|
||||||
|
 | |
 | |
 | Differntial gene expression between RNA A-to-I edited tumor samples versus normal samples |
 | |
 | |
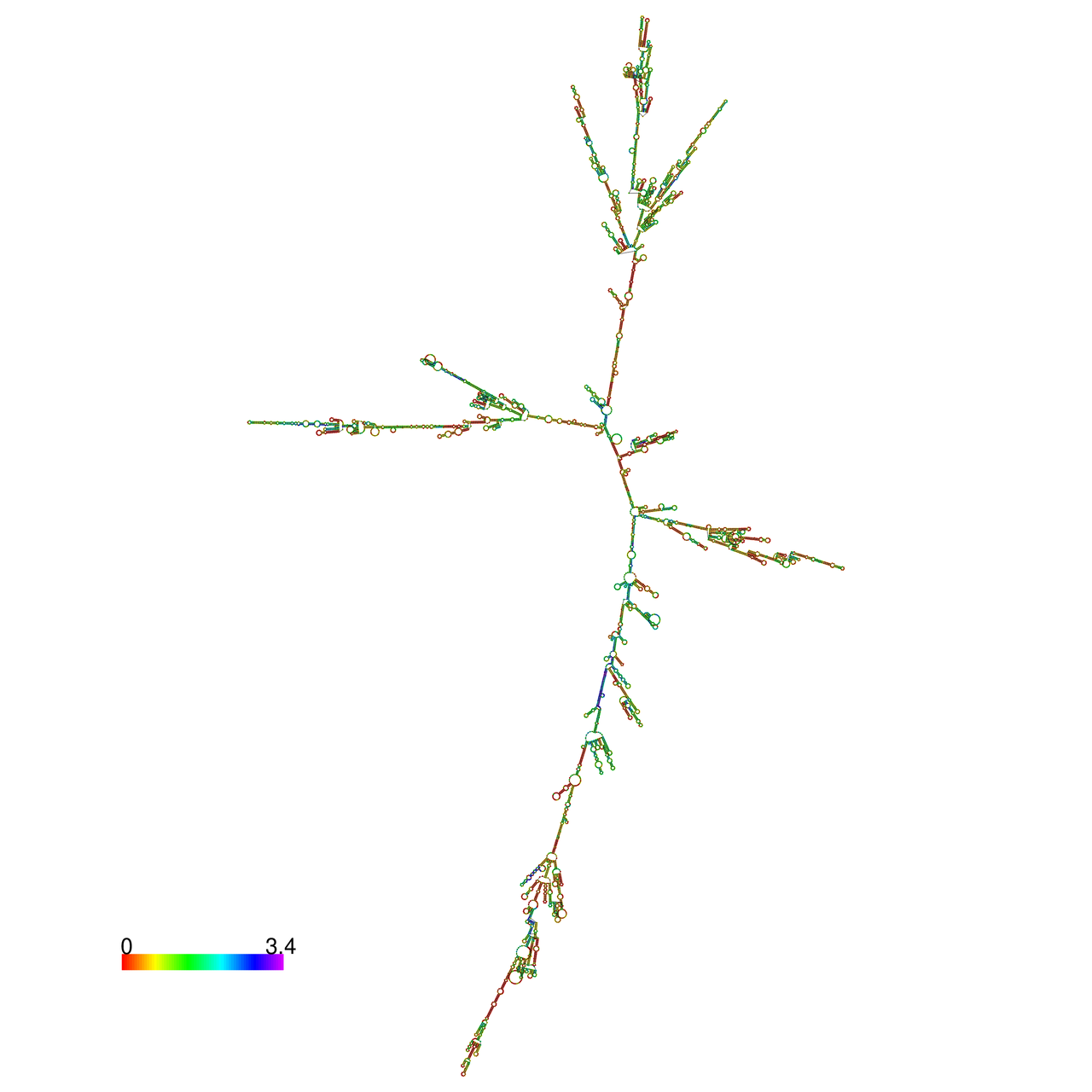
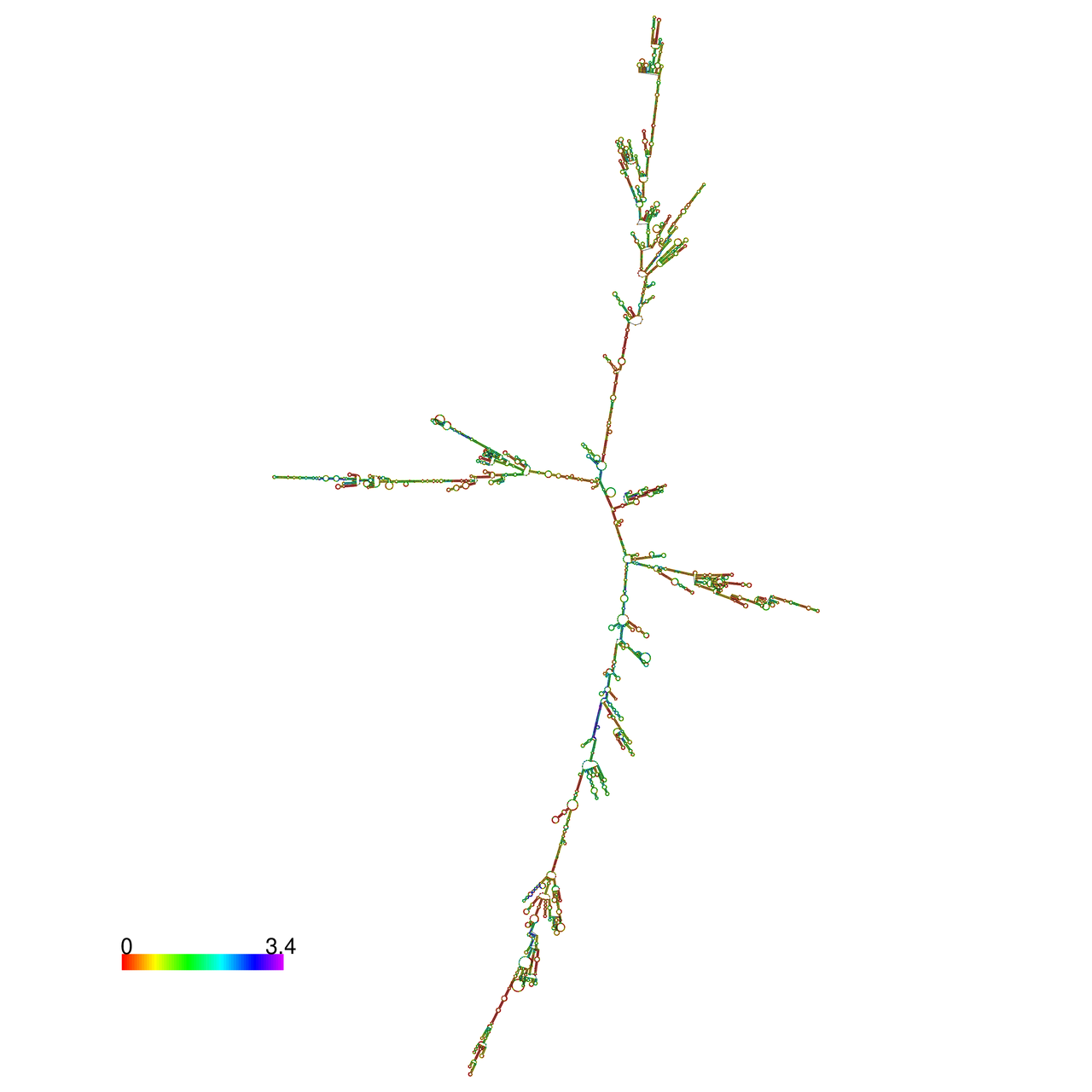
 | The effects of the RNA editing to the stability of the RNA structures |
 | |
 | |
 | |
 | |
 |
Editing gene: GIGYF2 (CAeditome ID:26058) |
Gene summary for GIGYF2 |
 Gene summary Gene summary |
| Gene information | Gene symbol | GIGYF2 | Gene ID | 26058 |
| Gene name | GRB10 interacting GYF protein 2 | |
| Synonyms | GYF2|PARK11|PERQ2|PERQ3|TNRC15 | |
| Cytomap | 2q37.1 | |
| Type of gene | protein-coding | |
| Description | GRB10-interacting GYF protein 2PERQ amino acid rich, with GYF domain 3PERQ amino acid-rich with GYF domain-containing protein 2Parkinson disease (autosomal recessive, early onset) 11trinucleotide repeat-containing gene 15 protein | |
| Modification date | 20210518 | |
| UniProtAcc | Q6Y7W6 | |
| Context |
 Gene ontology of each this gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each this gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Gene | GO ID | GO term | PubMed ID |
| GIGYF2 | GO:0070064 | GRB10-interacting GYF protein 2 | 20696395 |
| GIGYF2 | GO:0016441 | GRB10-interacting GYF protein 2 | 27157137 |
| GIGYF2 | GO:0061157 | GRB10-interacting GYF protein 2 | 27157137 |
| GIGYF2 | GO:0005768 | GRB10-interacting GYF protein 2 | 20670374 |
| GIGYF2 | GO:0005783 | GRB10-interacting GYF protein 2 | 20696395 |
| GIGYF2 | GO:0005794 | GRB10-interacting GYF protein 2 | 20696395 |
| GIGYF2 | GO:0005829 | GRB10-interacting GYF protein 2 | 0000052 |
| GIGYF2 | GO:0010494 | GRB10-interacting GYF protein 2 | 20696395 |
| GIGYF2 | GO:0043204 | GRB10-interacting GYF protein 2 | 20670374 |
| GIGYF2 | GO:1990635 | GRB10-interacting GYF protein 2 | 20670374 |
| GIGYF2 | GO:0032991 | GRB10-interacting GYF protein 2 | 20878056 |
Top |
RNA A-to-I events for GIGYF2 |
 RNA A-to-I editing events across TCGA 33 cancers data sets based on Ensembl gene isoform structure. RNA A-to-I editing events across TCGA 33 cancers data sets based on Ensembl gene isoform structure. |
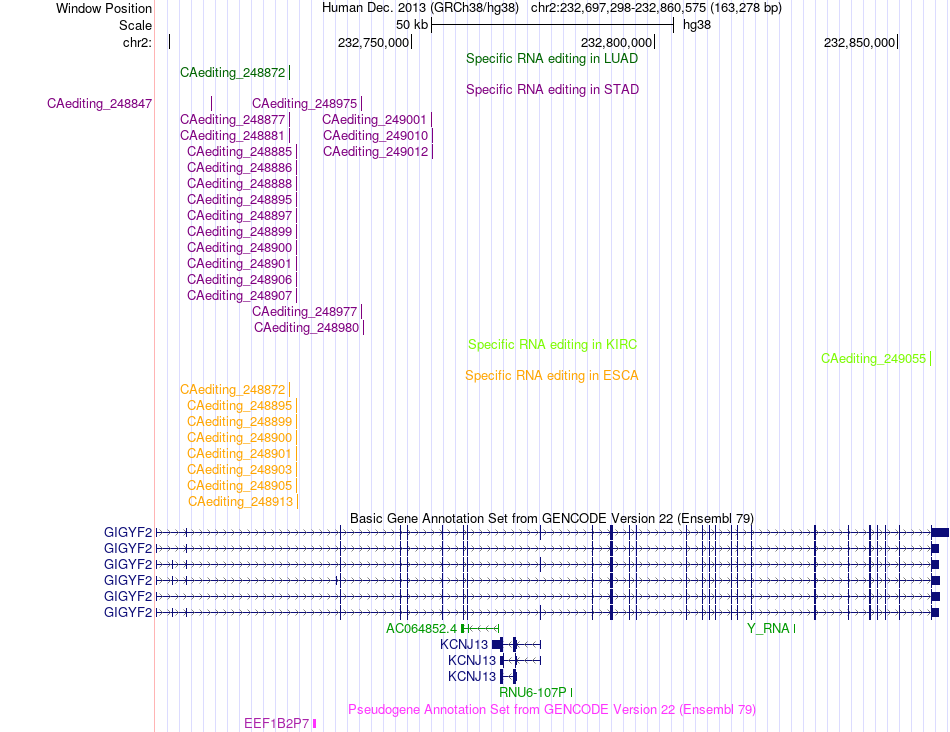 |
 RNA editing frequencies across 33 cancers of gene: ENSG00000204120.13 RNA editing frequencies across 33 cancers of gene: ENSG00000204120.13 |
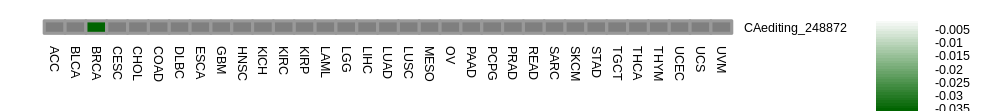 |
 RNA A-to-I editing events in Cancer. RNA A-to-I editing events in Cancer. |
| CAediting_248800(232701493, +), CAediting_248801(232701497, +), CAediting_248802(232701555, +), CAediting_248803(232701621, +), CAediting_248804(232701658, +), CAediting_248805(232701659, +), CAediting_248806(232701685, +), CAediting_248807(232701698, +), CAediting_248808(232701700, +), CAediting_248809(232701899, +), CAediting_248810(232701929, +), CAediting_248811(232701945, +), CAediting_248812(232702345, +), CAediting_248813(232703047, +), CAediting_248814(232703054, +), CAediting_248815(232704474, +), CAediting_248816(232704479, +), CAediting_248817(232704534, +), CAediting_248818(232704544, +), CAediting_248819(232704577, +), CAediting_248820(232704580, +), CAediting_248821(232704583, +), CAediting_248822(232704659, +), CAediting_248823(232705535, +), CAediting_248824(232705927, +), CAediting_248825(232705952, +), CAediting_248826(232706631, +), CAediting_248827(232706657, +), CAediting_248828(232706700, +), CAediting_248829(232706701, +), CAediting_248830(232706721, +), CAediting_248831(232706767, +), CAediting_248832(232706813, +), CAediting_248833(232706826, +), CAediting_248834(232706827, +), CAediting_248835(232706865, +), CAediting_248836(232706891, +), CAediting_248837(232708094, +), CAediting_248838(232708160, +), CAediting_248839(232708228, +), CAediting_248840(232708558, +), CAediting_248841(232708564, +), CAediting_248842(232708580, +), CAediting_248843(232708581, +), CAediting_248844(232708586, +), CAediting_248845(232708595, +), CAediting_248846(232708596, +), CAediting_248847(232708606, +), CAediting_248848(232708636, +), CAediting_248849(232708867, +), CAediting_248850(232708884, +), CAediting_248851(232708912, +), CAediting_248852(232708913, +), CAediting_248853(232708956, +), CAediting_248854(232708974, +), CAediting_248855(232709363, +), CAediting_248856(232709711, +), CAediting_248857(232710029, +), CAediting_248858(232710166, +), CAediting_248859(232710221, +), CAediting_248860(232710233, +), CAediting_248861(232710294, +), CAediting_248862(232710317, +), CAediting_248863(232710323, +), CAediting_248864(232710324, +), CAediting_248865(232713176, +), CAediting_248866(232717178, +), CAediting_248867(232717251, +), CAediting_248868(232718119, +), CAediting_248869(232718237, +), CAediting_248870(232724576, +), CAediting_248871(232724592, +), CAediting_248872(232724601, +), CAediting_248873(232724643, +), CAediting_248874(232724649, +), CAediting_248875(232724650, +), CAediting_248876(232724651, +), CAediting_248877(232724662, +), CAediting_248878(232724667, +), CAediting_248879(232724675, +), CAediting_248880(232724676, +), CAediting_248881(232724679, +), CAediting_248882(232724690, +), CAediting_248883(232724805, +), CAediting_248884(232724815, +), CAediting_248885(232726063, +), CAediting_248886(232726072, +), CAediting_248887(232726116, +), CAediting_248888(232726132, +), CAediting_248889(232726137, +), CAediting_248890(232726150, +), CAediting_248891(232726160, +), CAediting_248892(232726163, +), CAediting_248893(232726183, +), CAediting_248894(232726184, +), CAediting_248895(232726186, +), CAediting_248896(232726190, +), CAediting_248897(232726193, +), CAediting_248898(232726204, +), CAediting_248899(232726208, +), CAediting_248900(232726209, +), CAediting_248901(232726210, +), CAediting_248902(232726211, +), CAediting_248903(232726213, +), CAediting_248904(232726215, +), CAediting_248905(232726216, +), CAediting_248906(232726217, +), CAediting_248907(232726218, +), CAediting_248908(232726219, +), CAediting_248909(232726248, +), CAediting_248910(232726254, +), CAediting_248911(232726258, +), CAediting_248912(232726275, +), CAediting_248913(232726277, +), CAediting_248914(232726278, +), CAediting_248915(232726289, +), CAediting_248916(232726290, +), CAediting_248917(232726297, +), CAediting_248918(232726326, +), CAediting_248919(232726485, +), CAediting_248920(232726495, +), CAediting_248921(232727064, +), CAediting_248922(232727075, +), CAediting_248923(232727082, +), CAediting_248924(232727094, +), CAediting_248925(232727095, +), CAediting_248926(232727104, +), CAediting_248927(232727126, +), CAediting_248928(232727131, +), CAediting_248929(232727146, +), CAediting_248930(232727168, +), CAediting_248931(232730484, +), CAediting_248932(232730485, +), CAediting_248933(232