBA1A,GNL1,COL8A2,MTMR11,REX1BD,LGALS1,RCBTB2,ASB2,TPM2,FAM20C,TRAPPC2 | | Exist in public source | | NA |
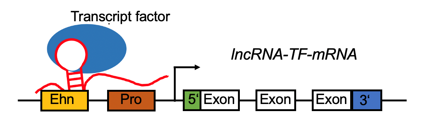
 lncRNA targets the promoter region and positively regulates the gene expression. lncRNA targets the promoter region and positively regulates the gene expression. |
| Only predicted by TDF | | SGSM3,C22orf39,CLMP,ECSCR,SYDE1,RAB23,DCHS1,PMF1,CA11,CNPY4,CSPG4,DMPK,RASD2,MYL6B,EFHC1,MSRB3,TCEAL8,GPC4,EDIL3,SMOC2,TNFSF12,CUEDC2,TADA3,GLT8D1,TNS1,HTRA3,TMEM9,P3H3,COL6A2,FKBP8,PPP1R12B,SV2A,GPX7,MBNL1,ZER1,MAOB,HACD1,TBC1D1,ORAI3,OAF,TPM2,TLE2,SYNGR1,PRICKLE2,ARHGEF17,LGALS1,FOXF1,NPR2,HSPA12B,PTGIS,OSCP1,TCEAL2,SRP14,MEF2D,GAS1,ZBTB4,LRRN4CL,KCTD12,GAB1,SRF,MFGE8,CNTNAP1,FOXF2,NUDT7,MANBAL,IGFBP6,VSTM4,PRELP,PER1,TBKBP1,MYL6,CRTC1,RBM42,CIRBP,RARRES2,ATP8B2,ELAC1,NDRG2,FAM234A,LOXL1,MEIS3,THOC6,SCNM1,ZNF48,S100A13,SLC22A17,DES,PDZRN3,CFH,TRIOBP,CYR61,PPDPF,SLC27A1,WDR13,HCFC1R1,ITPKB,EMILIN1,TXNIP,MAP7D1,CRTAP,LDB2,LPAR1,FBXO32,PDE3A,CYGB,SHC2,TAZ,PMP22,RRAGB | | Exist in public source | | NA |
 lncRNA targets the promoter region and negatively regulates the gene expression. lncRNA targets the promoter region and negatively regulates the gene expression. |
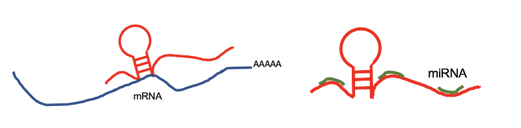
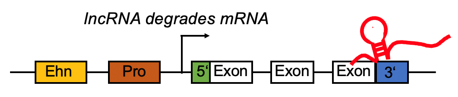
 lncRNA targets the 3'UTR region and negatively regulates the mRNA. lncRNA targets the 3'UTR region and negatively regulates the mRNA. |
| Only predicted by lncTar | | CNOT1,PGAM5,KPNB1,TRAP1,GTF3C2,PNPO,PPM1G,RACGAP1,AREL1,NAT10,QTRT2,PUS7,NOP14,BUB1B,BUB1,POLR1B,LARP1,PNPT1,TUBG1,PLEKHB2,EIF4G1,TUBA1C,BZW1,DPP3,DNAJC11,NADK,NSUN2,MRPL19,GARS1,CCDC47,MCCC2,CDCA5,SNX5,CCT5,HSPA4,EME1,USP32,NCAPH,PGD,G3BP1,PCCB,DKC1,SPAG5,FEN1,NOL6,CLPB,POLR1A,PRMT5,FANCD2,SRP54,GFM1,MFSD9,PSME3,MAPK1,SEC61A1,AHCY,RAD54L,AGFG1,HROB,R3HDM1,NIPA2,ORC6,HSPA9,RREB1,ATP5F1B,FARSA,TTF2,NUP98,NOP16,CCNF,DCAF7,CIAO1,RPN2,TRIP12,FANCI,RCC2,EFTUD2,SEC23B,ACLY,CSNK1A1,LARS2,NCAPD3,NAA50,CS,CAB39,ATIC,UGGT1,RPE,LSM12,ABCE1,MARS1,PRC1,DNAJA3,HJURP,CCNB1,CHEK1,EXO1,CSTF2,NDUFS1 | | Exist in public source | | NA |
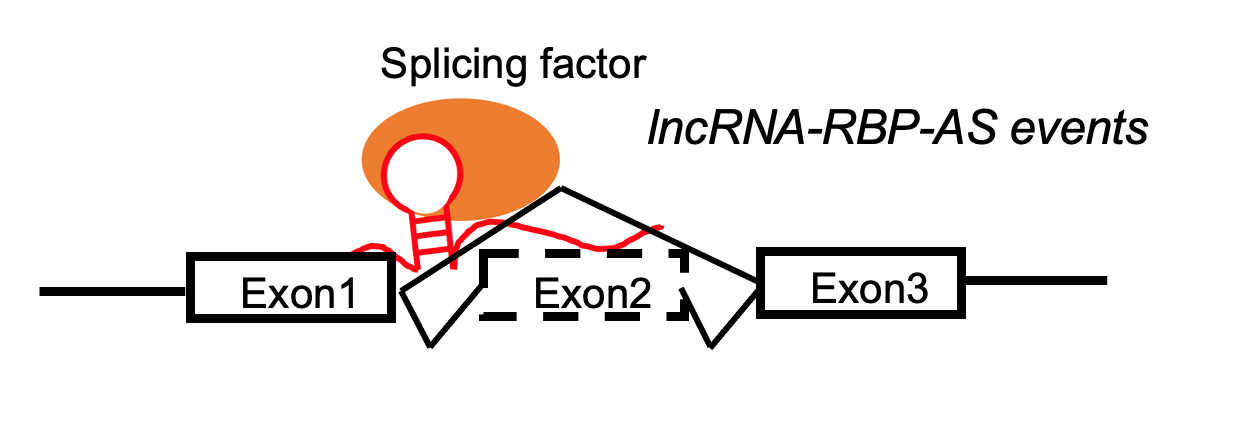
 lncRNA targets the skipped exon region. lncRNA targets the skipped exon region. | | -lncRNA and exonskipping events are positively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | ORF mutation | | ENSG00000203706 | ENST00000480052 | exon_skip_21610 | chr1:9737497-9737554 | -8.96 | -0.1906 | CLSTN1 | ENST00000377298 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_21628 | chr1:9756480-9756510 | -6.82 | -0.4012 | CLSTN1 | ENST00000377298 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_30682 | chr1:151310210-151310255 | -6.47 | -0.2311 | PI4KB | ENST00000368873 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_43507 | chr10:84499874-84499959 | -9.56 | -0.4780 | CCSER2 | ENST00000224756 | Frame-shift | | ENSG00000203706 | ENST00000480052 | exon_skip_75223 | chr11:72012400-72012442 | -6.38 | -0.1724 | NUMA1 | ENST00000393695 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_97739 | chr12:121244572-121244615 | -8.16 | -0.1990 | CAMKK2 | ENST00000324774,ENST00000402834,ENST00000404169 | Frame-shift | | ENSG00000203706 | ENST00000480052 | exon_skip_287874 | chr17:29085603-29085648 | -6.94 | -0.3305 | MYO18A | ENST00000527372 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_335378 | chr2:237751199-237751271 | -6.15 | -0.3844 | LRRFIP1 | ENST00000392000 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_436488 | chr5:96726793-96726859 | -8.47 | -0.2921 | CAST | ENST00000341926,ENST00000395813 | In-frame | | ENSG00000203706 | ENST00000437764 | exon_skip_470719 | chr7:117134123-117134192 | -7.44 | -0.3382 | ST7 | ENST00000265437 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_33516 | chr1:156938417-156938513 | -10.32 | -0.1306 | ARHGEF11 | ENST00000361409 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_48327 | chr10:26771074-26771089 | -4.72 | -0.4720 | ABI1 | ENST00000376142 | In-frame | | ENSG00000203706 | ENST00000437764 | exon_skip_111672 | chr14:23089981-23090101 | -19.74 | -0.1862 | ACIN1 | ENST00000262710 | In-frame |
| -lncRNA and exonskipping events are negatively correlated. |
 |
| LncRNA Ensembl ID | LncRNA ENST ID | Exon ID | Skipped Exon | dG | ndG | Gene name with skipped exon | TransID with skipped exon | LOF | | ENSG00000203706 | ENST00000437764 | exon_skip_7523 | chr1:78660553-78660654 | -10.99 | -0.1208 | IFI44 | ENST00000370747 | Frame-shift | | ENSG00000203706 | ENST00000437764 | exon_skip_132355 | chr16:1786756-1786955 | -36.75 | -0.2162 | NUBP2 | ENST00000262302 | Frame-shift | | ENSG00000203706 | ENST00000480052 | exon_skip_145776 | chr16:70665465-70665540 | -7.39 | -0.1680 | MTSS1L | ENST00000338779 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_289983 | chr17:42587833-42587864 | -6.29 | -0.5718 | RETREG3 | ENST00000309428 | Frame-shift | | ENSG00000203706 | ENST00000437764 | exon_skip_368399 | chr22:29825621-29825780 | -25.75 | -0.1966 | ASCC2 | ENST00000307790,ENST00000397771 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_27384 | chr1:54258068-54258149 | -10.46 | -0.1341 | SSBP3 | ENST00000371320 | In-frame | | ENSG00000203706 | ENST00000437764 | exon_skip_44853 | chr10:104010815-104010908 | -13.30 | -0.1446 | SLK | ENST00000369755 | In-frame | | ENSG00000203706 | ENST00000437764 | exon_skip_44998 | chr10:110132304-110132400 | -10.61 | -0.1632 | ADD3 | ENST00000356080 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_48713 | chr10:34336198-34336243 | -6.47 | -0.2022 | PARD3 | ENST00000374789 | In-frame | | ENSG00000203706 | ENST00000437764 | exon_skip_114200 | chr14:73279280-73279424 | -28.21 | -0.2029 | NUMB | ENST00000355058,ENST00000555238 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_148110 | chr17:4892401-4892512 | -11.49 | -0.1915 | MINK1 | ENST00000355280 | In-frame | | ENSG00000203706 | ENST00000480052 | exon_skip_324294 | chr2:27774423-27774530 | -11.37 | -0.1137 | MRPL33 | ENST00000296102 | Frame-shift | | ENSG00000203706 | ENST00000437764 | exon_skip_370224 | chr22:43137080-43137298 | -25.61 | -0.1231 | MCAT | ENST00000290429 | Frame-shift | | ENSG00000203706 | ENST00000480052 | exon_skip_456460 | chr6:17771113-17771218 | -13.82 | -0.1410 | KIF13A | ENST00000259711 | In-frame | | ENSG00000203706 | ENST00000437764 | exon_skip_462144 | chr6:125298713-125298816 | -14.36 | -0.1465 | HDDC2 | ENST00000398153 | Frame-shift | | ENSG00000203706 | ENST00000480052 | exon_skip_499771 | chr9:128592982-128593042 | -11.61 | -0.2322 | SPTAN1 | ENST00000372731 | In-frame |
 lncRNA targets by miRNA. lncRNA targets by miRNA. |
 |
| LncRNA Ensembl ID | miRNA ID | LncRNA ENST ID | Binding site in lncRNA | Score | Energy | Align Len | Public source | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000475406 | chr1:210231738-210231819 | 158.00 | -60.81 | 75 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000475406 | chr1:210231483-210231566 | 156.00 | -86.18 | 68 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000475406 | chr1:210231891-210231967 | 142.00 | -50.67 | 78 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000475406 | chr1:210231652-210231721 | 140.00 | -61.35 | 72 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000437764 | chr1:210231703-210231790 | 167.00 | -71.47 | 84 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000437764 | chr1:210231873-210231954 | 158.00 | -60.81 | 75 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000437764 | chr1:210231537-210231615 | 142.00 | -59.02 | 46 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000437764 | chr1:210232026-210232102 | 142.00 | -50.67 | 78 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000480052 | chr1:210231560-210231641 | 158.00 | -60.81 | 75 | NA | | ENSG00000203706 | hsa-mir-103a-1 | ENST00000480052 | chr1:210231713-210231789 | 142.00 | -50.67 | 78 | NA | | ENSG00000203706 | hsa-mir-103a-2 | ENST00000475406 | chr1:210231738-210231826 | 164.00 | -62.39 | 84 | NA | | ENSG00000203706 | hsa-mir-103a-2 | ENST00000437764 | chr1:210231873-210231961 | 164.00 | -62.39 | 84 | NA | | ENSG00000203706 | hsa-mir-103a-2 | ENST00000437764 | chr1:210231747-210231832 | 160.00 | -62.25 | 77 | NA | | ENSG00000203706 | hsa-mir-103a-2 | ENST00000437764 | chr1:210231611-210231689 | 142.00 | -60.55 | 69 | NA | | ENSG00000203706 | hsa-mir-103a-2 | ENST00000437764 | chr1:210231689-210231760 | 141.00 | -68.82 | 66 | NA | | ENSG00000203706 | hsa-mir-103a-2 | ENST00000480052 | chr1:210231560-210231648 | 164.00 | -62.39 | 84 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000475406 | chr1:210231883-210231969 | 165.00 | -68.82 | 82 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000475406 | chr1:210231572-210231646 | 157.00 | -89.12 | 84 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000475406 | chr1:210231722-210231801 | 141.00 | -78.10 | 75 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000437764 | chr1:210232018-210232104 | 165.00 | -68.82 | 82 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000437764 | chr1:210231857-210231936 | 145.00 | -82.76 | 83 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000437764 | chr1:210231532-210231623 | 142.00 | -81.91 | 85 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000480052 | chr1:210231705-210231791 | 165.00 | -68.82 | 82 | NA | | ENSG00000203706 | hsa-mir-142 | ENST00000480052 | chr1:210231544-210231623 | 149.00 | -78.25 | 83 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000475406 | chr1:210231700-210231778 | 176.00 | -84.10 | 79 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000475406 | chr1:210231531-210231610 | 163.00 | -111.34 | 79 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000475406 | chr1:210231457-210231534 | 146.00 | -97.99 | 70 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000475406 | chr1:210231611-210231693 | 142.00 | -82.34 | 77 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000437764 | chr1:210231821-210231913 | 199.00 | -100.66 | 92 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000437764 | chr1:210231544-210231624 | 182.00 | -85.22 | 83 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000437764 | chr1:210231678-210231759 | 167.00 | -100.60 | 82 | NA | | ENSG00000203706 | hsa-mir-185 | ENST00000480052 | chr1:210231519-210231600 | 146.00 | -76.07 | 69 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000475406 | chr1:210231832-210231965 | 213.00 | -86.90 | 129 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000475406 | chr1:210231729-210231833 | 171.00 | -88.96 | 109 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000475406 | chr1:210231589-210231704 | 165.00 | -104.71 | 114 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000437764 | chr1:210231967-210232100 | 213.00 | -86.90 | 129 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000437764 | chr1:210231457-210231549 | 186.00 | -71.40 | 92 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000437764 | chr1:210231859-210231968 | 175.00 | -91.45 | 111 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000437764 | chr1:210231734-210231852 | 145.00 | -110.22 | 104 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000480052 | chr1:210231654-210231787 | 213.00 | -86.90 | 129 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000480052 | chr1:210231457-210231552 | 186.00 | -73.04 | 92 | NA | | ENSG00000203706 | hsa-mir-7-1 | ENST00000480052 | chr1:210231551-210231655 | 171.00 | -88.79 | 109 | NA |
 RNA A-to-I editing events in lncRNA. RNA A-to-I editing events in lncRNA. |
| LncRNAediting ID | LncRNA Ensembl ID | Chromosome | Editing Position | Strand | Gene Type | Gene Name | Transcript ID | Transcript Type | Transcript Name |
 Edited-associated DElncRNAs in cancer. Edited-associated DElncRNAs in cancer. |
| LncRNA Ensembl ID | LncRNA Index | Cancer Type | Chr_Postion_Strand | AVE1 | AVE2 | log2FC | W-value | P-value | Adjc.p-value | Change |
 Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. Correlation between RNA A-to-I editing events's frequecy and lncRNA expression. |
| LncRNA Ensembl ID | LncRNA Index | Correlation | P-value | Adjc.p-value |
 Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. Cis-expression quantitative trait loci(cis-eQTL) of lncRNA. |
| LncRNA Ensembl ID | LncRNA Name | SNP info | Number of Positive corelated Cancer | Positive corelated Cancer | Number of Negative corelated Cancer | Negative corelated Cancer |
 lncRNA regulates differentially expressed genes by function as enhancer. lncRNA regulates differentially expressed genes by function as enhancer. |
| LncRNA Ensembl ID | PC Gene ID | PC Gene Name | Positive correlated cancers | Cancer with PC gene up-regulation | Cancer with PC gene down-regulation | | ENSG00000203706 | ENSG00000021762 | OSBPL5 | BLCA,PAAD,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000023902 | PLEKHO1 | BLCA,COAD,ESCA,HNSC,PRAD,READ,STAD,UCS | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000024422 | EHD2 | BLCA,COAD,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000035862 | TIMP2 | BLCA,CESC,COAD,ESCA,PRAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000079150 | FKBP7 | BLCA,COAD,ESCA,HNSC,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000082014 | SMARCD3 | BLCA,COAD,ESCA,HNSC,KIRP,READ,SARC,STAD,TGCT,THYM | KIRP | READ,STAD | | ENSG00000203706 | ENSG00000082497 | SERTAD4 | BLCA,BRCA,CESC,ESCA,HNSC,KICH,KIRC,KIRP,LUAD,LUSC,MESO,OV,PAAD,PCPG,PRAD,SARC,STAD,TGCT,THCA,THYM,UCEC,UCS | PCPG | UCEC | | ENSG00000203706 | ENSG00000084636 | COL16A1 | BLCA,MESO,PRAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000088543 | C3orf18 | BLCA,MESO,PRAD,READ,SARC,STAD,TGCT | | READ,STAD | | ENSG00000203706 | ENSG00000091986 | CCDC80 | BLCA,COAD,READ,STAD,TGCT,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000092841 | MYL6 | BLCA,COAD,KIRP,PRAD,READ,SARC,STAD,TGCT | | READ,STAD | | ENSG00000203706 | ENSG00000102181 | CD99L2 | BLCA,COAD,ESCA,HNSC,PRAD,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000203706 | ENSG00000105270 | CLIP3 | BLCA,COAD,ESCA,PCPG,PRAD,READ,SARC,STAD,TGCT,UCEC | PCPG | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000113296 | THBS4 | BLCA,COAD,READ,TGCT,UCS | | COAD,READ | | ENSG00000203706 | ENSG00000120820 | GLT8D2 | BLCA,COAD,ESCA,GBM,PRAD,READ,STAD,TGCT,THYM,UCEC,UCS | | UCEC | | ENSG00000203706 | ENSG00000126091 | ST3GAL3 | BLCA,COAD,HNSC,KIRP,MESO,READ,STAD,TGCT | | READ,STAD | | ENSG00000203706 | ENSG00000126458 | RRAS | BLCA,COAD,PRAD,READ,STAD,TGCT,THCA,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000127990 | SGCE | BLCA,CESC,COAD,ESCA,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000132563 | REEP2 | BLCA,SARC,STAD,TGCT,THCA | | STAD | | ENSG00000203706 | ENSG00000132718 | SYT11 | BLCA,COAD,PRAD,READ,STAD,TGCT | | COAD,READ | | ENSG00000203706 | ENSG00000137070 | IL11RA | BLCA,COAD,GBM,HNSC,KIRP,READ,STAD,TGCT,UCS | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000140682 | TGFB1I1 | BLCA,BRCA,CESC,COAD,ESCA,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,THCA,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000158270 | COLEC12 | BLCA,COAD,GBM,READ,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000159335 | PTMS | BLCA,HNSC,MESO,PCPG,STAD,THCA,THYM,UCS | PCPG | | | ENSG00000203706 | ENSG00000159884 | CCDC107 | BLCA,BRCA,COAD,KIRP,READ,SARC,STAD | | STAD | | ENSG00000203706 | ENSG00000159899 | NPR2 | BLCA,BRCA,CESC,COAD,ESCA,READ,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000203706 | ENSG00000163453 | IGFBP7 | BLCA,COAD,ESCA,PRAD,STAD,TGCT,THYM,UCEC | | UCEC | | ENSG00000203706 | ENSG00000164176 | EDIL3 | BLCA,COAD,ESCA,HNSC,READ,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000166341 | DCHS1 | BLCA,CESC,COAD,ESCA,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000169071 | ROR2 | BLCA,COAD,READ,SARC,STAD,THYM | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000170464 | DNAJC18 | BLCA,STAD,TGCT,THYM,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000171703 | TCEA2 | BLCA,ESCA,HNSC,KIRC,KIRP,MESO,PCPG,PRAD,STAD,TGCT,THCA | PCPG | | | ENSG00000203706 | ENSG00000177469 | CAVIN1 | BLCA,COAD,KIRP,PRAD,READ,STAD,TGCT,THCA,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000179954 | SSC5D | BLCA,COAD,GBM,PRAD,READ,STAD,TGCT,UCEC,UCS | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000183386 | FHL3 | BLCA,COAD,PRAD,STAD,THCA,UCEC | | UCEC | | ENSG00000203706 | ENSG00000196465 | MYL6B | BLCA,CESC,KIRC,KIRP,MESO,PCPG | PCPG | | | ENSG00000203706 | ENSG00000196923 | PDLIM7 | BLCA,BRCA,COAD,PAAD,READ,SARC,STAD,TGCT,THCA,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000203778 | FAM229B | BLCA,CESC,ESCA,HNSC,KIRP,READ,STAD,TGCT,UCS | | READ,STAD | | ENSG00000203706 | ENSG00000240771 | ARHGEF25 | BLCA,BRCA,COAD,ESCA,HNSC,PAAD,PCPG,PRAD,READ,STAD,TGCT,UCEC,UCS | PCPG | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000243279 | PRAF2 | BLCA,CESC,ESCA,HNSC,MESO,PCPG,STAD,TGCT | PCPG | | | ENSG00000203706 | ENSG00000005075 | POLR2J | BRCA,HNSC,KIRC,KIRP,UCS | KIRP | | | ENSG00000203706 | ENSG00000072163 | LIMS2 | BRCA,CESC,COAD,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000101335 | MYL9 | BRCA,CESC,COAD,ESCA,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000105701 | FKBP8 | BRCA,KIRC,MESO,PCPG,PRAD,THCA,UCS | PCPG | | | ENSG00000203706 | ENSG00000124074 | ENKD1 | BRCA,CESC,ESCA,HNSC,KIRC,KIRP,LUAD,MESO,PCPG,PRAD,THCA | KIRP | | | ENSG00000203706 | ENSG00000145936 | KCNMB1 | BRCA,CESC,COAD,PAAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000149591 | TAGLN | BRCA,CESC,COAD,ESCA,HNSC,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000159840 | ZYX | BRCA,KIRP,STAD,THCA,UCEC | KIRP | UCEC | | ENSG00000203706 | ENSG00000167641 | PPP1R14A | BRCA,CESC,COAD,ESCA,HNSC,LGG,PAAD,PRAD,READ,SARC,STAD | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000176014 | TUBB6 | BRCA,COAD,KIRP,PRAD,READ,STAD,TGCT,THCA,THYM | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000176533 | GNG7 | BRCA,STAD,TGCT,UCEC,UCS | | STAD,UCEC | | ENSG00000203706 | ENSG00000213398 | LCAT | BRCA,KIRP,MESO,PRAD,STAD,THYM | KIRP | | | ENSG00000203706 | ENSG00000004776 | HSPB6 | CESC,COAD,PAAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000014914 | MTMR11 | CESC,KIRP,PRAD,SARC,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000018408 | WWTR1 | CESC,COAD,PRAD,READ,STAD,TGCT,THYM | | COAD,READ | | ENSG00000203706 | ENSG00000069702 | TGFBR3 | CESC,COAD,GBM,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000071242 | RPS6KA2 | CESC,COAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000073712 | FERMT2 | CESC,COAD,HNSC,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000076555 | ACACB | CESC,COAD,LGG,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000079308 | TNS1 | CESC,COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000090006 | LTBP4 | CESC,COAD,PRAD,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000101938 | CHRDL1 | CESC,COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000103710 | RASL12 | CESC,COAD,GBM,MESO,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000107796 | ACTA2 | CESC,COAD,HNSC,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000109113 | RAB34 | CESC,COAD,ESCA,HNSC,KIRP,PRAD,READ,STAD,TGCT,THCA | KIRP | | | ENSG00000203706 | ENSG00000111077 | TNS2 | CESC,COAD,PRAD,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000116774 | OLFML3 | CESC,COAD,HNSC,PRAD,READ,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000128266 | GNAZ | CESC,MESO,PRAD,READ,SARC,STAD | | READ,STAD | | ENSG00000203706 | ENSG00000131831 | RAI2 | CESC,COAD,READ,STAD,TGCT,THYM | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000133392 | MYH11 | CESC,COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000134198 | TSPAN2 | CESC,COAD,GBM,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000134533 | RERG | CESC,COAD,GBM,PRAD,READ,SARC,STAD,TGCT,UCEC,UCS | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000138036 | DYNC2LI1 | CESC,GBM,HNSC,KIRP,LGG,TGCT | KIRP | | | ENSG00000203706 | ENSG00000138172 | CALHM2 | CESC,COAD,READ,TGCT,UCEC,UCS | | UCEC | | ENSG00000203706 | ENSG00000140092 | FBLN5 | CESC,COAD,PRAD,READ,STAD,UCEC,UCS | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000142871 | CCN1 | CESC,ESCA,HNSC,PRAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000146966 | DENND2A | CESC,ESCA,READ,STAD,TGCT,THYM,UCEC,UCS | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000147027 | TMEM47 | CESC,COAD,GBM,LGG,PRAD,READ,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000147113 | DIPK2B | CESC,COAD,PRAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000150281 | CTF1 | CESC,COAD,HNSC,KIRC,KIRP,LGG,MESO,PAAD,PRAD,SARC,STAD,TGCT,THCA,THYM | | STAD | | ENSG00000203706 | ENSG00000152583 | SPARCL1 | CESC,COAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000154309 | DISP1 | CESC,COAD,READ,STAD,TGCT,THYM,UCS | | STAD | | ENSG00000203706 | ENSG00000159239 | RP11-523H20.5 | CESC,KIRC,KIRP,LGG,LUAD,MESO,PCPG,TGCT,THCA | PCPG | | | ENSG00000203706 | ENSG00000160408 | ST6GALNAC6 | CESC,COAD,KICH,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000163069 | SGCB | CESC,COAD,READ,SARC,STAD,TGCT,THYM | | READ | | ENSG00000203706 | ENSG00000163431 | LMOD1 | CESC,COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000163815 | CLEC3B | CESC,COAD,KICH,PRAD,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000164741 | DLC1 | CESC,COAD,READ,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000165757 | JCAD | CESC,COAD,READ,THYM,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000165996 | HACD1 | CESC,COAD,KIRP,READ,SARC,STAD | | READ,STAD | | ENSG00000203706 | ENSG00000166313 | APBB1 | CESC,COAD,ESCA,HNSC,READ,SARC,STAD,UCEC,UCS | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000166333 | ILK | CESC,COAD,PAAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000166482 | MFAP4 | CESC,COAD,ESCA,GBM,HNSC,LGG,PRAD,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000166780 | BMERB1 | CESC,COAD,HNSC,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000166831 | RBPMS2 | CESC,COAD,LGG,PAAD,PRAD,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000167930 | FAM234A | CESC,HNSC,KIRP,MESO,PAAD,UCS | KIRP | | | ENSG00000203706 | ENSG00000169744 | LDB2 | CESC,COAD,READ,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000170271 | FAXDC2 | CESC,COAD,READ,SARC,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000172935 | MRGPRF | CESC,COAD,PAAD,READ,SARC,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000174080 | CTSF | CESC,COAD,ESCA,HNSC,READ,STAD,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000177363 | LRRN4CL | CESC,COAD,PAAD,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000177697 | CD151 | CESC,KIRP,TGCT,THCA,THYM | KIRP | | | ENSG00000203706 | ENSG00000178026 | LRRC75B | CESC,KIRC,KIRP,MESO,THYM,UCS | KIRP | | | ENSG00000203706 | ENSG00000179476 | C14orf28 | CESC,COAD,KIRP,PRAD,SARC,STAD,THYM,UCEC | | UCEC | | ENSG00000203706 | ENSG00000180354 | MTURN | CESC,COAD,READ,STAD,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000182809 | CRIP2 | CESC,COAD,KIRP,MESO,PCPG,TGCT,THCA | PCPG | | | ENSG00000203706 | ENSG00000185339 | TCN2 | CESC,COAD,MESO,PRAD,READ | | COAD,READ | | ENSG00000203706 | ENSG00000185437 | SH3BGR | CESC,COAD,GBM,LGG,PAAD,PRAD,READ,SARC,STAD | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000187824 | TMEM220 | CESC,GBM,LGG,READ,THYM | | READ | | ENSG00000203706 | ENSG00000197256 | KANK2 | CESC,COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000198121 | LPAR1 | CESC,COAD,ESCA,READ,TGCT,THYM,UCS | | COAD,READ | | ENSG00000203706 | ENSG00000198467 | TPM2 | CESC,COAD,ESCA,PAAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000205078 | SYCE1L | CESC,HNSC,KIRP,MESO,OV,TGCT,THCA | KIRP | | | ENSG00000203706 | ENSG00000220205 | VAMP2 | CESC,COAD,HNSC,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000255302 | EID1 | CESC,COAD,HNSC,KICH,READ,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000005243 | COPZ2 | COAD,GBM,HNSC,LUSC,MESO,PRAD,READ,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000010295 | IFFO1 | COAD,KIRP,PRAD,READ,STAD,UCEC | KIRP | UCEC | | ENSG00000203706 | ENSG00000012822 | CALCOCO1 | COAD,KICH,READ,SARC,STAD,TGCT,THYM,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000031081 | ARHGAP31 | COAD,READ,TGCT,UCEC,UCS | | UCEC | | ENSG00000203706 | ENSG00000049130 | KITLG | COAD,GBM,READ,TGCT,THYM | | COAD,READ | | ENSG00000203706 | ENSG00000049246 | PER3 | COAD,READ,STAD,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000055118 | KCNH2 | COAD,KIRP,READ,SARC,STAD | KIRP | STAD | | ENSG00000203706 | ENSG00000059915 | PSD | COAD,PAAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000063180 | CA11 | COAD,ESCA,MESO,PRAD,READ,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000068024 | HDAC4 | COAD,PAAD,READ,STAD,TGCT | | READ | | ENSG00000203706 | ENSG00000069535 | MAOB | COAD,LGG,PAAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000070778 | PTPN21 | COAD,READ,STAD,TGCT,THYM | | COAD,READ | | ENSG00000203706 | ENSG00000072110 | ACTN1 | COAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000075073 | TACR2 | COAD,PAAD,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000076706 | MCAM | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000077782 | FGFR1 | COAD,ESCA,HNSC,READ,STAD,TGCT,THYM | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000095637 | SORBS1 | COAD,PAAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000100307 | CBX7 | COAD,KICH,PRAD,READ,SARC,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000100628 | ASB2 | COAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000104332 | SFRP1 | COAD,GBM,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000104936 | DMPK | COAD,KIRP,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000106772 | PRUNE2 | COAD,PAAD,READ,SARC,STAD,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000109099 | PMP22 | COAD,HNSC,LGG,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000109846 | CRYAB | COAD,MESO,READ,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000111554 | MDM1 | COAD,GBM,LGG,READ,STAD | | READ | | ENSG00000203706 | ENSG00000112208 | BAG2 | COAD,ESCA,PAAD,READ,SARC,STAD,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000112210 | RAB23 | COAD,ESCA,PAAD,READ,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000112769 | LAMA4 | COAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000113657 | DPYSL3 | COAD,ESCA,HNSC,READ,SARC,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000114698 | PLSCR4 | COAD,PRAD,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000117020 | AKT3 | COAD,ESCA,READ,SARC,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000117640 | MTFR1L | COAD,KIRP,LUAD,PRAD,READ,STAD,TGCT | | STAD | | ENSG00000203706 | ENSG00000119865 | CNRIP1 | COAD,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000119938 | PPP1R3C | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000120885 | CLU | COAD,SARC,STAD,TGCT,THCA,THYM | | COAD,STAD | | ENSG00000203706 | ENSG00000121361 | KCNJ8 | COAD,PRAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000121440 | PDZRN3 | COAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000121577 | POPDC2 | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000121898 | CPXM2 | COAD,READ,SARC,STAD,UCS | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000123096 | SSPN | COAD,GBM,PRAD,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000124212 | PTGIS | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000124942 | AHNAK | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000125503 | PPP1R12C | COAD,KIRP,PAAD,PRAD,READ,STAD,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000125648 | SLC25A23 | COAD,PAAD,READ,SARC,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000125868 | DSTN | COAD,PAAD,PRAD,READ,SARC,STAD,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000126950 | TMEM35A | COAD,PRAD,READ,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000131471 | AOC3 | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000131477 | RAMP2 | COAD,HNSC,PRAD,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000135269 | TES | COAD,KIRP,READ,SARC,TGCT | KIRP | | | ENSG00000203706 | ENSG00000135842 | NIBAN1 | COAD,READ,SARC,STAD,TGCT,UCS | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000137094 | DNAJB5 | COAD,PAAD,READ,SARC,STAD,TGCT,THYM | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000141447 | OSBPL1A | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000141753 | IGFBP4 | COAD,ESCA,HNSC,KIRP,PRAD,STAD,TGCT,UCEC,UCS | | UCEC | | ENSG00000203706 | ENSG00000142227 | EMP3 | COAD,PRAD,READ,STAD,TGCT | | READ,STAD | | ENSG00000203706 | ENSG00000143196 | DPT | COAD,MESO,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000143515 | ATP8B2 | COAD,ESCA,READ,STAD,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000143772 | ITPKB | COAD,PAAD,READ,SARC,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000148660 | CAMK2G | COAD,PAAD,READ,SARC,STAD,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000149090 | PAMR1 | COAD,GBM,LGG,READ,STAD,TGCT,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000149596 | JPH2 | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000150764 | DIXDC1 | COAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000150995 | ITPR1 | COAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000154553 | PDLIM3 | COAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000155760 | FZD7 | COAD,LGG,PAAD,PRAD,READ,THYM | | READ | | ENSG00000203706 | ENSG00000156113 | KCNMA1 | COAD,PAAD,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000156804 | FBXO32 | COAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000157570 | TSPAN18 | COAD,HNSC,PAAD,READ,STAD,THYM,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000160307 | S100B | COAD,GBM,LGG,PRAD,READ,STAD | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000161544 | CYGB | COAD,GBM,MESO,PCPG,PRAD,READ | PCPG | | | ENSG00000203706 | ENSG00000162614 | NEXN | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000162616 | DNAJB4 | COAD,GBM,READ,STAD,THYM,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000162998 | FRZB | COAD,READ,STAD,TGCT,THYM,UCS | | READ,STAD | | ENSG00000203706 | ENSG00000163171 | CDC42EP3 | COAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000163297 | ANTXR2 | COAD,GBM,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000163820 | FYCO1 | COAD,READ,SARC,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000164035 | EMCN | COAD,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000164949 | GEM | COAD,GBM,READ,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000166250 | CLMP | COAD,GBM,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000166444 | DENND2B | COAD,READ,SARC,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000167994 | RAB3IL1 | COAD,HNSC,TGCT,THYM,UCEC | | UCEC | | ENSG00000203706 | ENSG00000168386 | FILIP1L | COAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000168497 | CAVIN2 | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000170962 | PDGFD | COAD,KIRP,PRAD,READ,TGCT | | COAD,READ | | ENSG00000203706 | ENSG00000171476 | HOPX | COAD,PRAD,READ,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000173546 | CSPG4 | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000173641 | HSPB7 | COAD,PAAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000175899 | A2M | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000176435 | CLEC14A | COAD,PRAD,READ,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000178498 | DTX3 | COAD,ESCA,HNSC,KIRP,PCPG,PRAD,READ,STAD,TGCT,THYM,UCEC,UCS | PCPG | READ,STAD | | ENSG00000203706 | ENSG00000178695 | KCTD12 | COAD,READ,TGCT,THYM,UCEC,UCS | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000180447 | GAS1 | COAD,READ,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000180875 | GREM2 | COAD,PAAD,READ,SARC,STAD,TGCT | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000182534 | MXRA7 | COAD,HNSC,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000183111 | ARHGEF37 | COAD,READ,SARC,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000183578 | TNFAIP8L3 | COAD,PRAD,READ,SARC,TGCT,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000183722 | LHFPL6 | COAD,ESCA,PRAD,READ,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000184113 | CLDN5 | COAD,PRAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000185585 | OLFML2A | COAD,ESCA,READ,TGCT,UCEC | | COAD,READ,UCEC | | ENSG00000203706 | ENSG00000185909 | KLHDC8B | COAD,READ,STAD,TGCT,THYM | | COAD,READ,STAD | | ENSG00000203706 | ENSG00000188549 | CCDC9B | COAD,READ,SARC,STAD,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000196557 | CACNA1H | COAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000196924 | FLNA | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000197321 | SVIL | COAD,PAAD,READ,SARC,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000197380 | DACT3 | COAD,PAAD,PRAD,READ,SARC,STAD,TGCT,THYM,UCEC,UCS | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000198624 | CCDC69 | COAD,LGG,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000204381 | LAYN | COAD,PRAD,READ,STAD,TGCT,THYM,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000213088 | ACKR1 | COAD,PRAD,READ,STAD,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000213694 | S1PR3 | COAD,READ,STAD,TGCT,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000249751 | ECSCR | COAD,READ,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000256043 | CTSO | COAD,READ,STAD,TGCT,THYM,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000265972 | TXNIP | COAD,READ,STAD,TGCT,UCEC | | COAD,READ,STAD,UCEC | | ENSG00000203706 | ENSG00000090971 | NAT14 | ESCA,HNSC,KIRP,MESO,PRAD | KIRP | | | ENSG00000203706 | ENSG00000103202 | NME4 | ESCA,HNSC,PAAD,SARC,STAD,TGCT | | STAD | | ENSG00000203706 | ENSG00000143434 | SEMA6C | ESCA,KIRP,MESO,PAAD,PCPG | PCPG | | | ENSG00000203706 | ENSG00000166848 | TERF2IP | ESCA,KIRP,PCPG,STAD,UCS | PCPG | | | ENSG00000203706 | ENSG00000175938 | ORAI3 | ESCA,GBM,HNSC,KIRP,TGCT,UCS | KIRP | | | ENSG00000203706 | ENSG00000164764 | SBSPON | GBM,SARC,STAD,TGCT,THYM | | STAD | | ENSG00000203706 | ENSG00000172572 | PDE3A | GBM,READ,STAD,TGCT,UCS | | READ,STAD | | ENSG00000203706 | ENSG00000198680 | TUSC1 | GBM,STAD,TGCT,THYM,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000205758 | CRYZL1 | GBM,PRAD,READ,SARC,STAD | | READ | | ENSG00000203706 | ENSG00000246705 | H2AJ | GBM,KIRP,LGG,MESO,TGCT | KIRP | | | ENSG00000203706 | ENSG00000006015 | REX1BD | HNSC,KIRP,MESO,PCPG,THCA | PCPG | | | ENSG00000203706 | ENSG00000007392 | LUC7L | HNSC,KIRP,PCPG,THCA,UCS | KIRP | | | ENSG00000203706 | ENSG00000102109 | PCSK1N | HNSC,KIRP,PCPG,THCA,UCS | PCPG | | | ENSG00000203706 | ENSG00000129946 | SHC2 | HNSC,KIRP,PCPG,STAD,THYM | KIRP,PCPG | | | ENSG00000203706 | ENSG00000167543 | TP53I13 | HNSC,KIRC,KIRP,MESO,PRAD,THCA | KIRP | | | ENSG00000203706 | ENSG00000168096 | ANKS3 | HNSC,KIRC,KIRP,MESO,PAAD,PCPG,TGCT | KIRP | | | ENSG00000203706 | ENSG00000204681 | GABBR1 | HNSC,KIRP,SARC,STAD,UCEC | KIRP | UCEC | | ENSG00000203706 | ENSG00000205795 | CYS1 | HNSC,LUSC,PRAD,READ,STAD,TGCT,THYM | | READ,STAD | | ENSG00000203706 | ENSG00000160445 | ZER1 | KICH,READ,SARC,STAD,TGCT | | STAD | | ENSG00000203706 | ENSG00000196459 | TRAPPC2 | KICH,MESO,PCPG,SARC,TGCT,UCS | PCPG | | | ENSG00000203706 | ENSG00000008710 | PKD1 | KIRP,MESO,PAAD,PRAD,SARC,STAD,TGCT,UCEC | | UCEC | | ENSG00000203706 | ENSG00000102125 | TAZ | KIRP,MESO,PCPG,PRAD,TGCT | KIRP | | | ENSG00000203706 | ENSG00000119242 | CCDC92 | KIRP,PCPG,STAD,TGCT,THYM | PCPG | | | ENSG00000203706 | ENSG00000136161 | RCBTB2 | KIRP,PRAD,READ,STAD,UCEC | | UCEC | | ENSG00000203706 | ENSG00000170876 | TMEM43 | KIRP,PRAD,STAD,TGCT,UCEC | KIRP | UCEC | | ENSG00000203706 | ENSG00000171791 | BCL2 | KIRP,READ,STAD,TGCT,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000145687 | SSBP2 | LGG,SARC,STAD,TGCT,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000067840 | PDZD4 | MESO,PCPG,PRAD,SARC,STAD | PCPG | STAD | | ENSG00000203706 | ENSG00000108823 | SGCA | PAAD,PRAD,READ,STAD,THYM,UCEC | | READ,STAD,UCEC | | ENSG00000203706 | ENSG00000116667 | C1orf21 | PAAD,READ,STAD,TGCT,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000151067 | CACNA1C | PAAD,READ,SARC,STAD,TGCT | | READ,STAD | | ENSG00000203706 | ENSG00000175183 | CSRP2 | PAAD,READ,STAD,TGCT,UCS | | STAD | | ENSG00000203706 | ENSG00000182963 | GJC1 | PAAD,PRAD,SARC,STAD,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000053702 | NRIP2 | PRAD,TGCT,THYM,UCEC,UCS | | UCEC | | ENSG00000203706 | ENSG00000163637 | PRICKLE2 | PRAD,SARC,STAD,TGCT,THYM,UCEC | | STAD,UCEC | | ENSG00000203706 | ENSG00000165795 | NDRG2 | PRAD,READ,TGCT,THYM,UCEC | | READ,UCEC | | ENSG00000203706 | ENSG00000167676 | PLIN4 | PRAD,READ,SARC,STAD,UCEC | | READ,STAD,UCEC |
 LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). LncRNA-TF complex positively regulates the gene expression by target promoter region (#cancer types with positive correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). LncRNA-TF complex negatively regulates the gene expression by target promoter region (#cancer types with negative correlation >= 5). |
| lncRNA ID | TF ID | TF Name | PCgene ID | PCgene Name | Number of Cancer | Canaer Types |
 LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex positively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. LncRNA-RBP complex negatively regulates the exon skippping events by target skipped eoxon region. |
| LncRNA Ensembl ID | RBP ID | RBP Gene Name | Exon Skipping ID | Skipped Exon | EX Gene Name | EX Affected TransID | ORF_anno | Cancer Type |
 lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. lncRNA regulates differential expressed mRNA by directly targeting 3' UTR region. |
| LncRNA Ensembl ID | LncRNA ENST ID | PC Gene Name | PC Gene ID | PC ENST ID | dG | nDG | Cancer with PC gene up-regulation | Cancer with PC gene Down-regulation | | ENSG00000203706 | ENST00000475406 | TUBA1C | ENSG00000167553 | ENST00000549818 | -0.1118 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | TUBA1C | ENSG00000167553 | ENST00000552125 | -0.1254 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | PGAM5 | ENSG00000247077 | ENST00000317555 | -0.1081 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | PGAM5 | ENSG00000247077 | ENST00000454808 | -0.1050 | 16 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | NOP16 | ENSG00000048162 | ENST00000619979 | -0.1578 | 398 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | NOP16 | ENSG00000048162 | ENST00000621444 | -0.5422 | 22 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | NOP16 | ENSG00000048162 | ENST00000614830 | -0.1147 | 69 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | NOP16 | ENSG00000048162 | ENST00000613081 | -0.5422 | 23 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | NOP16 | ENSG00000048162 | ENST00000502663 | -0.1151 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | SPAG5 | ENSG00000076382 | ENST00000321765 | -0.2770 | 1 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000475406 | SPAG5 | ENSG00000076382 | ENST00000582076 | -0.1077 | 1 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000475406 | SPAG5 | ENSG00000076382 | ENST00000378976 | -0.2699 | 1 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000475406 | SPAG5 | ENSG00000076382 | ENST00000578230 | -0.1107 | 17 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000475406 | SPAG5 | ENSG00000076382 | ENST00000580083 | -0.1159 | 1 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000475406 | RAD54L | ENSG00000085999 | ENST00000371975 | -0.1004 | 41 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | RAD54L | ENSG00000085999 | ENST00000488942 | -0.1266 | 8 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | RAD54L | ENSG00000085999 | ENST00000459678 | -0.1167 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | ORC6 | ENSG00000091651 | ENST00000563599 | -0.1011 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | PPM1G | ENSG00000115241 | ENST00000344034 | -0.1030 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | TTF2 | ENSG00000116830 | ENST00000427271 | -0.1171 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | NCAPH | ENSG00000121152 | ENST00000427946 | -0.1010 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | NCAPH | ENSG00000121152 | ENST00000435349 | -0.1299 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | HROB | ENSG00000125319 | ENST00000585683 | -0.1368 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | POLR1B | ENSG00000125630 | ENST00000424062 | -0.1329 | 1 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | TRAP1 | ENSG00000126602 | ENST00000246957 | -0.4368 | 228 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | TRAP1 | ENSG00000126602 | ENST00000575671 | -0.4368 | 229 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | TRAP1 | ENSG00000126602 | ENST00000538171 | -0.4368 | 229 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | TRAP1 | ENSG00000126602 | ENST00000574175 | -0.1453 | 1 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | TRAP1 | ENSG00000126602 | ENST00000571011 | -0.1113 | 2 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | MCCC2 | ENSG00000131844 | ENST00000509358 | -0.9000 | 554 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | NAT10 | ENSG00000135372 | ENST00000531159 | -0.1172 | 1 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | ATIC | ENSG00000138363 | ENST00000236959 | -0.1584 | 25 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | ATIC | ENSG00000138363 | ENST00000444305 | -0.1426 | 20 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | ATIC | ENSG00000138363 | ENST00000435675 | -0.1584 | 25 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | ATIC | ENSG00000138363 | ENST00000426233 | -0.3785 | 256 | COAD,READ | - | | ENSG00000203706 | ENST00000475406 | FANCI | ENSG00000140525 | ENST00000300027 | -0.1089 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | FANCI | ENSG00000140525 | ENST00000310775 | -0.1089 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | FANCI | ENSG00000140525 | ENST00000567996 | -0.1313 | 431 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | TUBA1C | ENSG00000167553 | ENST00000549818 | -0.1314 | 144 | UCEC | - | | ENSG00000203706 | ENST00000437764 | TUBA1C | ENSG00000167553 | ENST00000552125 | -0.1045 | 98 | UCEC | - | | ENSG00000203706 | ENST00000437764 | PGAM5 | ENSG00000247077 | ENST00000317555 | -0.1095 | 60 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | PGAM5 | ENSG00000247077 | ENST00000454808 | -0.1083 | 115 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | NOP16 | ENSG00000048162 | ENST00000619979 | -0.6040 | 269 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | NOP16 | ENSG00000048162 | ENST00000621444 | -0.2125 | 294 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | NOP16 | ENSG00000048162 | ENST00000614830 | -0.1077 | 131 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | NOP16 | ENSG00000048162 | ENST00000613081 | -0.2360 | 284 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | NOP16 | ENSG00000048162 | ENST00000616685 | -0.1097 | 181 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | NOP16 | ENSG00000048162 | ENST00000502663 | -0.1171 | 141 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | SPAG5 | ENSG00000076382 | ENST00000321765 | -0.1225 | 278 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000437764 | SPAG5 | ENSG00000076382 | ENST00000582076 | -0.1087 | 189 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000437764 | SPAG5 | ENSG00000076382 | ENST00000378976 | -0.1394 | 116 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000437764 | SPAG5 | ENSG00000076382 | ENST00000578230 | -0.1554 | 248 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000437764 | SPAG5 | ENSG00000076382 | ENST00000580083 | -0.1012 | 1 | UCEC,STAD | KIRP | | ENSG00000203706 | ENST00000437764 | RAD54L | ENSG00000085999 | ENST00000442598 | -0.1299 | 221 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | RAD54L | ENSG00000085999 | ENST00000371975 | -0.1018 | 122 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | RAD54L | ENSG00000085999 | ENST00000476687 | -0.1044 | 89 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | RAD54L | ENSG00000085999 | ENST00000488942 | -0.1452 | 155 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | RAD54L | ENSG00000085999 | ENST00000459678 | -0.1028 | 15 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | PPM1G | ENSG00000115241 | ENST00000344034 | -0.1125 | 129 | UCEC | - | | ENSG00000203706 | ENST00000437764 | TTF2 | ENSG00000116830 | ENST00000427271 | -0.1124 | 221 | UCEC | - | | ENSG00000203706 | ENST00000437764 | NCAPH | ENSG00000121152 | ENST00000427946 | -0.1037 | 202 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | NCAPH | ENSG00000121152 | ENST00000435349 | -0.1165 | 187 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | HJURP | ENSG00000123485 | ENST00000453122 | -0.1116 | 221 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | HROB | ENSG00000125319 | ENST00000585683 | -0.1120 | 127 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | TRAP1 | ENSG00000126602 | ENST00000246957 | -0.2159 | 363 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | TRAP1 | ENSG00000126602 | ENST00000575671 | -0.2159 | 364 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | TRAP1 | ENSG00000126602 | ENST00000538171 | -0.2159 | 364 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | TRAP1 | ENSG00000126602 | ENST00000574175 | -0.1622 | 333 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | TRAP1 | ENSG00000126602 | ENST00000571011 | -0.1343 | 205 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | MCCC2 | ENSG00000131844 | ENST00000509358 | -0.9000 | 689 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | CCNB1 | ENSG00000134057 | ENST00000507798 | -0.1065 | 219 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | ATIC | ENSG00000138363 | ENST00000236959 | -0.1242 | 128 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | ATIC | ENSG00000138363 | ENST00000444305 | -0.1603 | 324 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | ATIC | ENSG00000138363 | ENST00000435675 | -0.1242 | 128 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | ATIC | ENSG00000138363 | ENST00000426233 | -0.2533 | 309 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | FANCI | ENSG00000140525 | ENST00000567996 | -0.1313 | 566 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000480052 | PGAM5 | ENSG00000247077 | ENST00000454808 | -0.1090 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | NOP16 | ENSG00000048162 | ENST00000619979 | -0.5317 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | NOP16 | ENSG00000048162 | ENST00000613081 | -0.2629 | 229 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | TRAP1 | ENSG00000126602 | ENST00000575671 | -0.1164 | 51 | COAD,READ | - | | ENSG00000203706 | ENST00000480052 | TRAP1 | ENSG00000126602 | ENST00000538171 | -0.1164 | 51 | COAD,READ | - | | ENSG00000203706 | ENST00000480052 | MCCC2 | ENSG00000131844 | ENST00000509358 | -0.9000 | 376 | UCEC | PCPG | | ENSG00000203706 | ENST00000480052 | ATIC | ENSG00000138363 | ENST00000444305 | -0.1181 | 75 | COAD,READ | - | | ENSG00000203706 | ENST00000480052 | ATIC | ENSG00000138363 | ENST00000426233 | -0.5475 | 127 | COAD,READ | - | | ENSG00000203706 | ENST00000480052 | FANCI | ENSG00000140525 | ENST00000567996 | -0.1313 | 253 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | FANCI | ENSG00000140525 | ENST00000570225 | -0.1791 | 51 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | CDCA5 | ENSG00000146670 | ENST00000529290 | -0.2664 | 117 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | CDCA5 | ENSG00000146670 | ENST00000524733 | -0.1017 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | CDCA5 | ENSG00000146670 | ENST00000404147 | -1.3283 | 260 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | CHEK1 | ENSG00000149554 | ENST00000427383 | -0.1329 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | CHEK1 | ENSG00000149554 | ENST00000544373 | -0.1329 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | CHEK1 | ENSG00000149554 | ENST00000524737 | -0.1757 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | CHEK1 | ENSG00000149554 | ENST00000278916 | -0.1329 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | NCAPD3 | ENSG00000151503 | ENST00000534548 | -0.1097 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | EME1 | ENSG00000154920 | ENST00000338165 | -0.1130 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | EME1 | ENSG00000154920 | ENST00000393271 | -0.1130 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | EME1 | ENSG00000154920 | ENST00000511711 | -0.1042 | 18 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | EME1 | ENSG00000154920 | ENST00000511648 | -0.1130 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | BUB1B | ENSG00000156970 | ENST00000558715 | -0.1218 | 140 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | BUB1B | ENSG00000156970 | ENST00000559733 | -0.1041 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | RACGAP1 | ENSG00000161800 | ENST00000548961 | -0.6214 | 283 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | RACGAP1 | ENSG00000161800 | ENST00000548598 | -0.1078 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | RACGAP1 | ENSG00000161800 | ENST00000551260 | -0.1059 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000475406 | CCNF | ENSG00000162063 | ENST00000564236 | -0.1604 | 7 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | FEN1 | ENSG00000168496 | ENST00000535307 | -0.1458 | 1 | UCEC,STAD | - | | ENSG00000203706 | ENST00000475406 | BUB1 | ENSG00000169679 | ENST00000535254 | -0.1281 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | EXO1 | ENSG00000174371 | ENST00000518483 | -0.1454 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | EXO1 | ENSG00000174371 | ENST00000521202 | -0.2550 | 325 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | LARS2 | ENSG00000011376 | ENST00000414984 | -0.3179 | 88 | - | PCPG | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000294053 | -0.1127 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000437826 | -0.1127 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000538039 | -0.1127 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000535990 | -0.2422 | 64 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000340729 | -0.2422 | 64 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000543042 | -0.2739 | 234 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | CLPB | ENSG00000162129 | ENST00000536297 | -0.1383 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | DPP3 | ENSG00000254986 | ENST00000530165 | -0.2247 | 39 | UCEC | - | | ENSG00000203706 | ENST00000475406 | NDUFS1 | ENSG00000023228 | ENST00000449699 | -0.1925 | 477 | - | PCPG | | ENSG00000203706 | ENST00000475406 | NDUFS1 | ENSG00000023228 | ENST00000432169 | -0.1925 | 477 | - | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000251654 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000490504 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000483687 