730511, +), CAediting_248934(232730520, +), CAediting_248935(232730526, +), CAediting_248936(232730554, +), CAediting_248937(232732665, +), CAediting_248938(232732704, +), CAediting_248939(232732708, +), CAediting_248940(232732768, +), CAediting_248941(232732790, +), CAediting_248942(232732794, +), CAediting_248943(232732830, +), CAediting_248944(232733001, +), CAediting_248945(232733002, +), CAediting_248946(232733045, +), CAediting_248947(232733054, +), CAediting_248948(232733079, +), CAediting_248949(232733126, +), CAediting_248950(232733136, +), CAediting_248951(232733163, +), CAediting_248952(232733202, +), CAediting_248953(232733209, +), CAediting_248954(232734470, +), CAediting_248955(232734471, +), CAediting_248956(232734479, +), CAediting_248957(232738016, +), CAediting_248958(232738056, +), CAediting_248959(232738114, +), CAediting_248960(232739144, +), CAediting_248961(232739198, +), CAediting_248962(232739199, +), CAediting_248963(232739200, +), CAediting_248964(232739201, +), CAediting_248965(232739203, +), CAediting_248966(232739207, +), CAediting_248967(232739237, +), CAediting_248968(232739238, +), CAediting_248969(232739251, +), CAediting_248970(232739264, +), CAediting_248971(232739283, +), CAediting_248972(232739293, +), CAediting_248973(232739529, +), CAediting_248974(232739582, +), CAediting_248975(232739586, +), CAediting_248976(232739588, +), CAediting_248977(232739594, +), CAediting_248978(232739891, +), CAediting_248979(232739967, +), CAediting_248980(232739998, +), CAediting_248981(232740007, +), CAediting_248982(232740024, +), CAediting_248983(232740039, +), CAediting_248984(232741624, +), CAediting_248985(232742291, +), CAediting_248986(232742382, +), CAediting_248987(232742383, +), CAediting_248988(232742385, +), CAediting_248989(232742398, +), CAediting_248990(232743973, +), CAediting_248991(232744671, +), CAediting_248992(232744808, +), CAediting_248993(232750883, +), CAediting_248994(232750957, +), CAediting_248995(232753223, +), CAediting_248996(232753224, +), CAediting_248997(232753892, +), CAediting_248998(232753904, +), CAediting_248999(232753950, +), CAediting_249000(232753993, +), CAediting_249001(232753999, +), CAediting_249002(232754002, +), CAediting_249003(232754004, +), CAediting_249004(232754005, +), CAediting_249005(232754008, +), CAediting_249006(232754018, +), CAediting_249007(232754026, +), CAediting_249008(232754044, +), CAediting_249009(232754054, +), CAediting_249010(232754058, +), CAediting_249011(232754086, +), CAediting_249012(232754110, +), CAediting_249013(232754153, +), CAediting_249014(232782244, +), CAediting_249015(232782272, +), CAediting_249016(232784235, +), CAediting_249017(232784274, +), CAediting_249018(232797727, +), CAediting_249019(232801004, +), CAediting_249020(232807438, +), CAediting_249021(232807448, +), CAediting_249022(232809102, +), CAediting_249023(232812858, +), CAediting_249024(232812880, +), CAediting_249025(232812897, +), CAediting_249026(232812916, +), CAediting_249027(232812926, +), CAediting_249028(232812942, +), CAediting_249029(232812962, +), CAediting_249030(232814154, +), CAediting_249031(232820480, +), CAediting_249032(232823457, +), CAediting_249033(232828484, +), CAediting_249034(232828552, +), CAediting_249035(232828553, +), CAediting_249036(232828582, +), CAediting_249037(232828588, +), CAediting_249038(232828595, +), CAediting_249039(232830011, +), CAediting_249040(232830126, +), CAediting_249041(232830167, +), CAediting_249042(232830203, +), CAediting_249043(232836459, +), CAediting_249044(232836481, +), CAediting_249045(232848424, +), CAediting_249046(232848511, +), CAediting_249047(232851092, +), CAediting_249048(232851178, +), CAediting_249049(232851196, +), CAediting_249050(232854243, +), CAediting_249051(232855187, +), CAediting_249052(232855204, +), CAediting_249053(232856412, +), CAediting_249054(232856451, +), CAediting_249055(232856573, +), CAediting_249056(232856647, +), CAediting_249057(232860206, +), CAediting_249058(232860232, +), |
Top |
RNA editing positional annotations for GIGYF2 using Annovar |
 Protein coding RNA editing(s). Protein coding RNA editing(s). |
| CAeditome ID | Position | Variant type | ENST | NTchange | AAchange | SIFT_score | Polyphen2_HVAR_score | PROVEAN_pred |
 Gene structure information of RNA editing(s). Gene structure information of RNA editing(s). |
| CAeditome ID | Position | Variant type |
| CAediting_248800 | chr2_232701493_+ | ncRNA_intronic |
| CAediting_248801 | chr2_232701497_+ | ncRNA_intronic |
| CAediting_248802 | chr2_232701555_+ | ncRNA_intronic |
| CAediting_248803 | chr2_232701621_+ | ncRNA_intronic |
| CAediting_248804 | chr2_232701658_+ | ncRNA_intronic |
| CAediting_248805 | chr2_232701659_+ | ncRNA_intronic |
| CAediting_248806 | chr2_232701685_+ | ncRNA_intronic |
| CAediting_248807 | chr2_232701698_+ | ncRNA_intronic |
| CAediting_248808 | chr2_232701700_+ | ncRNA_intronic |
| CAediting_248809 | chr2_232701899_+ | ncRNA_intronic |
| CAediting_248810 | chr2_232701929_+ | ncRNA_intronic |
| CAediting_248811 | chr2_232701945_+ | ncRNA_intronic |
| CAediting_248812 | chr2_232702345_+ | ncRNA_intronic |
| CAediting_248813 | chr2_232703047_+ | ncRNA_intronic |
| CAediting_248814 | chr2_232703054_+ | ncRNA_intronic |
| CAediting_248815 | chr2_232704474_+ | ncRNA_intronic |
| CAediting_248816 | chr2_232704479_+ | ncRNA_intronic |
| CAediting_248817 | chr2_232704534_+ | ncRNA_intronic |
| CAediting_248818 | chr2_232704544_+ | ncRNA_intronic |
| CAediting_248819 | chr2_232704577_+ | ncRNA_intronic |
| CAediting_248820 | chr2_232704580_+ | ncRNA_intronic |
| CAediting_248821 | chr2_232704583_+ | ncRNA_intronic |
| CAediting_248822 | chr2_232704659_+ | ncRNA_intronic |
| CAediting_248823 | chr2_232705535_+ | ncRNA_exonic |
| CAediting_248824 | chr2_232705927_+ | ncRNA_exonic |
| CAediting_248825 | chr2_232705952_+ | ncRNA_intronic |
| CAediting_248826 | chr2_232706631_+ | ncRNA_intronic |
| CAediting_248827 | chr2_232706657_+ | ncRNA_intronic |
| CAediting_248828 | chr2_232706700_+ | ncRNA_intronic |
| CAediting_248829 | chr2_232706701_+ | ncRNA_intronic |
| CAediting_248830 | chr2_232706721_+ | ncRNA_intronic |
| CAediting_248831 | chr2_232706767_+ | ncRNA_intronic |
| CAediting_248832 | chr2_232706813_+ | ncRNA_intronic |
| CAediting_248833 | chr2_232706826_+ | ncRNA_intronic |
| CAediting_248834 | chr2_232706827_+ | ncRNA_intronic |
| CAediting_248835 | chr2_232706865_+ | ncRNA_intronic |
| CAediting_248836 | chr2_232706891_+ | ncRNA_intronic |
| CAediting_248837 | chr2_232708094_+ | ncRNA_intronic |
| CAediting_248838 | chr2_232708160_+ | ncRNA_intronic |
| CAediting_248839 | chr2_232708228_+ | ncRNA_intronic |
| CAediting_248840 | chr2_232708558_+ | ncRNA_intronic |
| CAediting_248841 | chr2_232708564_+ | ncRNA_intronic |
| CAediting_248842 | chr2_232708580_+ | ncRNA_intronic |
| CAediting_248843 | chr2_232708581_+ | ncRNA_intronic |
| CAediting_248844 | chr2_232708586_+ | ncRNA_intronic |
| CAediting_248845 | chr2_232708595_+ | ncRNA_intronic |
| CAediting_248846 | chr2_232708596_+ | ncRNA_intronic |
| CAediting_248847 | chr2_232708606_+ | ncRNA_intronic |
| CAediting_248848 | chr2_232708636_+ | ncRNA_intronic |
| CAediting_248849 | chr2_232708867_+ | ncRNA_intronic |
| CAediting_248850 | chr2_232708884_+ | ncRNA_intronic |
| CAediting_248851 | chr2_232708912_+ | ncRNA_intronic |
| CAediting_248852 | chr2_232708913_+ | ncRNA_intronic |
| CAediting_248853 | chr2_232708956_+ | ncRNA_intronic |
| CAediting_248854 | chr2_232708974_+ | ncRNA_intronic |
| CAediting_248855 | chr2_232709363_+ | ncRNA_intronic |
| CAediting_248856 | chr2_232709711_+ | ncRNA_intronic |
| CAediting_248857 | chr2_232710029_+ | ncRNA_intronic |
| CAediting_248858 | chr2_232710166_+ | ncRNA_intronic |
| CAediting_248859 | chr2_232710221_+ | ncRNA_intronic |
| CAediting_248860 | chr2_232710233_+ | ncRNA_intronic |
| CAediting_248861 | chr2_232710294_+ | ncRNA_intronic |
| CAediting_248862 | chr2_232710317_+ | ncRNA_intronic |
| CAediting_248863 | chr2_232710323_+ | ncRNA_intronic |
| CAediting_248864 | chr2_232710324_+ | ncRNA_intronic |
| CAediting_248865 | chr2_232713176_+ | ncRNA_intronic |
| CAediting_248866 | chr2_232717178_+ | ncRNA_intronic |
| CAediting_248867 | chr2_232717251_+ | ncRNA_intronic |
| CAediting_248868 | chr2_232718119_+ | ncRNA_intronic |
| CAediting_248869 | chr2_232718237_+ | ncRNA_intronic |
| CAediting_248870 | chr2_232724576_+ | ncRNA_exonic |
| CAediting_248871 | chr2_232724592_+ | ncRNA_exonic |
| CAediting_248872 | chr2_232724601_+ | ncRNA_exonic |
| CAediting_248873 | chr2_232724643_+ | ncRNA_exonic |
| CAediting_248874 | chr2_232724649_+ | ncRNA_exonic |
| CAediting_248875 | chr2_232724650_+ | ncRNA_exonic |
| CAediting_248876 | chr2_232724651_+ | ncRNA_exonic |
| CAediting_248877 | chr2_232724662_+ | ncRNA_exonic |
| CAediting_248878 | chr2_232724667_+ | ncRNA_exonic |
| CAediting_248879 | chr2_232724675_+ | ncRNA_exonic |
| CAediting_248880 | chr2_232724676_+ | ncRNA_exonic |
| CAediting_248881 | chr2_232724679_+ | ncRNA_exonic |
| CAediting_248882 | chr2_232724690_+ | ncRNA_exonic |
| CAediting_248883 | chr2_232724805_+ | ncRNA_intronic |
| CAediting_248884 | chr2_232724815_+ | ncRNA_intronic |
| CAediting_248885 | chr2_232726063_+ | ncRNA_intronic |
| CAediting_248886 | chr2_232726072_+ | ncRNA_intronic |
| CAediting_248887 | chr2_232726116_+ | ncRNA_intronic |
| CAediting_248888 | chr2_232726132_+ | ncRNA_intronic |
| CAediting_248889 | chr2_232726137_+ | ncRNA_intronic |
| CAediting_248890 | chr2_232726150_+ | ncRNA_intronic |
| CAediting_248891 | chr2_232726160_+ | ncRNA_intronic |
| CAediting_248892 | chr2_232726163_+ | ncRNA_intronic |
| CAediting_248893 | chr2_232726183_+ | ncRNA_intronic |
| CAediting_248894 | chr2_232726184_+ | ncRNA_intronic |
| CAediting_248895 | chr2_232726186_+ | ncRNA_intronic |
| CAediting_248896 | chr2_232726190_+ | ncRNA_intronic |
| CAediting_248897 | chr2_232726193_+ | ncRNA_intronic |
| CAediting_248898 | chr2_232726204_+ | ncRNA_intronic |
| CAediting_248899 | chr2_232726208_+ | ncRNA_intronic |
| CAediting_248900 | chr2_232726209_+ | ncRNA_intronic |
| CAediting_248901 | chr2_232726210_+ | ncRNA_intronic |
| CAediting_248902 | chr2_232726211_+ | ncRNA_intronic |
| CAediting_248903 | chr2_232726213_+ | ncRNA_intronic |
| CAediting_248904 | chr2_232726215_+ | ncRNA_intronic |
| CAediting_248905 | chr2_232726216_+ | ncRNA_intronic |
| CAediting_248906 | chr2_232726217_+ | ncRNA_intronic |
| CAediting_248907 | chr2_232726218_+ | ncRNA_intronic |
| CAediting_248908 | chr2_232726219_+ | ncRNA_intronic |
| CAediting_248909 | chr2_232726248_+ | ncRNA_intronic |
| CAediting_248910 | chr2_232726254_+ | ncRNA_intronic |
| CAediting_248911 | chr2_232726258_+ | ncRNA_intronic |
| CAediting_248912 | chr2_232726275_+ | ncRNA_intronic |
| CAediting_248913 | chr2_232726277_+ | ncRNA_intronic |
| CAediting_248914 | chr2_232726278_+ | ncRNA_intronic |
| CAediting_248915 | chr2_232726289_+ | ncRNA_intronic |
| CAediting_248916 | chr2_232726290_+ | ncRNA_intronic |
| CAediting_248917 | chr2_232726297_+ | ncRNA_intronic |
| CAediting_248918 | chr2_232726326_+ | ncRNA_intronic |
| CAediting_248919 | chr2_232726485_+ | ncRNA_intronic |
| CAediting_248920 | chr2_232726495_+ | ncRNA_intronic |
| CAediting_248921 | chr2_232727064_+ | ncRNA_intronic |
| CAediting_248922 | chr2_232727075_+ | ncRNA_intronic |
| CAediting_248923 | chr2_232727082_+ | ncRNA_intronic |
| CAediting_248924 | chr2_232727094_+ | ncRNA_intronic |
| CAediting_248925 | chr2_232727095_+ | ncRNA_intronic |
| CAediting_248926 | chr2_232727104_+ | ncRNA_intronic |
| CAediting_248927 | chr2_232727126_+ | ncRNA_intronic |
| CAediting_248928 | chr2_232727131_+ | ncRNA_intronic |
| CAediting_248929 | chr2_232727146_+ | ncRNA_intronic |
| CAediting_248930 | chr2_232727168_+ | ncRNA_intronic |
| CAediting_248931 | chr2_232730484_+ | ncRNA_intronic |
| CAediting_248932 | chr2_232730485_+ | ncRNA_intronic |
| CAediting_248933 | chr2_232730511_+ | ncRNA_intronic |
| CAediting_248934 | chr2_232730520_+ | ncRNA_intronic |
| CAediting_248935 | chr2_232730526_+ | ncRNA_intronic |
| CAediting_248936 | chr2_232730554_+ | ncRNA_intronic |
| CAediting_248937 | chr2_232732665_+ | ncRNA_intronic |
| CAediting_248938 | chr2_232732704_+ | ncRNA_intronic |
| CAediting_248939 | chr2_232732708_+ | ncRNA_intronic |
| CAediting_248940 | chr2_232732768_+ | ncRNA_intronic |
| CAediting_248941 | chr2_232732790_+ | ncRNA_intronic |
| CAediting_248942 | chr2_232732794_+ | ncRNA_intronic |
| CAediting_248943 | chr2_232732830_+ | ncRNA_intronic |
| CAediting_248944 | chr2_232733001_+ | ncRNA_intronic |
| CAediting_248945 | chr2_232733002_+ | ncRNA_intronic |
| CAediting_248946 | chr2_232733045_+ | ncRNA_intronic |
| CAediting_248947 | chr2_232733054_+ | ncRNA_intronic |
| CAediting_248948 | chr2_232733079_+ | ncRNA_intronic |
| CAediting_248949 | chr2_232733126_+ | ncRNA_intronic |
| CAediting_248950 | chr2_232733136_+ | ncRNA_intronic |
| CAediting_248951 | chr2_232733163_+ | ncRNA_intronic |
| CAediting_248952 | chr2_232733202_+ | ncRNA_intronic |
| CAediting_248953 | chr2_232733209_+ | ncRNA_intronic |
| CAediting_248954 | chr2_232734470_+ | ncRNA_exonic |
| CAediting_248955 | chr2_232734471_+ | ncRNA_exonic |
| CAediting_248956 | chr2_232734479_+ | ncRNA_exonic |
| CAediting_248957 | chr2_232738016_+ | ncRNA_intronic |
| CAediting_248958 | chr2_232738056_+ | ncRNA_intronic |