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000468777 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | FANCI | ENSG00000140525 | ENST00000570225 | -0.1933 | 94 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | FANCD2 | ENSG00000144554 | ENST00000435522 | -0.1268 | 1 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000437764 | CDCA5 | ENSG00000146670 | ENST00000529290 | -0.4100 | 286 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | CDCA5 | ENSG00000146670 | ENST00000404147 | -0.8712 | 147 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | CHEK1 | ENSG00000149554 | ENST00000524737 | -0.1198 | 292 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000437764 | EME1 | ENSG00000154920 | ENST00000511711 | -0.1026 | 182 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | BUB1B | ENSG00000156970 | ENST00000558715 | -0.1610 | 139 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000437764 | RACGAP1 | ENSG00000161800 | ENST00000548961 | -0.6214 | 418 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000437764 | CCNF | ENSG00000162063 | ENST00000564236 | -0.1512 | 199 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | EXO1 | ENSG00000174371 | ENST00000521202 | -0.2550 | 4 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | RCC2 | ENSG00000179051 | ENST00000375433 | -0.1144 | 1 | UCEC,STAD | - | | ENSG00000203706 | ENST00000437764 | LARS2 | ENSG00000011376 | ENST00000414984 | -0.4583 | 173 | - | PCPG | | ENSG00000203706 | ENST00000437764 | CLPB | ENSG00000162129 | ENST00000535990 | -0.2073 | 329 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | CLPB | ENSG00000162129 | ENST00000340729 | -0.2073 | 329 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | CLPB | ENSG00000162129 | ENST00000543042 | -0.2112 | 299 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | DPP3 | ENSG00000254986 | ENST00000530165 | -0.2397 | 290 | UCEC | - | | ENSG00000203706 | ENST00000437764 | NDUFS1 | ENSG00000023228 | ENST00000449699 | -0.1891 | 245 | - | PCPG | | ENSG00000203706 | ENST00000437764 | NDUFS1 | ENSG00000023228 | ENST00000432169 | -0.1891 | 245 | - | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000251654 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000490504 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000483687 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000468777 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000480052 | CDCA5 | ENSG00000146670 | ENST00000529290 | -0.4343 | 57 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000480052 | CDCA5 | ENSG00000146670 | ENST00000404147 | -0.3753 | 97 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000480052 | RACGAP1 | ENSG00000161800 | ENST00000548961 | -0.6214 | 105 | UCEC,COAD,STAD | - | | ENSG00000203706 | ENST00000480052 | CCNF | ENSG00000162063 | ENST00000564236 | -0.1147 | 75 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000480052 | EXO1 | ENSG00000174371 | ENST00000521202 | -0.2550 | 7 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000480052 | LARS2 | ENSG00000011376 | ENST00000414984 | -0.1394 | 97 | - | PCPG | | ENSG00000203706 | ENST00000480052 | CLPB | ENSG00000162129 | ENST00000535990 | -0.1494 | 133 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | CLPB | ENSG00000162129 | ENST00000340729 | -0.1494 | 133 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | CLPB | ENSG00000162129 | ENST00000543042 | -0.1638 | 168 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | DPP3 | ENSG00000254986 | ENST00000531863 | -0.1239 | 1 | UCEC | - | | ENSG00000203706 | ENST00000480052 | DPP3 | ENSG00000254986 | ENST00000532677 | -0.1239 | 1 | UCEC | - | | ENSG00000203706 | ENST00000480052 | DPP3 | ENSG00000254986 | ENST00000541961 | -0.1239 | 1 | UCEC | - | | ENSG00000203706 | ENST00000480052 | DPP3 | ENSG00000254986 | ENST00000530165 | -0.1432 | 107 | UCEC | - | | ENSG00000203706 | ENST00000480052 | NDUFS1 | ENSG00000023228 | ENST00000449699 | -0.1925 | 299 | - | PCPG | | ENSG00000203706 | ENST00000480052 | NDUFS1 | ENSG00000023228 | ENST00000432169 | -0.1925 | 299 | - | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000462637 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000466072 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000482086 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | PCCB | ENSG00000114054 | ENST00000469217 | -0.1023 | 51 | UCEC | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000346169 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000352767 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000414031 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000392537 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000382330 | -0.3030 | 55 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000350481 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000342981 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000424196 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000434061 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000319274 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000435046 | -0.1858 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | EIF4G1 | ENSG00000114867 | ENST00000422614 | -0.1685 | 33 | - | PCPG | | ENSG00000203706 | ENST00000475406 | NAA50 | ENSG00000121579 | ENST00000493900 | -0.1025 | 20 | - | PCPG | | ENSG00000203706 | ENST00000475406 | CCT5 | ENSG00000150753 | ENST00000511700 | -0.1043 | 9 | UCEC | - | | ENSG00000203706 | ENST00000475406 | PRC1 | ENSG00000198901 | ENST00000556972 | -0.1109 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | PRC1 | ENSG00000198901 | ENST00000555455 | -0.2164 | 1 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000475406 | DNAJC11 | ENSG00000007923 | ENST00000451196 | -0.1237 | 1 | - | KIRP,PCPG | | ENSG00000203706 | ENST00000475406 | NADK | ENSG00000008130 | ENST00000498806 | -0.1668 | 4 | - | PCPG | | ENSG00000203706 | ENST00000475406 | NADK | ENSG00000008130 | ENST00000489538 | -0.1149 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | BZW1 | ENSG00000082153 | ENST00000452790 | -0.7360 | 188 | - | PCPG | | ENSG00000203706 | ENST00000475406 | SNX5 | ENSG00000089006 | ENST00000377759 | -0.1329 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | SNX5 | ENSG00000089006 | ENST00000377768 | -0.1329 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | SNX5 | ENSG00000089006 | ENST00000606557 | -0.1381 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | SNX5 | ENSG00000089006 | ENST00000606602 | -0.1381 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | SNX5 | ENSG00000089006 | ENST00000481323 | -0.1275 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | SNX5 | ENSG00000089006 | ENST00000486039 | -0.2377 | 21 | - | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000462637 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000466072 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000482086 | -0.1209 | 286 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | PCCB | ENSG00000114054 | ENST00000469217 | -0.1855 | 171 | UCEC | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000346169 | -0.1308 | 108 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000352767 | -0.1308 | 104 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000414031 | -0.1308 | 108 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000392537 | -0.1308 | 110 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000382330 | -0.2572 | 112 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000350481 | -0.1308 | 108 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000342981 | -0.1308 | 103 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000424196 | -0.1308 | 110 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000434061 | -0.1308 | 110 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000319274 | -0.1024 | 110 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000435046 | -0.1341 | 114 | - | PCPG | | ENSG00000203706 | ENST00000437764 | EIF4G1 | ENSG00000114867 | ENST00000422614 | -0.1123 | 222 | - | PCPG | | ENSG00000203706 | ENST00000437764 | NAA50 | ENSG00000121579 | ENST00000493900 | -0.1680 | 310 | - | PCPG | | ENSG00000203706 | ENST00000437764 | PRC1 | ENSG00000198901 | ENST00000555455 | -0.1819 | 153 | UCEC,COAD,STAD,READ | - | | ENSG00000203706 | ENST00000437764 | DNAJC11 | ENSG00000007923 | ENST00000451196 | -0.1144 | 38 | - | KIRP,PCPG | | ENSG00000203706 | ENST00000437764 | NADK | ENSG00000008130 | ENST00000498806 | -0.1373 | 267 | - | PCPG | | ENSG00000203706 | ENST00000437764 | NADK | ENSG00000008130 | ENST00000489538 | -0.1300 | 231 | - | PCPG | | ENSG00000203706 | ENST00000437764 | BZW1 | ENSG00000082153 | ENST00000452790 | -0.1587 | 353 | - | PCPG | | ENSG00000203706 | ENST00000437764 | SNX5 | ENSG00000089006 | ENST00000481323 | -0.1200 | 69 | - | PCPG | | ENSG00000203706 | ENST00000437764 | SNX5 | ENSG00000089006 | ENST00000486039 | -0.2132 | 224 | - | PCPG | | ENSG00000203706 | ENST00000480052 | EIF4G1 | ENSG00000114867 | ENST00000382330 | -0.1596 | 42 | - | PCPG | | ENSG00000203706 | ENST00000480052 | SNX5 | ENSG00000089006 | ENST00000486039 | -0.1076 | 40 | - | PCPG | | ENSG00000203706 | ENST00000475406 | MFSD9 | ENSG00000135953 | ENST00000428085 | -0.1205 | 6 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MFSD9 | ENSG00000135953 | ENST00000421966 | -0.1002 | 68 | UCEC | - | | ENSG00000203706 | ENST00000475406 | DNAJA3 | ENSG00000103423 | ENST00000570857 | -0.1013 | 1 | - | PCPG | | ENSG00000203706 | ENST00000475406 | RPN2 | ENSG00000118705 | ENST00000437329 | -0.1313 | 436 | UCEC,READ | - | | ENSG00000203706 | ENST00000475406 | RPN2 | ENSG00000118705 | ENST00000456400 | -0.4067 | 427 | UCEC,READ | - | | ENSG00000203706 | ENST00000475406 | TUBG1 | ENSG00000131462 | ENST00000251413 | -0.1832 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000262027 | -0.1974 | 41 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000547501 | -0.1113 | 59 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000551431 | -0.1335 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000549074 | -0.1410 | 34 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000552007 | -0.1124 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000548944 | -0.1425 | 35 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000552914 | -0.1097 | 88 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000547665 | -0.1293 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | MARS1 | ENSG00000166986 | ENST00000551172 | -0.1211 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | PGD | ENSG00000142657 | ENST00000477958 | -0.1066 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | PGD | ENSG00000142657 | ENST00000270776 | -0.1279 | 1 | UCEC | - | | ENSG00000203706 | ENST00000475406 | AHCY | ENSG00000101444 | ENST00000217426 | -0.1255 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000475406 | AHCY | ENSG00000101444 | ENST00000538132 | -0.1255 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | MFSD9 | ENSG00000135953 | ENST00000428085 | -0.1113 | 83 | UCEC | - | | ENSG00000203706 | ENST00000437764 | MFSD9 | ENSG00000135953 | ENST00000421966 | -0.1379 | 117 | UCEC | - | | ENSG00000203706 | ENST00000437764 | RPN2 | ENSG00000118705 | ENST00000437329 | -0.1313 | 571 | UCEC,READ | - | | ENSG00000203706 | ENST00000437764 | RPN2 | ENSG00000118705 | ENST00000456400 | -1.4900 | 35 | UCEC,READ | - | | ENSG00000203706 | ENST00000437764 | MARS1 | ENSG00000166986 | ENST00000262027 | -0.1500 | 263 | UCEC | - | | ENSG00000203706 | ENST00000437764 | MARS1 | ENSG00000166986 | ENST00000547501 | -0.1564 | 299 | UCEC | - | | ENSG00000203706 | ENST00000437764 | MARS1 | ENSG00000166986 | ENST00000551431 | -0.1298 | 138 | UCEC | - | | ENSG00000203706 | ENST00000437764 | MARS1 | ENSG00000166986 | ENST00000549074 | -0.1356 | 124 | UCEC | - | | ENSG00000203706 | ENST00000437764 | MARS1 | ENSG00000166986 | ENST00000552007 | -0.1347 | 294 | UCEC | - | | ENSG00000203706 | ENST00000437764 | MARS1 | ENSG00000166986 | ENST00000552914 | -0.1500 | 263 | UCEC | - | | ENSG00000203706 | ENST00000437764 | GARS1 | ENSG00000106105 | ENST00000389266 | -0.1060 | 354 | COAD,READ | - | | ENSG00000203706 | ENST00000437764 | PGD | ENSG00000142657 | ENST00000477958 | -0.1209 | 112 | UCEC | - | | ENSG00000203706 | ENST00000437764 | AHCY | ENSG00000101444 | ENST00000217426 | -0.1066 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000437764 | AHCY | ENSG00000101444 | ENST00000538132 | -0.1066 | 1 | UCEC,COAD,READ | - | | ENSG00000203706 | ENST00000480052 | MFSD9 | ENSG00000135953 | ENST00000421966 | -0.1036 | 1 | UCEC | - | | ENSG00000203706 | ENST00000480052 | RPN2 | ENSG00000118705 | ENST00000437329 | -0.1313 | 258 | UCEC,READ | - | | ENSG00000203706 | ENST00000480052 | RPN2 | ENSG00000118705 | ENST00000456400 | -1.4900 | 38 | UCEC,READ | - | | ENSG00000203706 | ENST00000480052 | MARS1 | ENSG00000166986 | ENST00000262027 | -0.1252 | 274 | UCEC | - | | ENSG00000203706 | ENST00000480052 | MARS1 | ENSG00000166986 | ENST00000548944 | -0.1138 | 268 | UCEC | - | | ENSG00000203706 | ENST00000480052 | MARS1 | ENSG00000166986 | ENST00000552914 | -0.1252 | 268 | UCEC | - |
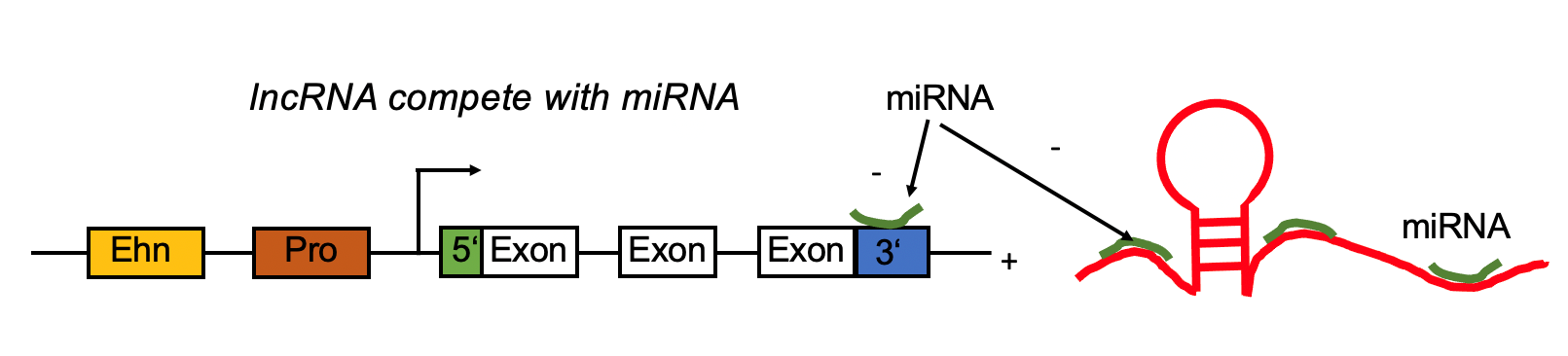
 lncRNA regulates mRNA by competing the miRNA binding site with mRNA. lncRNA regulates mRNA by competing the miRNA binding site with mRNA. |
| LncRNA Ensembl ID | lncRNA-miRNA-mRNA | Cancer Types | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,FYCO1 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,CFL2 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,FBXO32 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,RASL12 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,FAXDC2 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,MYLK | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,CSRP1 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,MCAM | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-7-1,HSPB8 | UCEC,TGCT,READ,GBM,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,CNN1 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,TCEAL4 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,ILK | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,RERG | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,TPM1 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,GPRASP2 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,SPARCL1 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,INPP5A | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,TCEAL3 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,PGM5 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,FAXDC2 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,KANK2 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,TAGLN | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,PLN | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,MYH11 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,KCNMB1 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,CAP2 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,LMOD1 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,SORBS1 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,HSPB6 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-142,CFL2 | UCEC,CESC,TGCT,READ,SARC | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,SYNPO2 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,FERMT2 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,KANK2 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,CALD1 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,MSRB3 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,DACT3 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,TGFB1I1 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,PDLIM7 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,PBX1 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,CAP2 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,PGM5 | UCEC,TGCT,READ,SARC,THYM | | ENSG00000203706 | SERTAD4-AS1,hsa-mir-185,TNS1 | UCEC,TGCT,READ,SARC,THYM |
 ORFfinder result for the gencode.v22.lncRNA.transcript.fa. ORFfinder result for the gencode.v22.lncRNA.transcript.fa. |
| lncRNA Ensembl ID | lncRNA ENST ID | length(AA) | start at transcript | end at transcript | | ENSG00000203706.7 | ENST00000475406.4 | 60 | 192 | 10 | | ENSG00000203706.7 | ENST00000437764.4 | 58 | 16 | 192 | | ENSG00000203706.7 | ENST00000480052.1 | 100 | 124 | 426 |
|

 Gene summary
Gene summary Differentially expressed gene analysis between cancer and normal samples.
Differentially expressed gene analysis between cancer and normal samples. Correlation between lncRNAs and cancer stages.
Correlation between lncRNAs and cancer stages.
 lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5).
lncRNA-epigenetic factor interaction (#cancer types with positive correlation >= 5). lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5).
lncRNA-transcript factor interaction (#cancer types with positive correlation >= 5). lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).
lncRNA-RNA binding protein interaction (#cancer types with positive correlation >= 5).