| CAediting_248959 | chr2_232738114_+ | ncRNA_intronic |
| CAediting_248960 | chr2_232739144_+ | ncRNA_intronic |
| CAediting_248961 | chr2_232739198_+ | ncRNA_intronic |
| CAediting_248962 | chr2_232739199_+ | ncRNA_intronic |
| CAediting_248963 | chr2_232739200_+ | ncRNA_intronic |
| CAediting_248964 | chr2_232739201_+ | ncRNA_intronic |
| CAediting_248965 | chr2_232739203_+ | ncRNA_intronic |
| CAediting_248966 | chr2_232739207_+ | ncRNA_intronic |
| CAediting_248967 | chr2_232739237_+ | ncRNA_intronic |
| CAediting_248968 | chr2_232739238_+ | ncRNA_intronic |
| CAediting_248969 | chr2_232739251_+ | ncRNA_intronic |
| CAediting_248970 | chr2_232739264_+ | ncRNA_intronic |
| CAediting_248971 | chr2_232739283_+ | ncRNA_intronic |
| CAediting_248972 | chr2_232739293_+ | ncRNA_intronic |
| CAediting_248973 | chr2_232739529_+ | ncRNA_intronic |
| CAediting_248974 | chr2_232739582_+ | ncRNA_intronic |
| CAediting_248975 | chr2_232739586_+ | ncRNA_intronic |
| CAediting_248976 | chr2_232739588_+ | ncRNA_intronic |
| CAediting_248977 | chr2_232739594_+ | ncRNA_intronic |
| CAediting_248978 | chr2_232739891_+ | ncRNA_intronic |
| CAediting_248979 | chr2_232739967_+ | ncRNA_intronic |
| CAediting_248980 | chr2_232739998_+ | ncRNA_intronic |
| CAediting_248981 | chr2_232740007_+ | ncRNA_intronic |
| CAediting_248982 | chr2_232740024_+ | ncRNA_intronic |
| CAediting_248983 | chr2_232740039_+ | ncRNA_intronic |
| CAediting_248984 | chr2_232741624_+ | ncRNA_intronic |
| CAediting_248985 | chr2_232742291_+ | ncRNA_intronic |
| CAediting_248986 | chr2_232742382_+ | ncRNA_intronic |
| CAediting_248987 | chr2_232742383_+ | ncRNA_intronic |
| CAediting_248988 | chr2_232742385_+ | ncRNA_intronic |
| CAediting_248989 | chr2_232742398_+ | ncRNA_intronic |
| CAediting_248990 | chr2_232743973_+ | ncRNA_intronic |
| CAediting_248991 | chr2_232744671_+ | ncRNA_intronic |
| CAediting_248992 | chr2_232744808_+ | ncRNA_intronic |
| CAediting_248993 | chr2_232750883_+ | ncRNA_intronic |
| CAediting_248994 | chr2_232750957_+ | ncRNA_intronic |
| CAediting_248995 | chr2_232753223_+ | ncRNA_intronic |
| CAediting_248996 | chr2_232753224_+ | ncRNA_intronic |
| CAediting_248997 | chr2_232753892_+ | ncRNA_intronic |
| CAediting_248998 | chr2_232753904_+ | ncRNA_intronic |
| CAediting_248999 | chr2_232753950_+ | ncRNA_intronic |
| CAediting_249000 | chr2_232753993_+ | ncRNA_intronic |
| CAediting_249001 | chr2_232753999_+ | ncRNA_intronic |
| CAediting_249002 | chr2_232754002_+ | ncRNA_intronic |
| CAediting_249003 | chr2_232754004_+ | ncRNA_intronic |
| CAediting_249004 | chr2_232754005_+ | ncRNA_intronic |
| CAediting_249005 | chr2_232754008_+ | ncRNA_intronic |
| CAediting_249006 | chr2_232754018_+ | ncRNA_intronic |
| CAediting_249007 | chr2_232754026_+ | ncRNA_intronic |
| CAediting_249008 | chr2_232754044_+ | ncRNA_intronic |
| CAediting_249009 | chr2_232754054_+ | ncRNA_intronic |
| CAediting_249010 | chr2_232754058_+ | ncRNA_intronic |
| CAediting_249011 | chr2_232754086_+ | ncRNA_intronic |
| CAediting_249012 | chr2_232754110_+ | ncRNA_intronic |
| CAediting_249013 | chr2_232754153_+ | ncRNA_intronic |
| CAediting_249014 | chr2_232782244_+ | intronic |
| CAediting_249015 | chr2_232782272_+ | intronic |
| CAediting_249016 | chr2_232784235_+ | intronic |
| CAediting_249017 | chr2_232784274_+ | intronic |
| CAediting_249018 | chr2_232797727_+ | intronic |
| CAediting_249019 | chr2_232801004_+ | intronic |
| CAediting_249020 | chr2_232807438_+ | ncRNA_intronic |
| CAediting_249021 | chr2_232807448_+ | ncRNA_intronic |
| CAediting_249022 | chr2_232809102_+ | ncRNA_intronic |
| CAediting_249023 | chr2_232812858_+ | ncRNA_intronic |
| CAediting_249024 | chr2_232812880_+ | ncRNA_intronic |
| CAediting_249025 | chr2_232812897_+ | ncRNA_intronic |
| CAediting_249026 | chr2_232812916_+ | ncRNA_intronic |
| CAediting_249027 | chr2_232812926_+ | ncRNA_intronic |
| CAediting_249028 | chr2_232812942_+ | ncRNA_intronic |
| CAediting_249029 | chr2_232812962_+ | ncRNA_intronic |
| CAediting_249030 | chr2_232814154_+ | ncRNA_intronic |
| CAediting_249031 | chr2_232820480_+ | ncRNA_intronic |
| CAediting_249032 | chr2_232823457_+ | ncRNA_intronic |
| CAediting_249033 | chr2_232828484_+ | ncRNA_intronic |
| CAediting_249034 | chr2_232828552_+ | ncRNA_intronic |
| CAediting_249035 | chr2_232828553_+ | ncRNA_intronic |
| CAediting_249036 | chr2_232828582_+ | ncRNA_intronic |
| CAediting_249037 | chr2_232828588_+ | ncRNA_intronic |
| CAediting_249038 | chr2_232828595_+ | ncRNA_intronic |
| CAediting_249039 | chr2_232830011_+ | ncRNA_intronic |
| CAediting_249040 | chr2_232830126_+ | ncRNA_intronic |
| CAediting_249041 | chr2_232830167_+ | ncRNA_intronic |
| CAediting_249042 | chr2_232830203_+ | ncRNA_intronic |
| CAediting_249043 | chr2_232836459_+ | ncRNA_intronic |
| CAediting_249044 | chr2_232836481_+ | ncRNA_intronic |
| CAediting_249045 | chr2_232848424_+ | ncRNA_intronic |
| CAediting_249046 | chr2_232848511_+ | ncRNA_intronic |
| CAediting_249047 | chr2_232851092_+ | ncRNA_intronic |
| CAediting_249048 | chr2_232851178_+ | ncRNA_intronic |
| CAediting_249049 | chr2_232851196_+ | ncRNA_intronic |
| CAediting_249050 | chr2_232854243_+ | ncRNA_intronic |
| CAediting_249051 | chr2_232855187_+ | ncRNA_intronic |
| CAediting_249052 | chr2_232855204_+ | ncRNA_intronic |
| CAediting_249053 | chr2_232856412_+ | ncRNA_intronic |
| CAediting_249054 | chr2_232856451_+ | ncRNA_intronic |
| CAediting_249055 | chr2_232856573_+ | ncRNA_intronic |
| CAediting_249056 | chr2_232856647_+ | ncRNA_intronic |
| CAediting_249057 | chr2_232860206_+ | UTR3 |
| CAediting_249058 | chr2_232860232_+ | UTR3 |
 Repat-regional RNA editing(s). Repat-regional RNA editing(s). |
| CAeditome ID | Position | Repeat family | Repeat sub family | Repeat name |
| CAediting_248800 | chr2_232701493_+ | SINE | Alu | AluJb |
| CAediting_248801 | chr2_232701497_+ | SINE | Alu | AluJb |
| CAediting_248802 | chr2_232701555_+ | SINE | Alu | AluJb |
| CAediting_248803 | chr2_232701621_+ | SINE | Alu | AluJb |
| CAediting_248804 | chr2_232701658_+ | SINE | Alu | AluJb |
| CAediting_248805 | chr2_232701659_+ | SINE | Alu | AluJb |
| CAediting_248806 | chr2_232701685_+ | SINE | Alu | AluJb |
| CAediting_248807 | chr2_232701698_+ | SINE | Alu | AluJb |
| CAediting_248808 | chr2_232701700_+ | SINE | Alu | AluJb |
| CAediting_248809 | chr2_232701899_+ | SINE | Alu | AluJr |
| CAediting_248810 | chr2_232701929_+ | SINE | Alu | AluJr |
| CAediting_248811 | chr2_232701945_+ | SINE | Alu | AluJr |
| CAediting_248812 | chr2_232702345_+ | SINE | Alu | AluSz6 |
| CAediting_248813 | chr2_232703047_+ | SINE | Alu | AluJb |
| CAediting_248814 | chr2_232703054_+ | SINE | Alu | AluJb |
| CAediting_248815 | chr2_232704474_+ | SINE | Alu | AluSz6 |
| CAediting_248816 | chr2_232704479_+ | SINE | Alu | AluSz6 |
| CAediting_248817 | chr2_232704534_+ | SINE | Alu | AluSz6 |
| CAediting_248818 | chr2_232704544_+ | SINE | Alu | AluSz6 |
| CAediting_248819 | chr2_232704577_+ | SINE | Alu | AluSz6 |
| CAediting_248820 | chr2_232704580_+ | SINE | Alu | AluSz6 |
| CAediting_248821 | chr2_232704583_+ | SINE | Alu | AluSz6 |
| CAediting_248822 | chr2_232704659_+ | SINE | Alu | AluSz6 |
| CAediting_248823 | chr2_232705535_+ | SINE | Alu | AluSp |
| CAediting_248824 | chr2_232705927_+ | SINE | Alu | AluSq2 |
| CAediting_248825 | chr2_232705952_+ | SINE | Alu | AluSq2 |
| CAediting_248826 | chr2_232706631_+ | SINE | Alu | AluSz |
| CAediting_248827 | chr2_232706657_+ | SINE | Alu | AluSz |
| CAediting_248828 | chr2_232706700_+ | SINE | Alu | AluSz |
| CAediting_248829 | chr2_232706701_+ | SINE | Alu | AluSz |
| CAediting_248830 | chr2_232706721_+ | SINE | Alu | AluSz |
| CAediting_248831 | chr2_232706767_+ | SINE | Alu | AluSz |
| CAediting_248832 | chr2_232706813_+ | SINE | Alu | AluJb |
| CAediting_248833 | chr2_232706826_+ | SINE | Alu | AluJb |
| CAediting_248834 | chr2_232706827_+ | SINE | Alu | AluJb |
| CAediting_248835 | chr2_232706865_+ | SINE | Alu | AluJb |
| CAediting_248836 | chr2_232706891_+ | SINE | Alu | AluJb |
| CAediting_248837 | chr2_232708094_+ | SINE | Alu | AluJb |
| CAediting_248838 | chr2_232708160_+ | SINE | Alu | AluJb |
| CAediting_248839 | chr2_232708228_+ | SINE | Alu | AluJb |
| CAediting_248840 | chr2_232708558_+ | SINE | Alu | AluJo |
| CAediting_248841 | chr2_232708564_+ | SINE | Alu | AluJo |
| CAediting_248842 | chr2_232708580_+ | SINE | Alu | AluJo |
| CAediting_248843 | chr2_232708581_+ | SINE | Alu | AluJo |
| CAediting_248844 | chr2_232708586_+ | SINE | Alu | AluJo |
| CAediting_248845 | chr2_232708595_+ | SINE | Alu | AluJo |
| CAediting_248846 | chr2_232708596_+ | SINE | Alu | AluJo |
| CAediting_248847 | chr2_232708606_+ | SINE | Alu | AluJo |
| CAediting_248848 | chr2_232708636_+ | SINE | Alu | AluJo |
| CAediting_248849 | chr2_232708867_+ | SINE | Alu | AluSx1 |
| CAediting_248850 | chr2_232708884_+ | SINE | Alu | AluSx1 |
| CAediting_248851 | chr2_232708912_+ | SINE | Alu | AluSx1 |
| CAediting_248852 | chr2_232708913_+ | SINE | Alu | AluSx1 |
| CAediting_248853 | chr2_232708956_+ | SINE | Alu | AluSx1 |
| CAediting_248854 | chr2_232708974_+ | SINE | Alu | AluSx1 |
| CAediting_248855 | chr2_232709363_+ | SINE | Alu | AluJo |
| CAediting_248856 | chr2_232709711_+ | SINE | Alu | AluSg |
| CAediting_248857 | chr2_232710029_+ | SINE | Alu | AluY |
| CAediting_248858 | chr2_232710166_+ | SINE | Alu | AluY |
| CAediting_248859 | chr2_232710221_+ | SINE | Alu | AluY |
| CAediting_248860 | chr2_232710233_+ | SINE | Alu | AluY |
| CAediting_248861 | chr2_232710294_+ | SINE | Alu | FLAM_C |
| CAediting_248862 | chr2_232710317_+ | SINE | Alu | FLAM_C |
| CAediting_248863 | chr2_232710323_+ | SINE | Alu | FLAM_C |
| CAediting_248864 | chr2_232710324_+ | SINE | Alu | FLAM_C |
| CAediting_248865 | chr2_232713176_+ | SINE | Alu | AluSx1 |
| CAediting_248866 | chr2_232717178_+ | SINE | Alu | AluJr4 |
| CAediting_248867 | chr2_232717251_+ | SINE | Alu | AluJr4 |
| CAediting_248868 | chr2_232718119_+ | SINE | Alu | AluSx1 |
| CAediting_248869 | chr2_232718237_+ | SINE | Alu | AluSx1 |
| CAediting_248870 | chr2_232724576_+ | SINE | Alu | AluJr |
| CAediting_248871 | chr2_232724592_+ | SINE | Alu | AluJr |
| CAediting_248872 | chr2_232724601_+ | SINE | Alu | AluJr |
| CAediting_248873 | chr2_232724643_+ | SINE | Alu | AluJr |
| CAediting_248874 | chr2_232724649_+ | SINE | Alu | AluJr |
| CAediting_248875 | chr2_232724650_+ | SINE | Alu | AluJr |
| CAediting_248876 | chr2_232724651_+ | SINE | Alu | AluJr |
| CAediting_248877 | chr2_232724662_+ | SINE | Alu | AluJr |
| CAediting_248878 | chr2_232724667_+ | SINE | Alu | AluJr |
| CAediting_248879 | chr2_232724675_+ | SINE | Alu | AluJr |
| CAediting_248880 | chr2_232724676_+ | SINE | Alu | AluJr |
| CAediting_248881 | chr2_232724679_+ | SINE | Alu | AluJr |
| CAediting_248882 | chr2_232724690_+ | SINE | Alu | AluJr |
| CAediting_248883 | chr2_232724805_+ | SINE | Alu | AluJr |
| CAediting_248884 | chr2_232724815_+ | SINE | Alu | AluJr |
| CAediting_248887 | chr2_232726116_+ | SINE | Alu | AluSx1 |
| CAediting_248888 | chr2_232726132_+ | SINE | Alu | AluSx1 |
| CAediting_248889 | chr2_232726137_+ | SINE | Alu | AluSx1 |
| CAediting_248890 | chr2_232726150_+ | SINE | Alu | AluSx1 |
| CAediting_248891 | chr2_232726160_+ | SINE | Alu | AluSx1 |
| CAediting_248892 | chr2_232726163_+ | SINE | Alu | AluSx1 |
| CAediting_248893 | chr2_232726183_+ | SINE | Alu | AluSx1 |
| CAediting_248894 | chr2_232726184_+ | SINE | Alu | AluSx1 |
| CAediting_248895 | chr2_232726186_+ | SINE | Alu | AluSx1 |
| CAediting_248896 | chr2_232726190_+ | SINE | Alu | AluSx1 |
| CAediting_248897 | chr2_232726193_+ | SINE | Alu | AluSx1 |
| CAediting_248898 | chr2_232726204_+ | SINE | Alu | AluSx1 |
| CAediting_248899 | chr2_232726208_+ | SINE | Alu | AluSx1 |
| CAediting_248900 | chr2_232726209_+ | SINE | Alu | AluSx1 |
| CAediting_248901 | chr2_232726210_+ | SINE | Alu | AluSx1 |
| CAediting_248902 | chr2_232726211_+ | SINE | Alu | AluSx1 |
| CAediting_248903 | chr2_232726213_+ | SINE | Alu | AluSx1 |
| CAediting_248904 | chr2_232726215_+ | SINE | Alu | AluSx1 |
| CAediting_248905 | chr2_232726216_+ | SINE | Alu | AluSx1 |
| CAediting_248906 | chr2_232726217_+ | SINE | Alu | AluSx1 |
| CAediting_248907 | chr2_232726218_+ | SINE | Alu | AluSx1 |
| CAediting_248908 | chr2_232726219_+ | SINE | Alu | AluSx1 |
| CAediting_248909 | chr2_232726248_+ | SINE | Alu | AluSx1 |
| CAediting_248910 | chr2_232726254_+ | SINE | Alu | AluSx1 |
| CAediting_248911 | chr2_232726258_+ | SINE | Alu | AluSx1 |
| CAediting_248912 | chr2_232726275_+ | SINE | Alu | AluSx1 |
| CAediting_248913 | chr2_232726277_+ | SINE | Alu | AluSx1 |
| CAediting_248914 | chr2_232726278_+ | SINE | Alu | AluSx1 |
| CAediting_248915 | chr2_232726289_+ | SINE | Alu | AluSx1 |
| CAediting_248916 | chr2_232726290_+ | SINE | Alu | AluSx1 |
| CAediting_248917 | chr2_232726297_+ | SINE | Alu | AluSx1 |
| CAediting_248918 | chr2_232726326_+ | SINE | Alu | AluSx1 |
| CAediting_248919 | chr2_232726485_+ | SINE | Alu | AluJb |
| CAediting_248920 | chr2_232726495_+ | SINE | Alu | AluJb |
| CAediting_248921 | chr2_232727064_+ | SINE | Alu | AluSz6 |
| CAediting_248922 | chr2_232727075_+ | SINE | Alu | AluSz6 |
| CAediting_248923 | chr2_232727082_+ | SINE | Alu | AluSz6 |
| CAediting_248924 | chr2_232727094_+ | SINE | Alu | AluSz6 |
| CAediting_248925 | chr2_232727095_+ | SINE | Alu | AluSz6 |
| CAediting_248926 | chr2_232727104_+ | SINE | Alu | AluSz6 |
| CAediting_248927 | chr2_232727126_+ | SINE | Alu | AluSz6 |
| CAediting_248928 | chr2_232727131_+ | SINE | Alu | AluSz6 |
| CAediting_248929 | chr2_232727146_+ | SINE | Alu | AluSz6 |
| CAediting_248930 | chr2_232727168_+ | SINE | Alu | AluSz6 |
| CAediting_248931 | chr2_232730484_+ | SINE | Alu | AluSz |
| CAediting_248932 | chr2_232730485_+ | SINE | Alu | AluSz |
| CAediting_248933 | chr2_232730511_+ | SINE | Alu | AluSz |
| CAediting_248934 | chr2_232730520_+ | SINE | Alu | AluSz |
| CAediting_248935 | chr2_232730526_+ | SINE | Alu | AluSz |
| CAediting_248936 | chr2_232730554_+ | SINE | Alu | AluSz |
| CAediting_248937 | chr2_232732665_+ | SINE | Alu | AluJb |
| CAediting_248938 | chr2_232732704_+ | SINE | Alu | AluJb |
| CAediting_248939 | chr2_232732708_+ | SINE | Alu | AluJb |
| CAediting_248940 | chr2_232732768_+ | SINE | Alu | AluJb |
| CAediting_248941 | chr2_232732790_+ | SINE | Alu | AluJb |
| CAediting_248942 | chr2_232732794_+ | SINE | Alu | AluJb |
| CAediting_248943 | chr2_232732830_+ | SINE | Alu | AluJb |
| CAediting_248944 | chr2_232733001_+ | SINE | Alu | AluY |
| CAediting_248945 | chr2_232733002_+ | SINE | Alu | AluY |
| CAediting_248946 | chr2_232733045_+ | SINE | Alu | AluY |
| CAediting_248947 | chr2_232733054_+ | SINE | Alu | AluY |
| CAediting_248948 | chr2_232733079_+ | SINE | Alu | AluY |
| CAediting_248949 | chr2_232733126_+ | SINE | Alu | AluY |
| CAediting_248950 | chr2_232733136_+ | SINE | Alu | AluY |
| CAediting_248951 | chr2_232733163_+ | SINE | Alu | AluY |
| CAediting_248952 | chr2_232733202_+ | SINE | Alu | AluY |
| CAediting_248953 | chr2_232733209_+ | SINE | Alu | AluY |
| CAediting_248954 | chr2_232734470_+ | SINE | Alu | AluJr |
| CAediting_248955 | chr2_232734471_+ | SINE | Alu | AluJr |
| CAediting_248956 | chr2_232734479_+ | SINE | Alu | AluJr |
| CAediting_248957 | chr2_232738016_+ | SINE | Alu | AluSx1 |
| CAediting_248958 | chr2_232738056_+ | SINE | Alu | AluSx1 |
| CAediting_248959 | chr2_232738114_+ | SINE | Alu | AluSx1 |
| CAediting_248960 | chr2_232739144_+ | SINE | Alu | AluSq2 |
| CAediting_248961 | chr2_232739198_+ | SINE | Alu | AluSq2 |
| CAediting_248962 | chr2_232739199_+ | SINE | Alu | AluSq2 |
| CAediting_248963 | chr2_232739200_+ | SINE | Alu | AluSq2 |
| CAediting_248964 | chr2_232739201_+ | SINE | Alu | AluSq2 |
| CAediting_248965 | chr2_232739203_+ | SINE | Alu | AluSq2 |
| CAediting_248966 | chr2_232739207_+ | SINE | Alu | AluSq2 |
| CAediting_248967 | chr2_232739237_+ | SINE | Alu | AluSq2 |
| CAediting_248968 | chr2_232739238_+ | SINE | Alu | AluSq2 |
| CAediting_248969 | chr2_232739251_+ | SINE | Alu | AluSq2 |
| CAediting_248970 | chr2_232739264_+ | SINE | Alu | AluSq2 |
| CAediting_248971 | chr2_232739283_+ | SINE | Alu | AluSq2 |
| CAediting_248972 | chr2_232739293_+ | SINE | Alu | AluSq2 |
| CAediting_248973 | chr2_232739529_+ | SINE | Alu | AluJb |
| CAediting_248974 | chr2_232739582_+ | SINE | Alu | AluJb |
| CAediting_248975 | chr2_232739586_+ | SINE | Alu | AluJb |
| CAediting_248976 | chr2_232739588_+ | SINE | Alu | AluJb |
| CAediting_248977 | chr2_232739594_+ | SINE | Alu | AluJb |
| CAediting_248978 | chr2_232739891_+ | SINE | Alu | AluSz |
| CAediting_248979 | chr2_232739967_+ | SINE | Alu | AluSz |
| CAediting_248980 | chr2_232739998_+ | SINE | Alu | AluSz |
| CAediting_248981 | chr2_232740007_+ | SINE | Alu | AluSz |
| CAediting_248982 | chr2_232740024_+ | SINE | Alu | AluSz |
| CAediting_248983 | chr2_232740039_+ | SINE | Alu | AluSz |
| CAediting_248984 | chr2_232741624_+ | SINE | Alu | AluSz |
| CAediting_248985 | chr2_232742291_+ | SINE | Alu | AluSx |
| CAediting_248986 | chr2_232742382_+ | SINE | Alu | AluSx |
| CAediting_248987 | chr2_232742383_+ | SINE | Alu | AluSx |
| CAediting_248988 | chr2_232742385_+ | SINE | Alu | AluSx |
| CAediting_248989 | chr2_232742398_+ | SINE | Alu | AluSx |
| CAediting_248990 | chr2_232743973_+ | SINE | Alu | AluJr |
| CAediting_248991 | chr2_232744671_+ | SINE | Alu | AluJb |
| CAediting_248992 | chr2_232744808_+ | SINE | Alu | AluJb |
| CAediting_248993 | chr2_232750883_+ | SINE | Alu | AluJr |
| CAediting_248994 | chr2_232750957_+ | SINE | Alu | AluJr |
| CAediting_248995 | chr2_232753223_+ | SINE | Alu | AluSx1 |
| CAediting_248996 | chr2_232753224_+ | SINE | Alu | AluSx1 |
| CAediting_248997 | chr2_232753892_+ | SINE | Alu | AluSc8 |
| CAediting_248998 | chr2_232753904_+ | SINE | Alu | AluSc8 |
| CAediting_248999 | chr2_232753950_+ | SINE | Alu | AluSc8 |
| CAediting_249000 | chr2_232753993_+ | SINE | Alu | AluSc8 |
| CAediting_249001 | chr2_232753999_+ | SINE | Alu | AluSc8 |
| CAediting_249002 | chr2_232754002_+ | SINE | Alu | AluSc8 |
| CAediting_249003 | chr2_232754004_+ | SINE | Alu | AluSc8 |
| CAediting_249004 | chr2_232754005_+ | SINE | Alu | AluSc8 |
| CAediting_249005 | chr2_232754008_+ | SINE | Alu | AluSc8 |
| CAediting_249006 | chr2_232754018_+ | SINE | Alu | AluSc8 |
| CAediting_249007 | chr2_232754026_+ | SINE | Alu | AluSc8 |
| CAediting_249008 | chr2_232754044_+ | SINE | Alu | AluSc8 |
| CAediting_249009 | chr2_232754054_+ | SINE | Alu | AluSc8 |
| CAediting_249010 | chr2_232754058_+ | SINE | Alu | AluSc8 |
| CAediting_249011 | chr2_232754086_+ | SINE | Alu | AluSc8 |
| CAediting_249012 | chr2_232754110_+ | SINE | Alu | AluSc8 |
| CAediting_249013 | chr2_232754153_+ | SINE | Alu | AluSc8 |
| CAediting_249015 | chr2_232782272_+ | SINE | Alu | AluJb |
| CAediting_249016 | chr2_232784235_+ | SINE | Alu | AluJr |
| CAediting_249017 | chr2_232784274_+ | SINE | Alu | AluJr |
| CAediting_249018 | chr2_232797727_+ | SINE | Alu | AluSz6 |
| CAediting_249019 | chr2_232801004_+ | SINE | Alu | AluSz6 |
| CAediting_249020 | chr2_232807438_+ | SINE | Alu | AluJo |
| CAediting_249021 | chr2_232807448_+ | SINE | Alu | AluJo |
| CAediting_249022 | chr2_232809102_+ | SINE | Alu | AluSg |
| CAediting_249023 | chr2_232812858_+ | SINE | Alu | AluJr |
| CAediting_249024 | chr2_232812880_+ | SINE | Alu | AluJr |
| CAediting_249025 | chr2_232812897_+ | SINE | Alu | AluJr |
| CAediting_249026 | chr2_232812916_+ | SINE | Alu | AluJr |
| CAediting_249027 | chr2_232812926_+ | SINE | Alu | AluJr |
| CAediting_249028 | chr2_232812942_+ | SINE | Alu | AluJr |
| CAediting_249029 | chr2_232812962_+ | SINE | Alu | AluJr |
| CAediting_249030 | chr2_232814154_+ | SINE | Alu | AluSx1 |
| CAediting_249031 | chr2_232820480_+ | SINE | Alu | AluY |
| CAediting_249032 | chr2_232823457_+ | SINE | Alu | AluJr |
| CAediting_249033 | chr2_232828484_+ | SINE | Alu | AluJr |
| CAediting_249034 | chr2_232828552_+ | SINE | Alu | AluJr |
| CAediting_249035 | chr2_232828553_+ | SINE | Alu | AluJr |
| CAediting_249036 | chr2_232828582_+ | SINE | Alu | AluJr |
| CAediting_249037 | chr2_232828588_+ | SINE | Alu | AluJr |
| CAediting_249038 | chr2_232828595_+ | SINE | Alu | AluJr |
| CAediting_249039 | chr2_232830011_+ | SINE | Alu | AluJr |
| CAediting_249040 | chr2_232830126_+ | SINE | Alu | AluJr |
| CAediting_249041 | chr2_232830167_+ | SINE | Alu | AluJr |
| CAediting_249042 | chr2_232830203_+ | SINE | Alu | AluJr |
| CAediting_249043 | chr2_232836459_+ | SINE | Alu | AluJb |
| CAediting_249044 | chr2_232836481_+ | SINE | Alu | AluJb |
| CAediting_249045 | chr2_232848424_+ | SINE | Alu | AluSg7 |
| CAediting_249046 | chr2_232848511_+ | SINE | Alu | AluSg7 |
| CAediting_249047 | chr2_232851092_+ | SINE | Alu | AluJr |
| CAediting_249048 | chr2_232851178_+ | SINE | Alu | AluJr |
| CAediting_249049 | chr2_232851196_+ | SINE | Alu | AluJr |
| CAediting_249050 | chr2_232854243_+ | SINE | Alu | AluJb |
| CAediting_249051 | chr2_232855187_+ | SINE | Alu | AluSz6 |
| CAediting_249052 | chr2_232855204_+ | SINE | Alu | AluSz6 |
| CAediting_249053 | chr2_232856412_+ | SINE | Alu | AluSz |
| CAediting_249054 | chr2_232856451_+ | SINE | Alu | AluSz |
| CAediting_249055 | chr2_232856573_+ | SINE | Alu | AluSz |
| CAediting_249056 | chr2_232856647_+ | SINE | Alu | AluSz |
| CAediting_249057 | chr2_232860206_+ | SINE | Alu | FLAM_A |
| CAediting_249058 | chr2_232860232_+ | SINE | Alu | FLAM_A |
Top |
RNA A-to-I editing events in the alternative splicing sites for GIGYF2 |
 RNA A-to-I editing(s) in the alternative splicing sites. RNA A-to-I editing(s) in the alternative splicing sites. |
| AStype | CAeditomID | Editing position | AS position | AS direction | Exonic location | Wildtype sequence | Wildtype splicing strength | RNA edited sequence | RNA edited splicing strength |
| ES | CAediting_248870 | chr2_232724576_+ | 232724555:232724577 | 3SS | 2+0i | ATTTTTTTTTTCCTTTTAGAGAC | -4.91 | ATTTTTTTTTTCCTTTTAGAGGC | -5.05 |
| MEX_1 | CAediting_248870 | chr2_232724576_+ | 232724555:232724577 | 3SS | 2+0i | ATTTTTTTTTTCCTTTTAGAGAC | -4.91 | ATTTTTTTTTTCCTTTTAGAGGC | -5.05 |
| MEX_2 | CAediting_248870 | chr2_232724576_+ | 232724555:232724577 | 3SS | 2+0i | ATTTTTTTTTTCCTTTTAGAGAC | -4.91 | ATTTTTTTTTTCCTTTTAGAGGC | -5.05 |
| ES | CAediting_248871 | chr2_232724592_+ | 232724577:232724599 | 3SS | 0+5i | CTGTCTCGCTCTGTCACCCAGGC | -5.97 | CTGTCTCGCTCTGTCGCCCAGGC | -7.06 |
| MEX_1 | CAediting_248871 | chr2_232724592_+ | 232724577:232724599 | 3SS | 0+5i | CTGTCTCGCTCTGTCACCCAGGC | -5.97 | CTGTCTCGCTCTGTCGCCCAGGC | -7.06 |
| ES | CAediting_248882 | chr2_232724690_+ | 232724689:232724697 | 5SS | 2+0i | AAGGCAAGT | 3.24 | AGGGCAAGT | 2.7 |
| MEX_1 | CAediting_248882 | chr2_232724690_+ | 232724689:232724697 | 5SS | 2+0i | AAGGCAAGT | 3.24 | AGGGCAAGT | 2.7 |
| MEX_2 | CAediting_248882 | chr2_232724690_+ | 232724689:232724697 | 5SS | 2+0i | AAGGCAAGT | 3.24 | AGGGCAAGT | 2.7 |
 Differential percent of spliced in (PSI) between RNA A-to-I edited and non-edited samples for ENSG00000204120.13.GIGYF2. Differential percent of spliced in (PSI) between RNA A-to-I edited and non-edited samples for ENSG00000204120.13.GIGYF2. |
 Correlation between RNA A-to-I editing and PSI for ENSG00000204120.13.GIGYF2. Correlation between RNA A-to-I editing and PSI for ENSG00000204120.13.GIGYF2. |
Top |
Differntial gene expression between RNA A-to-I edited versus non-edited samples for GIGYF2 |
 Differential gene expressions between RNA A-to-I edited and non-edited tumor samples. Differential gene expressions between RNA A-to-I edited and non-edited tumor samples.* The grey color means N/A. |
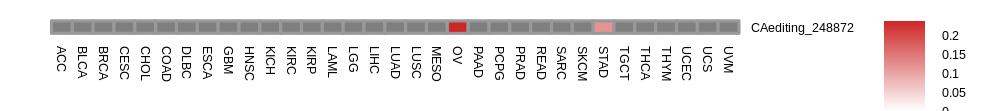 | |
 Correlation between RNA A-to-I editing and gene expression. Correlation between RNA A-to-I editing and gene expression. |
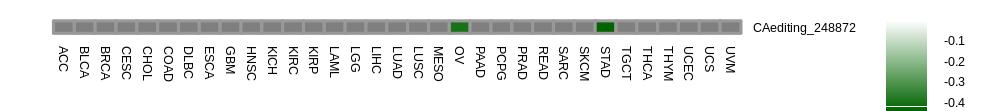 | |
 - Differentially expressed gene between RNA A-to-I edited samples versus non-edited samples. - Differentially expressed gene between RNA A-to-I edited samples versus non-edited samples.* Click on the image to enlarge it in a new window. |
Top |
Protein coding region RNA A-to-I editings for GIGYF2 |
 - Lollipop plot for RNA A-to-I editings across protein structure. - Lollipop plot for RNA A-to-I editings across protein structure. * Click on the image to enlarge it in a new window. |
Top |
The effects of the RNA editing to the miRNA binding sites for GIGYF2 |
| **If you are searching a miRNA gene with RNA editing events in its seed regions, please see the search pages of miRNA targets to discover the effects of this RNA editing event to miRNA regulations. For more information, please check the files in the Download page. |
 RNA A-to-I editing in the 3'-UTR regions of mRNA gained miRNA binding sites. RNA A-to-I editing in the 3'-UTR regions of mRNA gained miRNA binding sites. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-3940-3p | 232860203 | 232860209 | 7mer-m8 | 232860189 | 232860210 | 142.00 | -21.15 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-7112-5p | 232860200 | 232860205 | 6mer | 232860183 | 232860206 | 143.00 | -24.03 |
| CAediting_249058 | chr2_232860232_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-1199-5p | 232860225 | 232860231 | 7mer-m8 | 232860213 | 232860232 | 156.00 | -28.10 |
| CAediting_249058 | chr2_232860232_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-3622a-3p | 232860228 | 232860235 | 8mer-1a | 232860214 | 232860235 | 148.00 | -19.39 |
| CAediting_249058 | chr2_232860232_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-3622b-3p | 232860228 | 232860235 | 8mer-1a | 232860215 | 232860235 | 157.00 | -23.90 |
| CAediting_249058 | chr2_232860232_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-6751-3p | 232860225 | 232860231 | 7mer-m8 | 232860212 | 232860232 | 141.00 | -13.15 |
 RNA A-to-I editing in the 3'-UTR regions of mRNA lost miRNA binding sites. RNA A-to-I editing in the 3'-UTR regions of mRNA lost miRNA binding sites. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-3116 | 232860203 | 232860208 | 6mer | 232860186 | 232860209 | 147 | -31.34 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-3116 | 232860203 | 232860208 | 6mer | 232860186 | 232860209 | 147 | -31.34 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-5189-5p | 232860200 | 232860206 | 7mer-m8 | 232860184 | 232860207 | 150 | -20.87 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-612 | 232860200 | 232860206 | 7mer-m8 | 232860184 | 232860207 | 149 | -21.95 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-661 | 232860202 | 232860208 | 7mer-m8 | 232860185 | 232860209 | 142 | -18.63 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-6860 | 232860200 | 232860206 | 7mer-m8 | 232860186 | 232860207 | 148 | -22.51 |
| CAediting_249057 | chr2_232860206_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-939-3p | 232860201 | 232860207 | 7mer-m8 | 232860188 | 232860208 | 140 | -15.63 |
 RNA A-to-I editing in lncRNA gained miRNA binding sites. RNA A-to-I editing in lncRNA gained miRNA binding sites. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
 RNA A-to-I editing in lncRNA lost miRNA binding sites. RNA A-to-I editing in lncRNA lost miRNA binding sites. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
 RNA A-to-I editing in miRNAs gained binding to lncRNA. RNA A-to-I editing in miRNAs gained binding to lncRNA. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
 RNA A-to-I editing in miRNAs lost binding to lncRNA. RNA A-to-I editing in miRNAs lost binding to lncRNA. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
 RNA A-to-I editing in miRNAs gained binding to the 3'-UTR region of mRNA. RNA A-to-I editing in miRNAs gained binding to the 3'-UTR region of mRNA. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
| CAediting_748296 | chr10_51299626_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-605-3p | 232858877 | 232858884 | 8mer-1a | 232858863 | 232858884 | 152 | -16.18 |
| CAediting_980496 | chr14_31014685_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-624-3p | 232857072 | 232857078 | 7mer-m8 | 232857061 | 232857079 | 147 | -19.64 |
| CAediting_980496 | chr14_31014685_- | ENSG00000204120.13 | ENST00000409480 | hsa-miR-624-3p | 232857072 | 232857078 | 7mer-m8 | 232857061 | 232857079 | 147 | -19.64 |
| CAediting_980496 | chr14_31014685_- | ENSG00000204120.13 | ENST00000409547 | hsa-miR-624-3p | 232857072 | 232857078 | 7mer-m8 | 232857061 | 232857079 | 147 | -19.64 |
| CAediting_980496 | chr14_31014685_- | ENSG00000204120.13 | ENST00000409196 | hsa-miR-624-3p | 232857072 | 232857078 | 7mer-m8 | 232857061 | 232857079 | 147 | -19.64 |
| CAediting_980496 | chr14_31014685_- | ENSG00000204120.13 | ENST00000409451 | hsa-miR-624-3p | 232857072 | 232857078 | 7mer-m8 | 232857061 | 232857079 | 147 | -19.64 |
| CAediting_1229415 | chr17_67471547_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-548aa | 232860140 | 232860147 | 8mer-1a | 232860123 | 232860147 | 140 | -9.58 |
| CAediting_1229415 | chr17_67471547_+ | ENSG00000204120.13 | ENST00000629305 | hsa-miR-548aa | 232860333 | 232860340 | 8mer-1a | 232860316 | 232860340 | 168 | -11.76 |
| CAediting_1351755 | chr19_40282625_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-641 | 232857162 | 232857167 | 6mer | 232857145 | 232857168 | 145 | -19.4 |
| CAediting_1368054 | chr19_46709310_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-320e | 232859424 | 232859430 | 7mer-m8 | 232859413 | 232859431 | 147 | -19.97 |
| CAediting_524911 | chr7_5495852_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-589-3p | 232858282 | 232858289 | 8mer-1a | 232858266 | 232858289 | 156 | -23.41 |
| CAediting_1351755 | chr19_40282625_- | ENSG00000204120.13 | ENST00000409480 | hsa-miR-641 | 232857162 | 232857167 | 6mer | 232857145 | 232857168 | 145 | -19.4 |
| CAediting_524911 | chr7_5495852_- | ENSG00000204120.13 | ENST00000409480 | hsa-miR-589-3p | 232858282 | 232858289 | 8mer-1a | 232858266 | 232858289 | 156 | -23.41 |
| CAediting_1351755 | chr19_40282625_- | ENSG00000204120.13 | ENST00000409547 | hsa-miR-641 | 232857162 | 232857167 | 6mer | 232857145 | 232857168 | 145 | -19.4 |
| CAediting_524911 | chr7_5495852_- | ENSG00000204120.13 | ENST00000409547 | hsa-miR-589-3p | 232858282 | 232858289 | 8mer-1a | 232858266 | 232858289 | 156 | -23.41 |
| CAediting_1351755 | chr19_40282625_- | ENSG00000204120.13 | ENST00000409196 | hsa-miR-641 | 232857162 | 232857167 | 6mer | 232857145 | 232857168 | 145 | -19.4 |
| CAediting_524911 | chr7_5495852_- | ENSG00000204120.13 | ENST00000409196 | hsa-miR-589-3p | 232858282 | 232858289 | 8mer-1a | 232858266 | 232858289 | 156 | -23.41 |
| CAediting_1351755 | chr19_40282625_- | ENSG00000204120.13 | ENST00000409451 | hsa-miR-641 | 232857162 | 232857167 | 6mer | 232857145 | 232857168 | 145 | -19.4 |
| CAediting_524911 | chr7_5495852_- | ENSG00000204120.13 | ENST00000409451 | hsa-miR-589-3p | 232858282 | 232858289 | 8mer-1a | 232858266 | 232858289 | 156 | -23.41 |
| CAediting_77128 | chr1_85133844_+ | ENSG00000204120.13 | ENST00000458528 | hsa-miR-4423-3p | 232791427 | 232791433 | 7mer-m8 | 232791414 | 232791434 | 140 | -12.68 |
| CAediting_604450 | chr7_133034877_- | ENSG00000204120.13 | ENST00000458528 | hsa-miR-3654 | 232794761 | 232794768 | 8mer-1a | 232794748 | 232794768 | 147 | -16.58 |
 RNA A-to-I editing in miRNAs lost binding to the 3'-UTR mRNA. RNA A-to-I editing in miRNAs lost binding to the 3'-UTR mRNA. |
| CAeditom_ID | Position | ENSG | ENST | miRNA | Targetscan start | Targetscan end | Targetscan score | Miranda start | Miranda end | Miranda score | Miranda energy |
| CAediting_1368054 | chr19_46709310_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-320e | 232860290 | 232860296 | 7mer-m8 | 232860280 | 232860297 | 150 | -17.95 |
| CAediting_396018 | chr5_1062918_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-4635 | 232858074 | 232858080 | 7mer-m8 | 232858061 | 232858081 | 145 | -10.54 |
| CAediting_1351755 | chr19_40282625_- | ENSG00000204120.13 | ENST00000629305 | hsa-miR-641 | 232859658 | 232859664 | 7mer-m8 | 232859641 | 232859665 | 155 | -15.45 |
| CAediting_396018 | chr5_1062918_- | ENSG00000204120.13 | ENST00000409480 | hsa-miR-4635 | 232858074 | 232858080 | 7mer-m8 | 232858061 | 232858081 | 145 | -10.54 |
| CAediting_396018 | chr5_1062918_- | ENSG00000204120.13 | ENST00000409547 | hsa-miR-4635 | 232858074 | 232858080 | 7mer-m8 | 232858061 | 232858081 | 145 | -10.54 |
| CAediting_396018 | chr5_1062918_- | ENSG00000204120.13 | ENST00000409196 | hsa-miR-4635 | 232858074 | 232858080 | 7mer-m8 | 232858061 | 232858081 | 145 | -10.54 |
| CAediting_396018 | chr5_1062918_- | ENSG00000204120.13 | ENST00000409451 | hsa-miR-4635 | 232858074 | 232858080 | 7mer-m8 | 232858061 | 232858081 | 145 | -10.54 |
 Differentially expressed gene-miRNA network Differentially expressed gene-miRNA network |
Top |
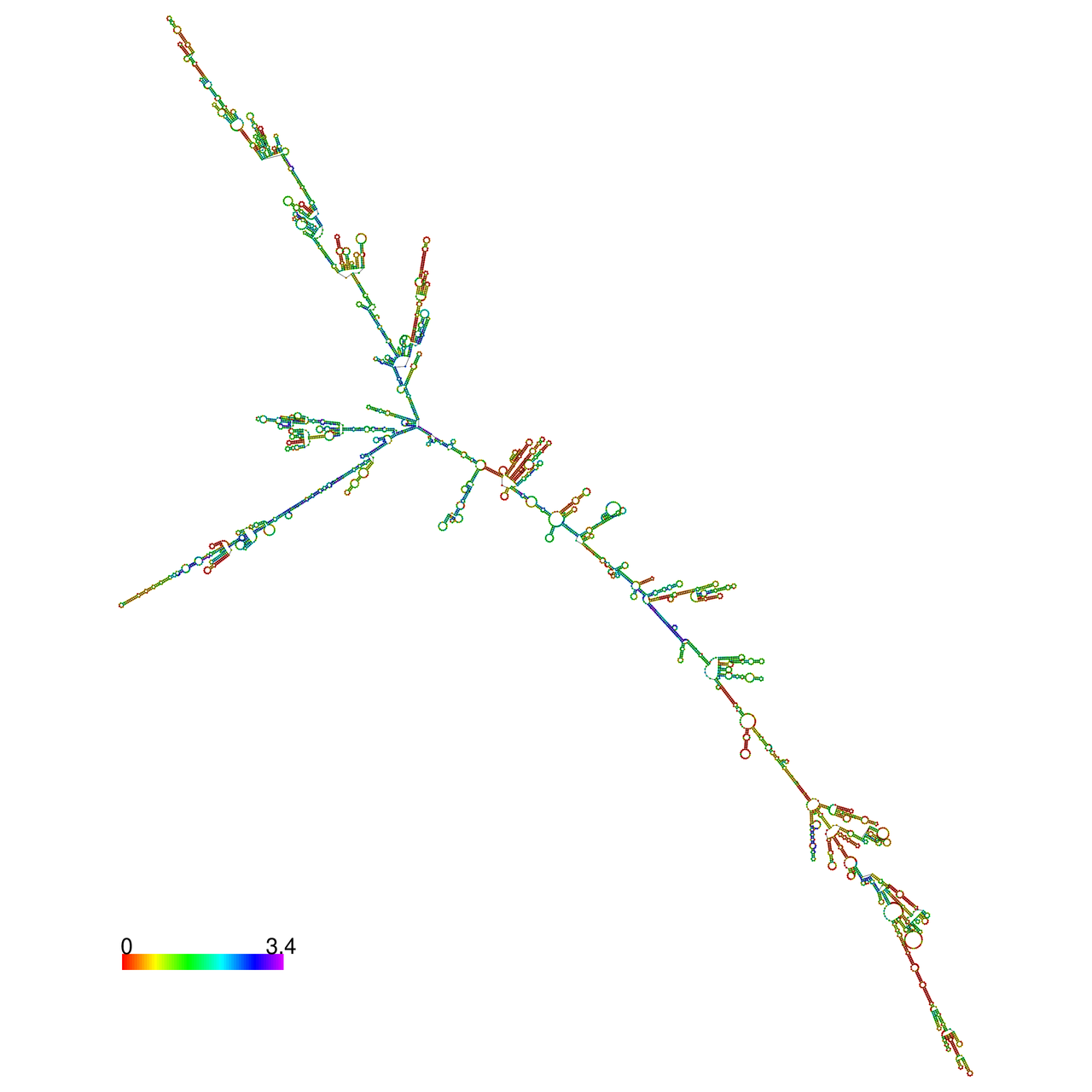
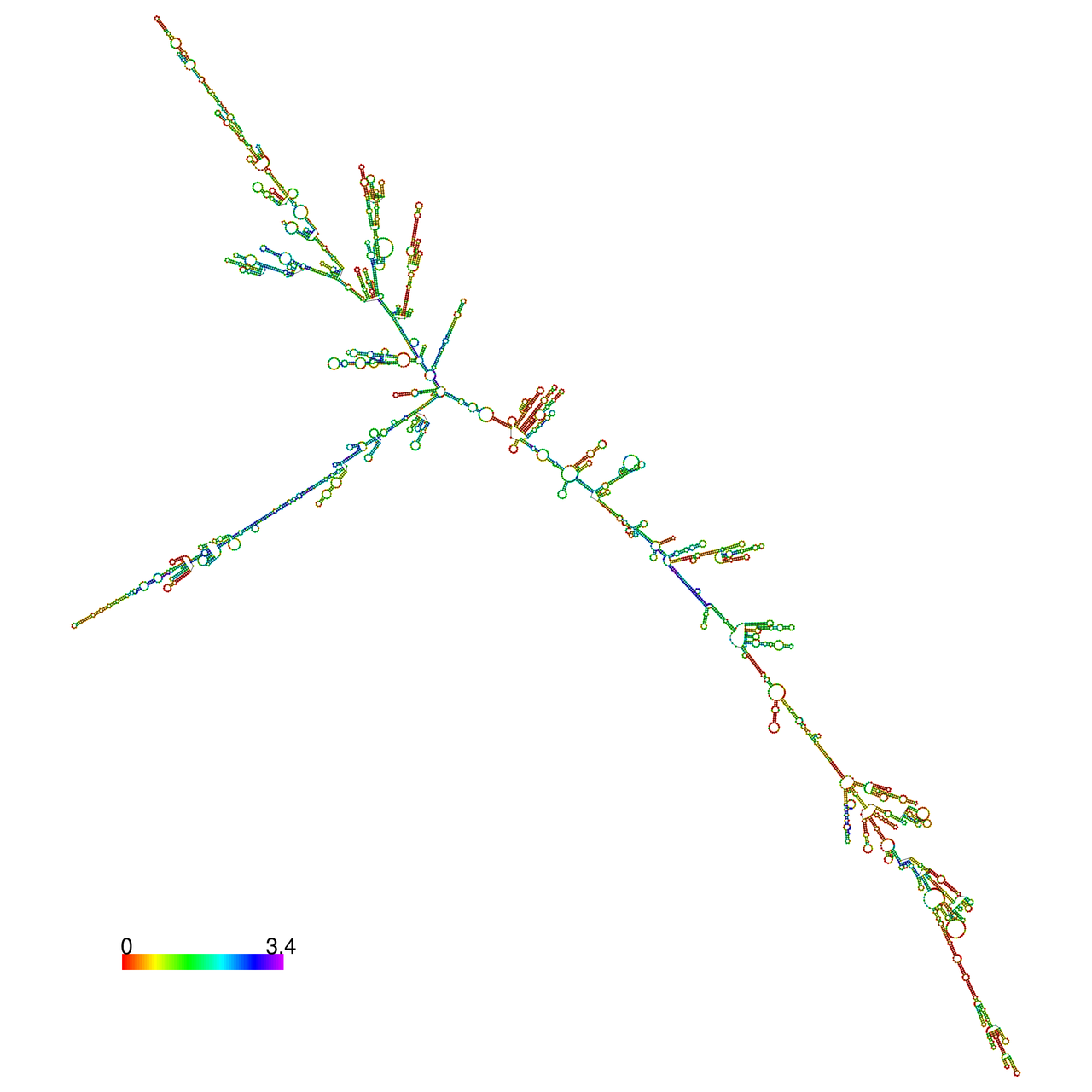
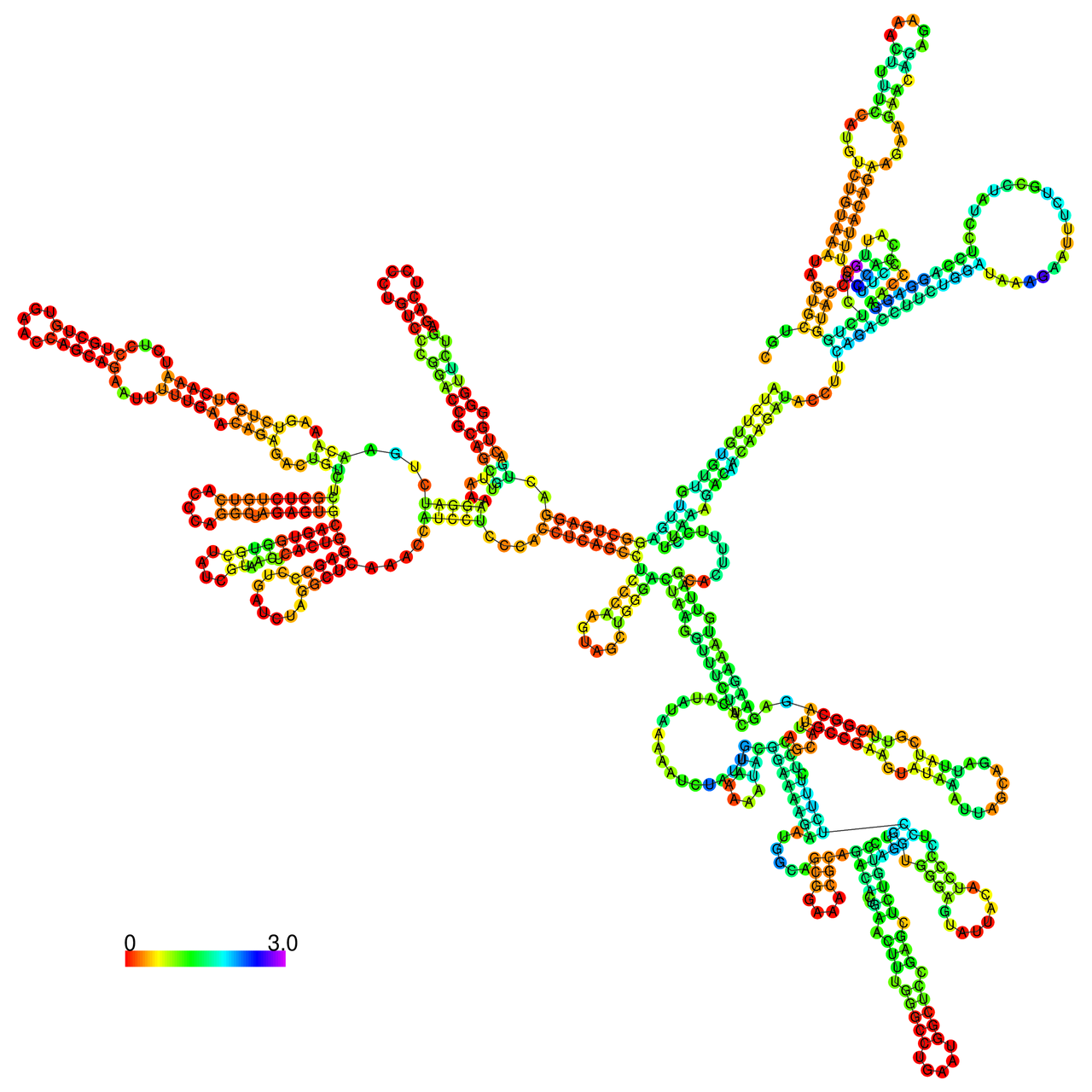
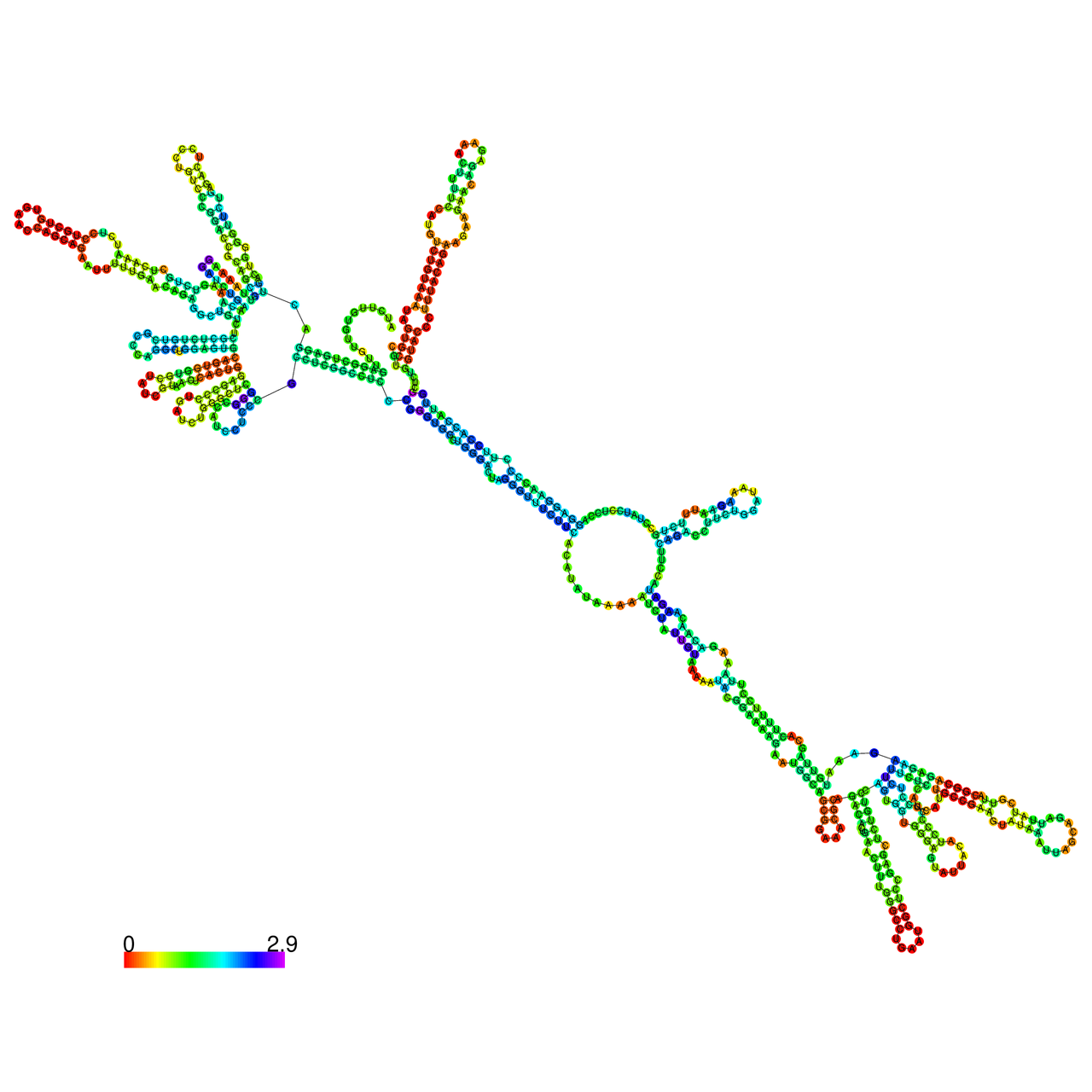
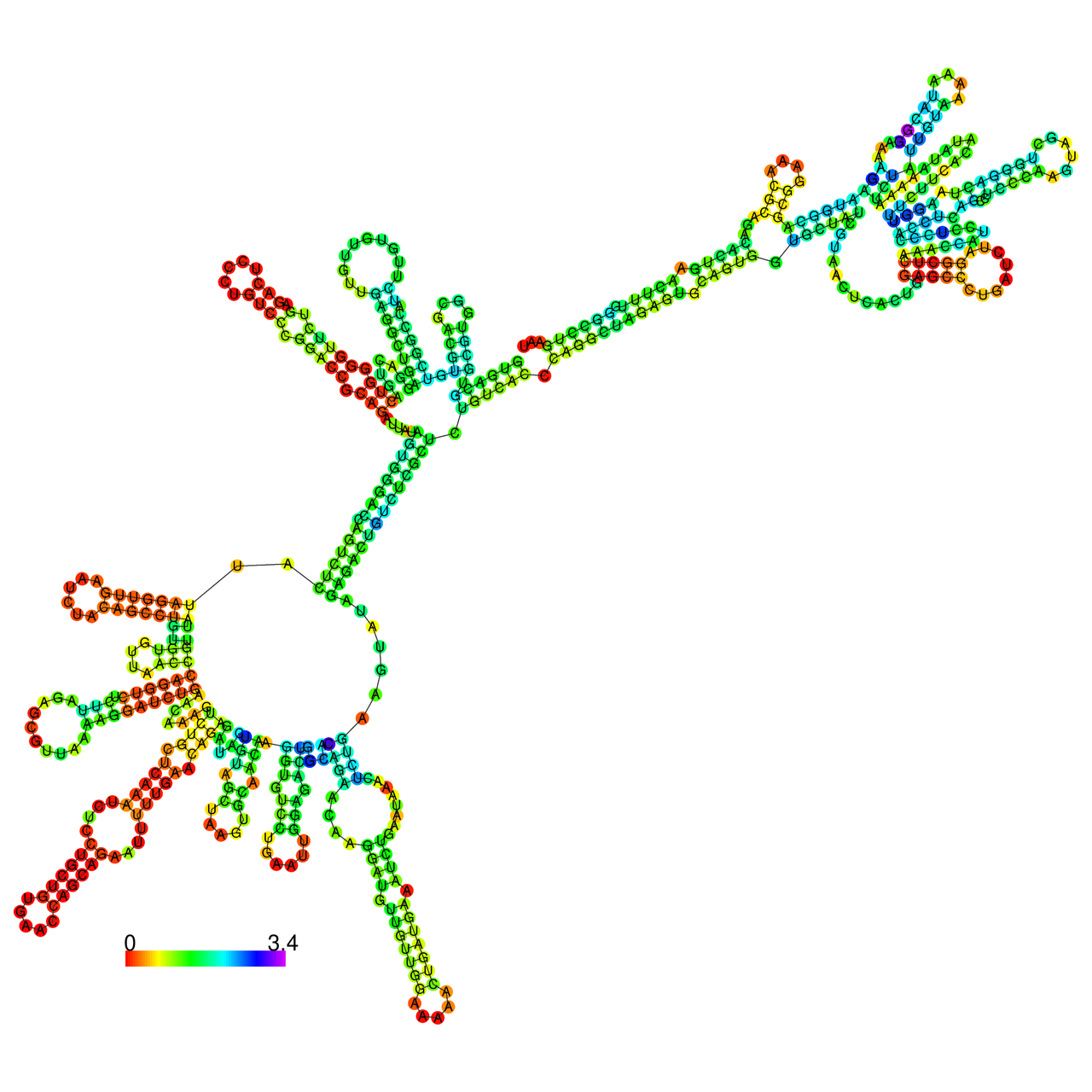
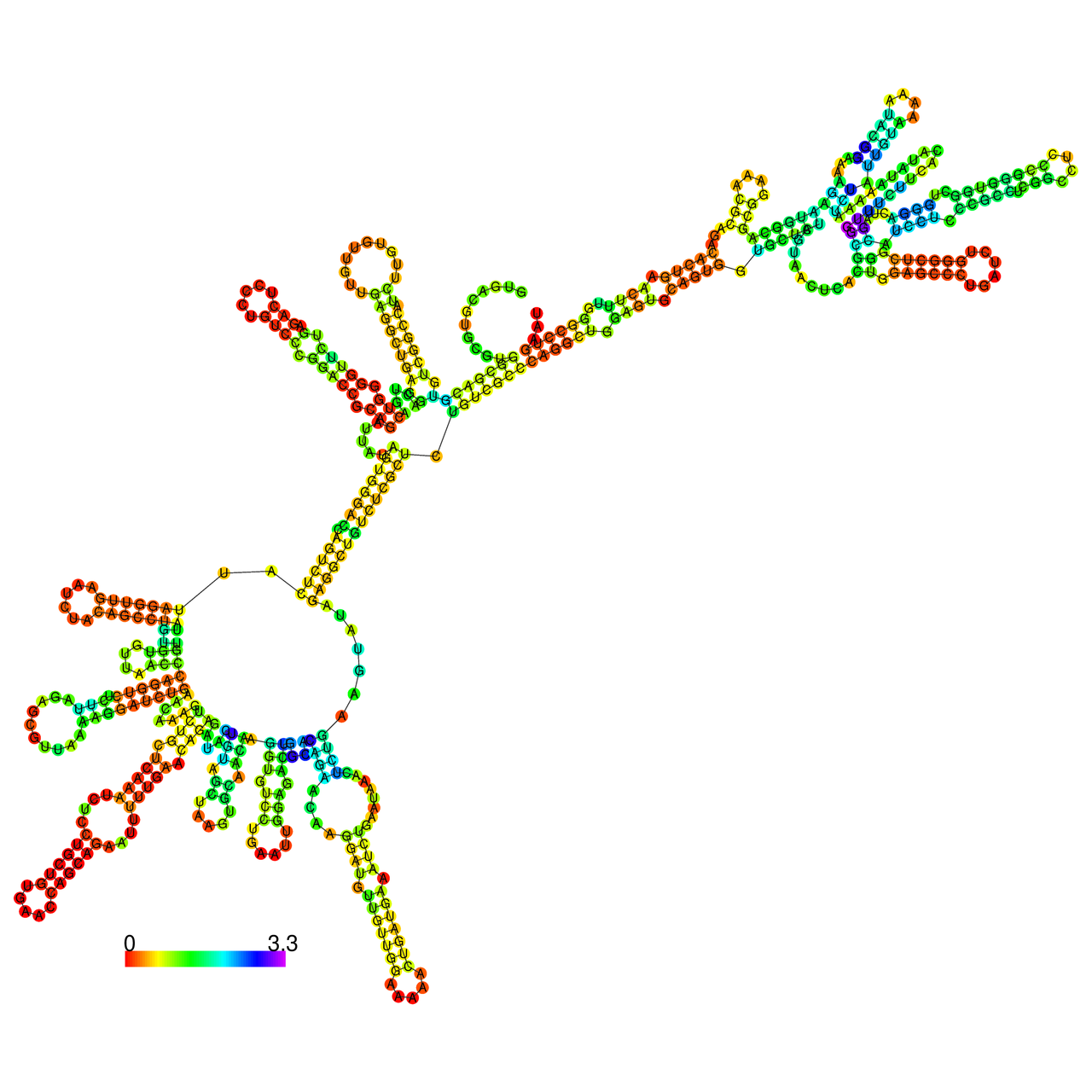
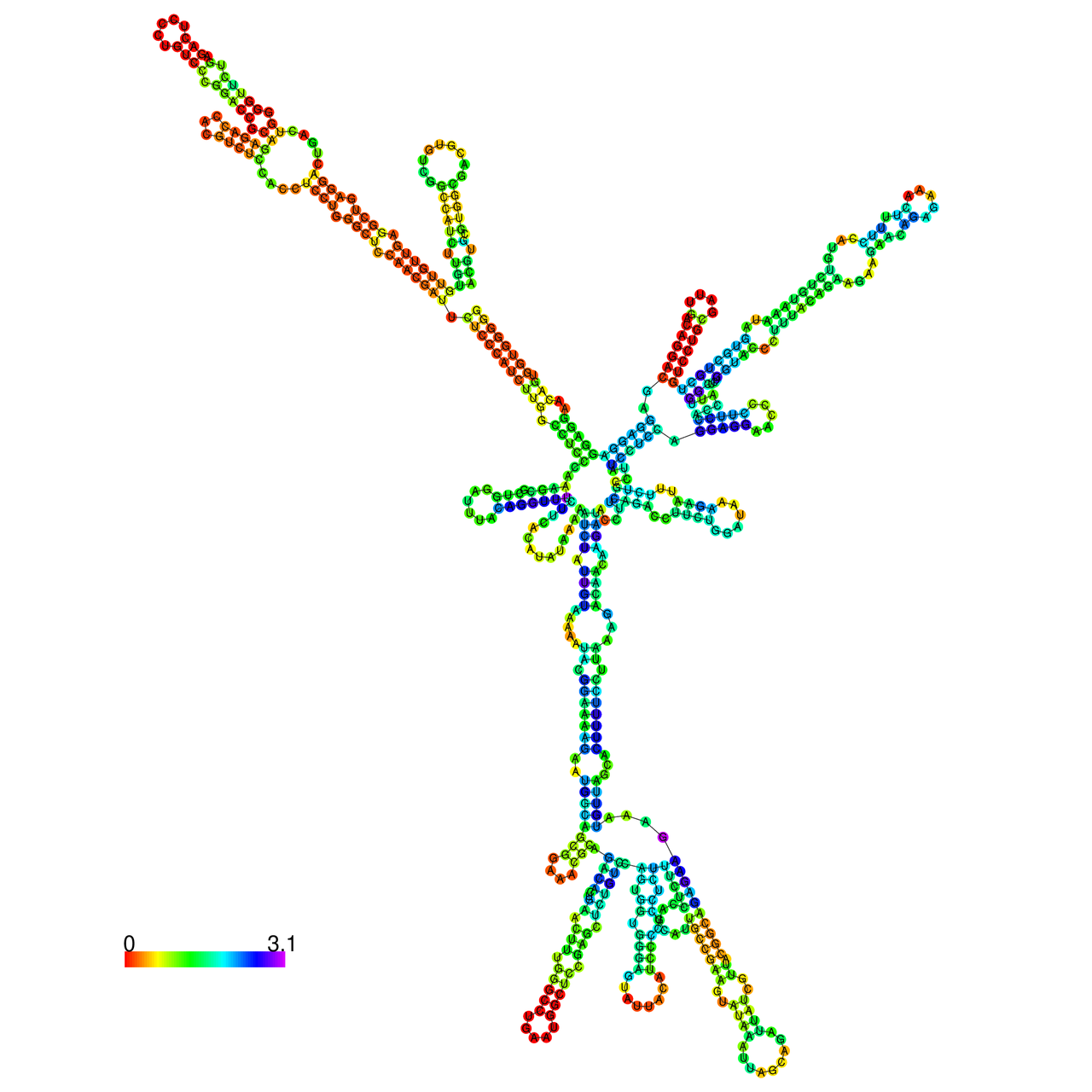
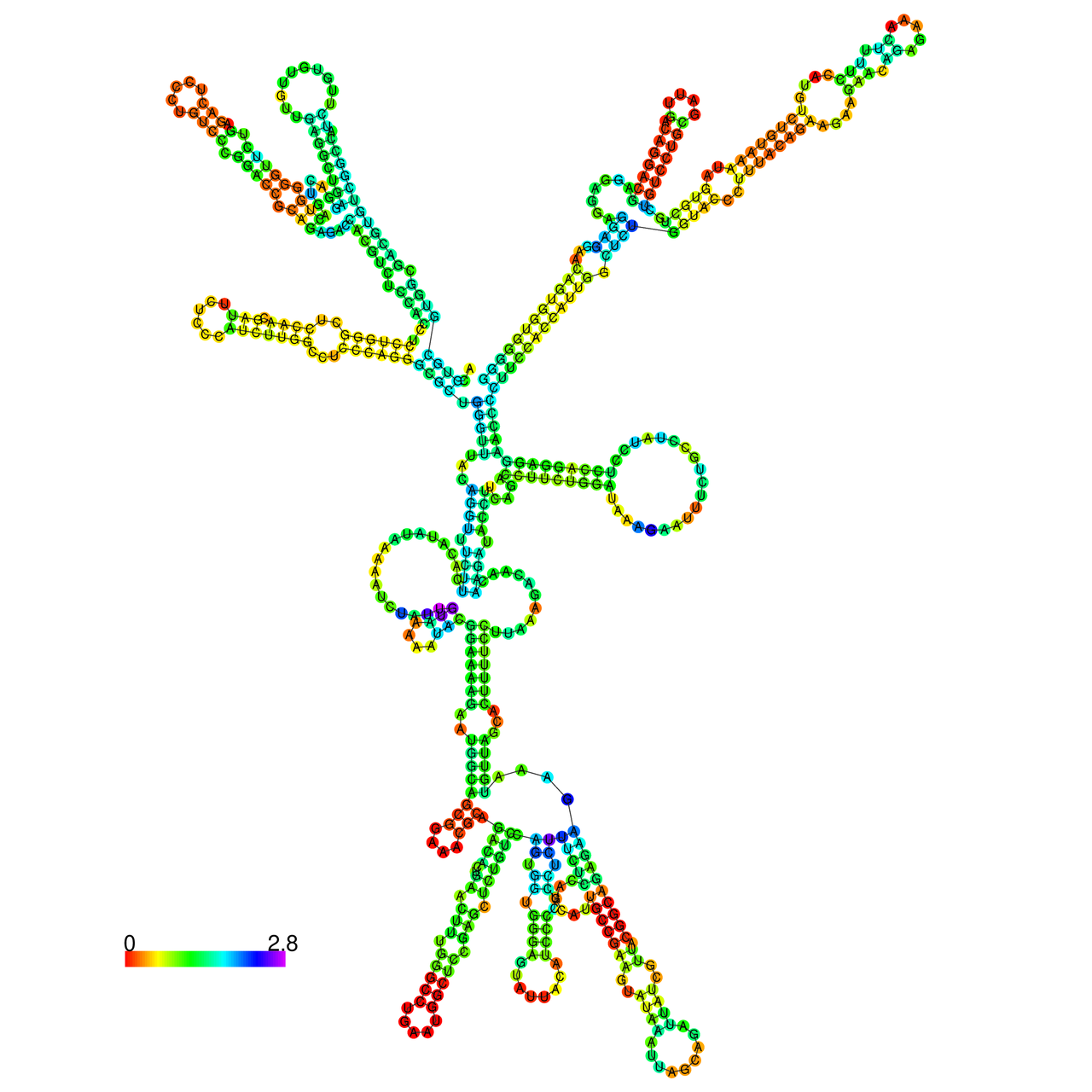
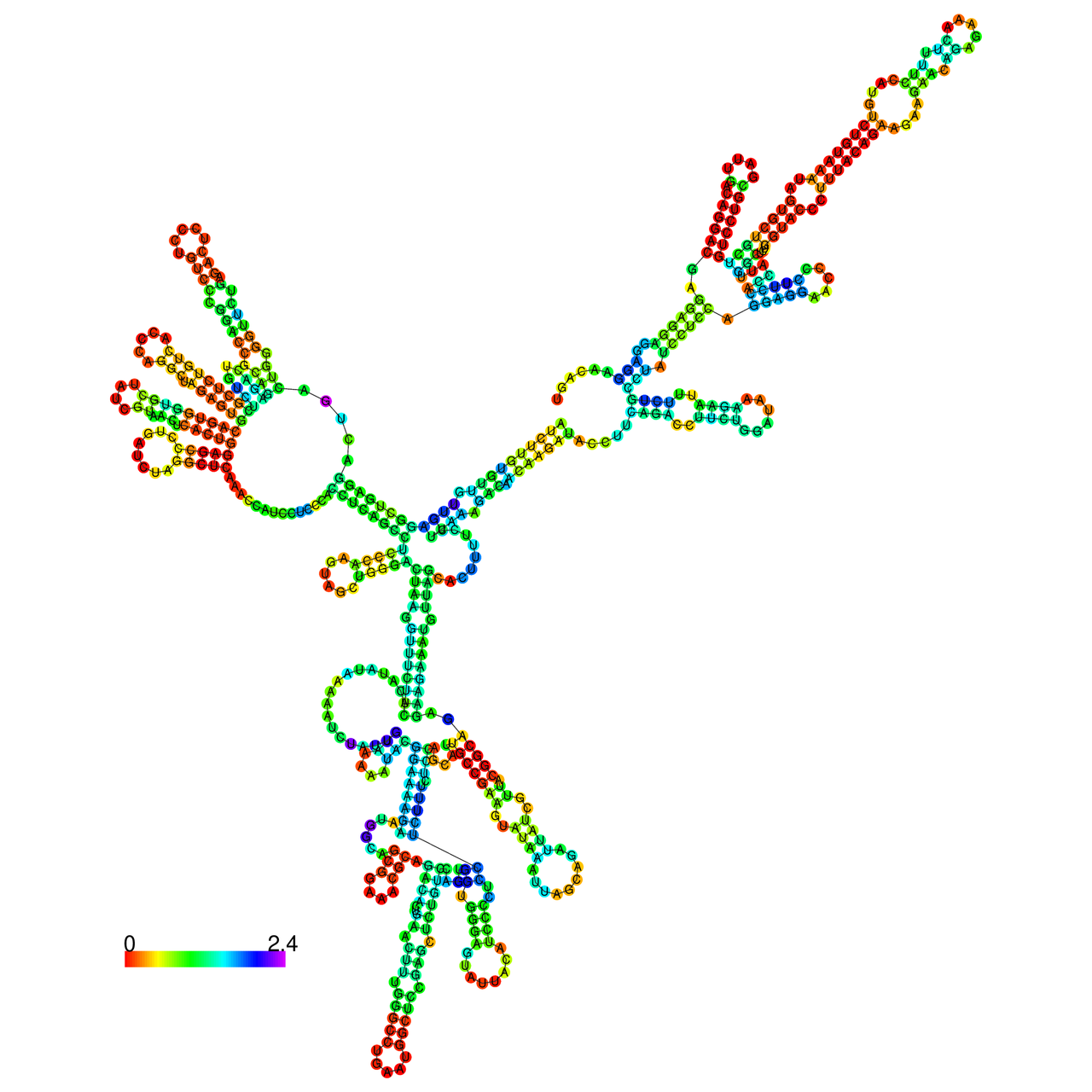
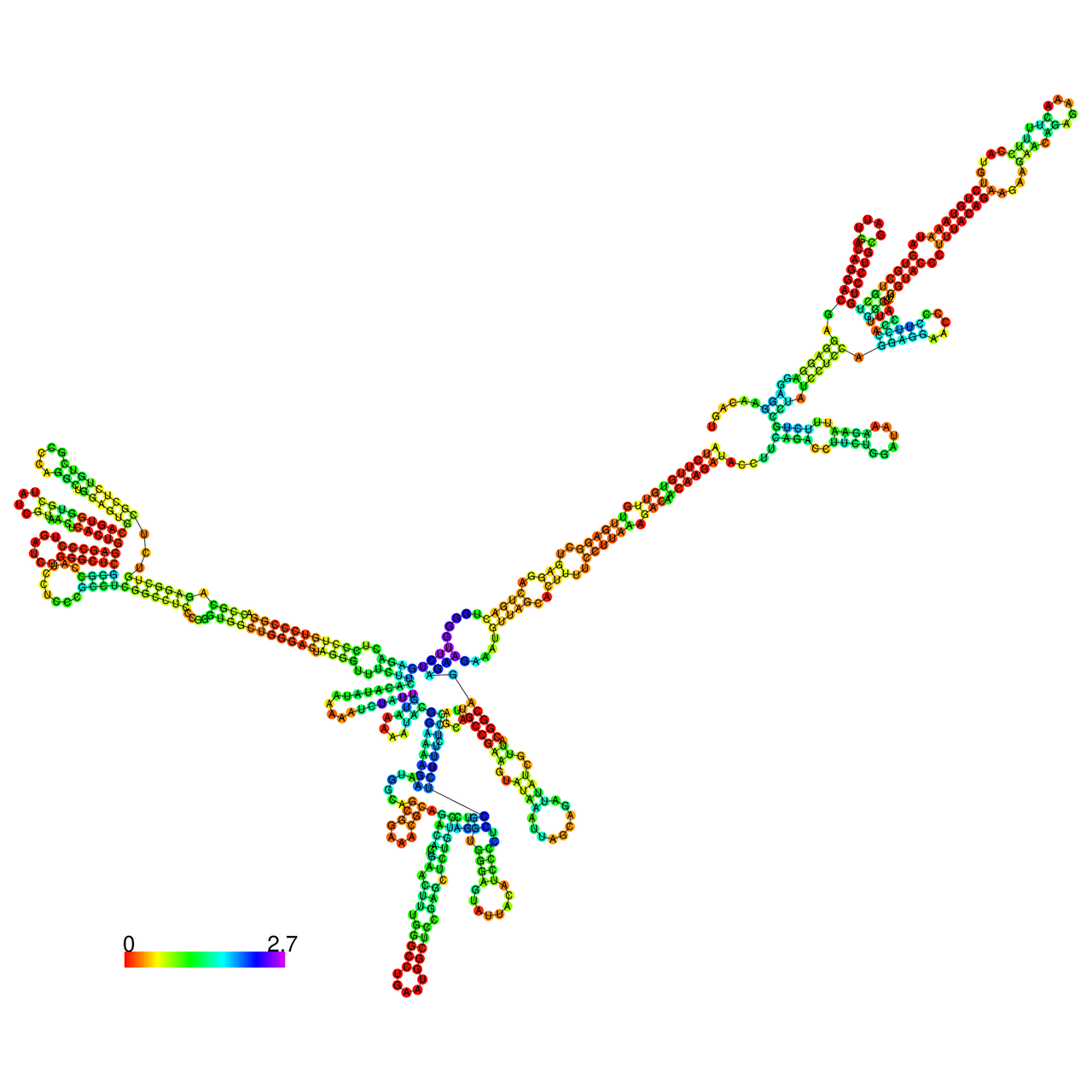
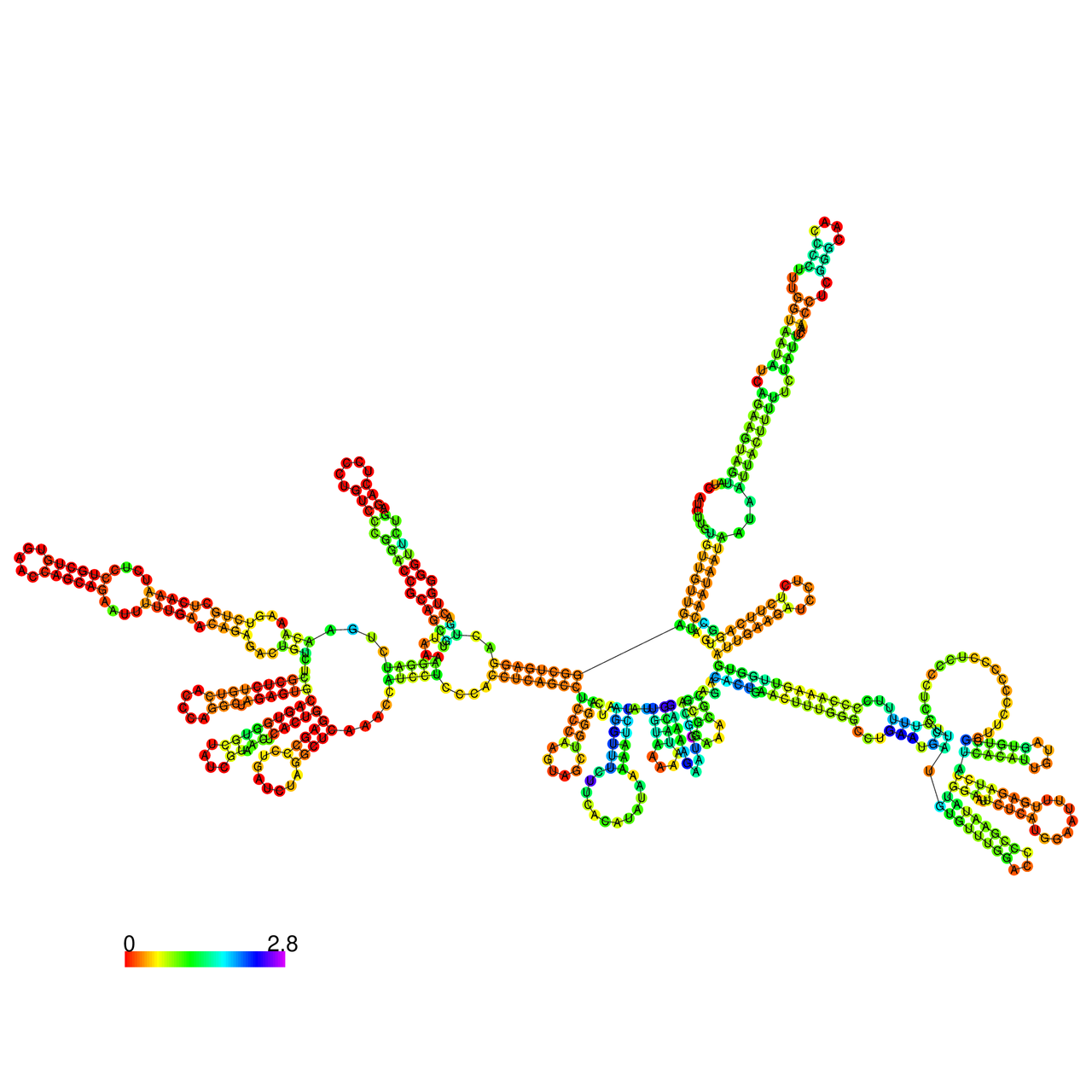
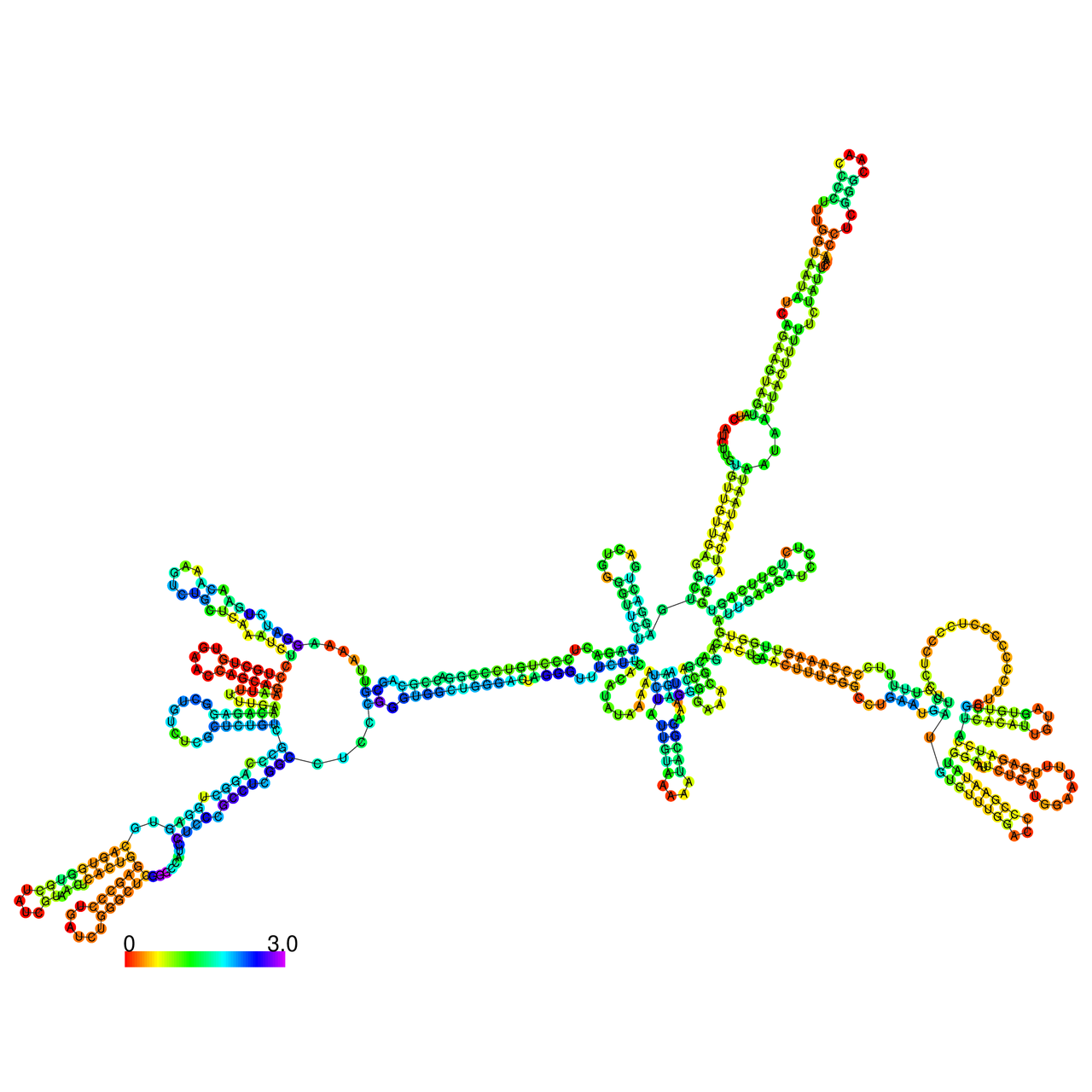
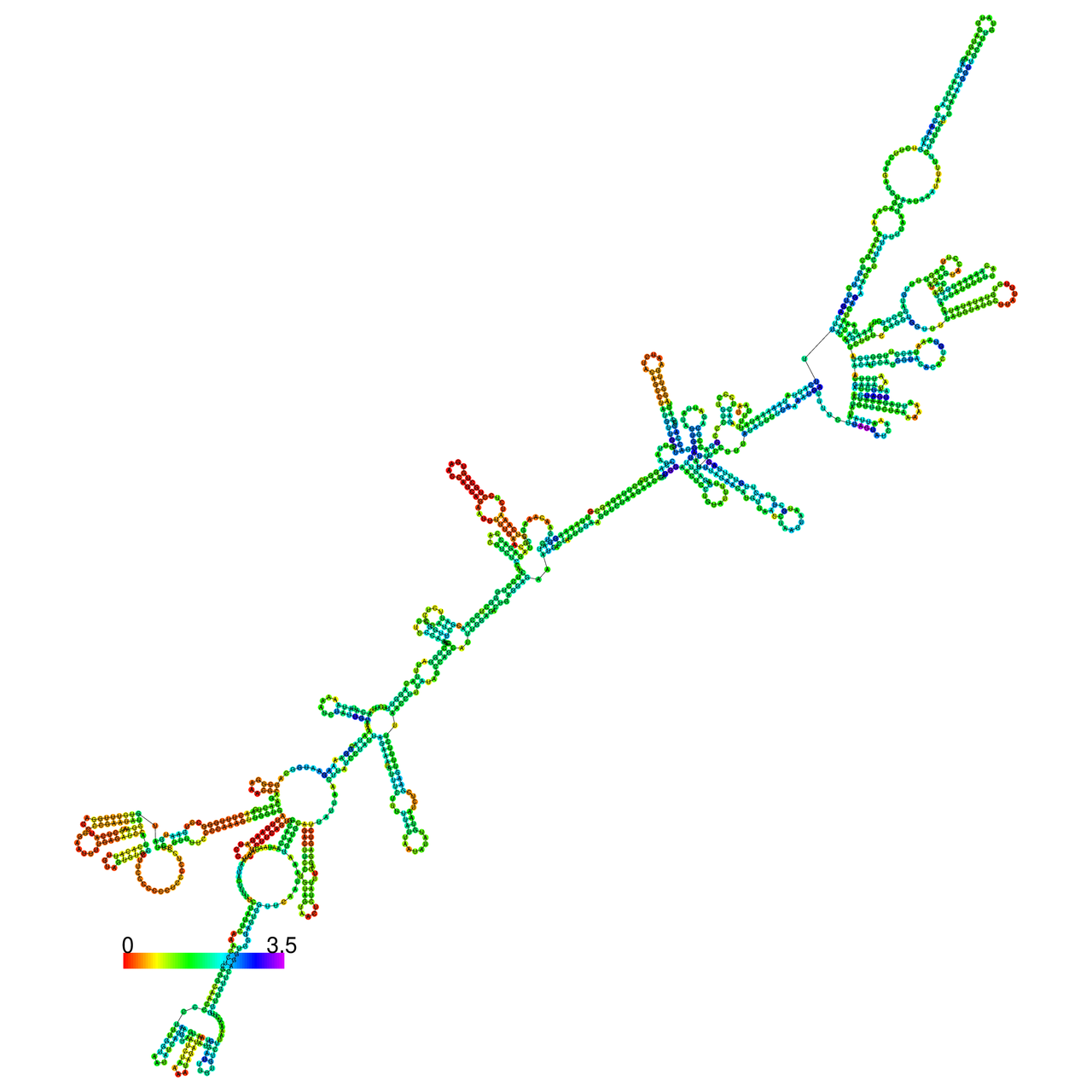
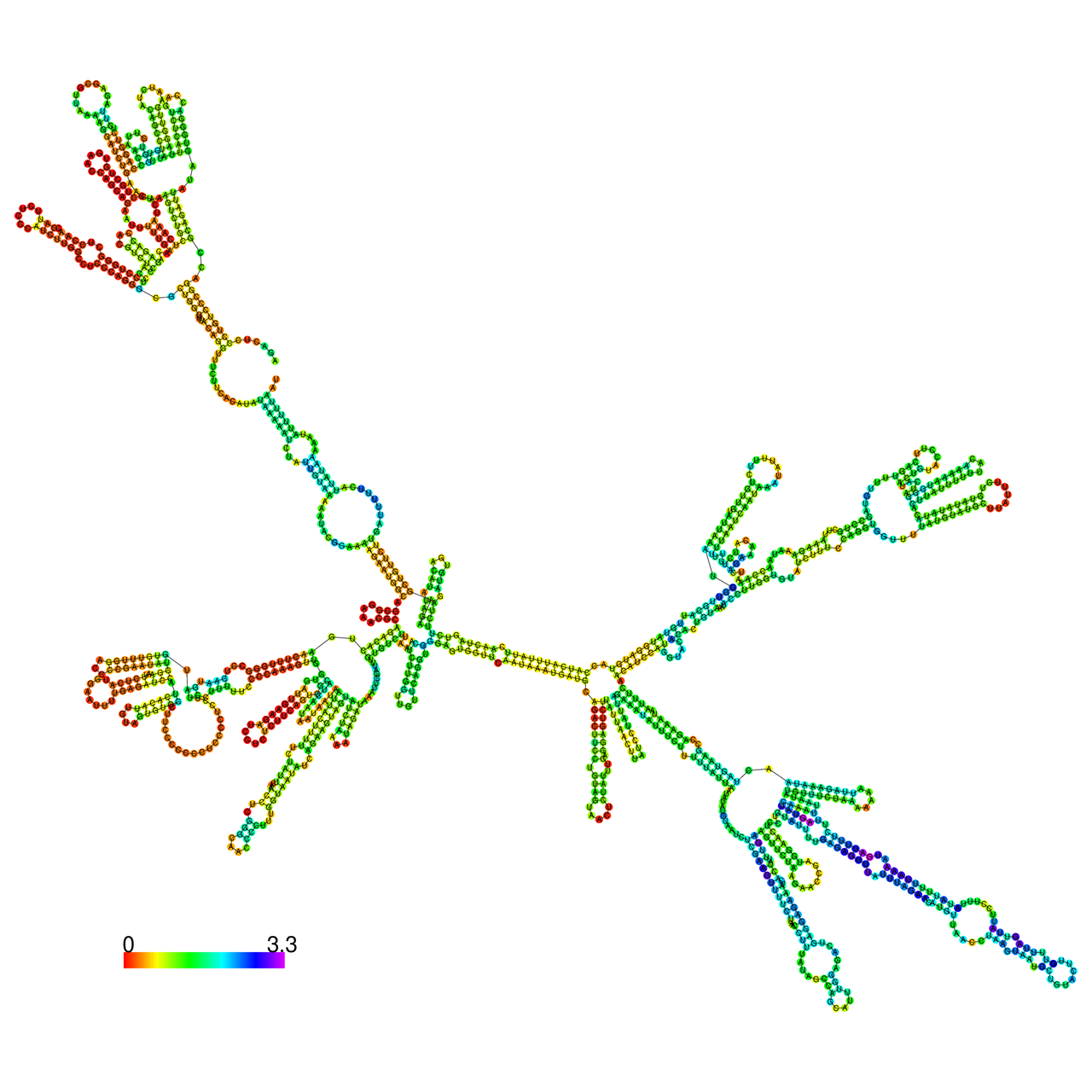
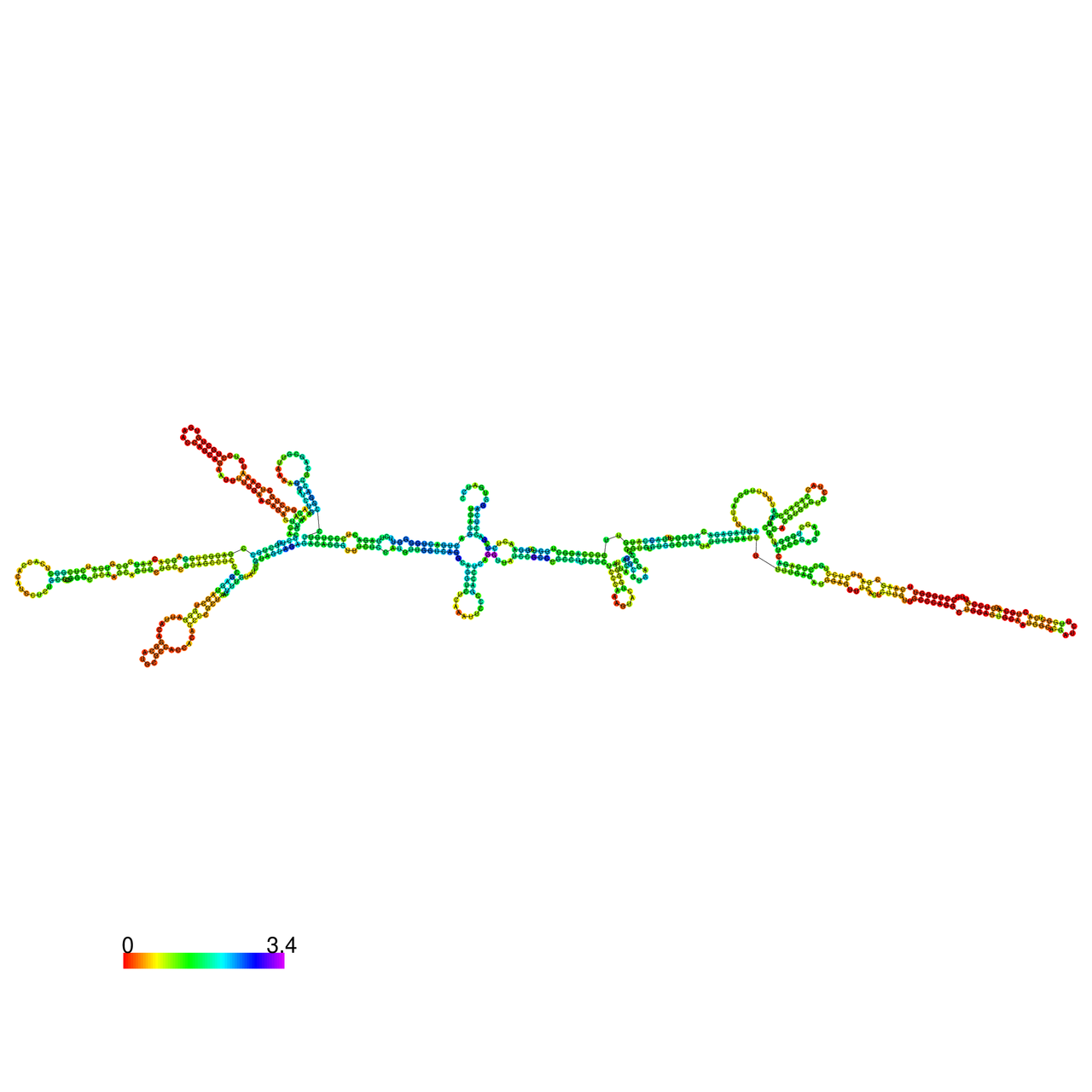
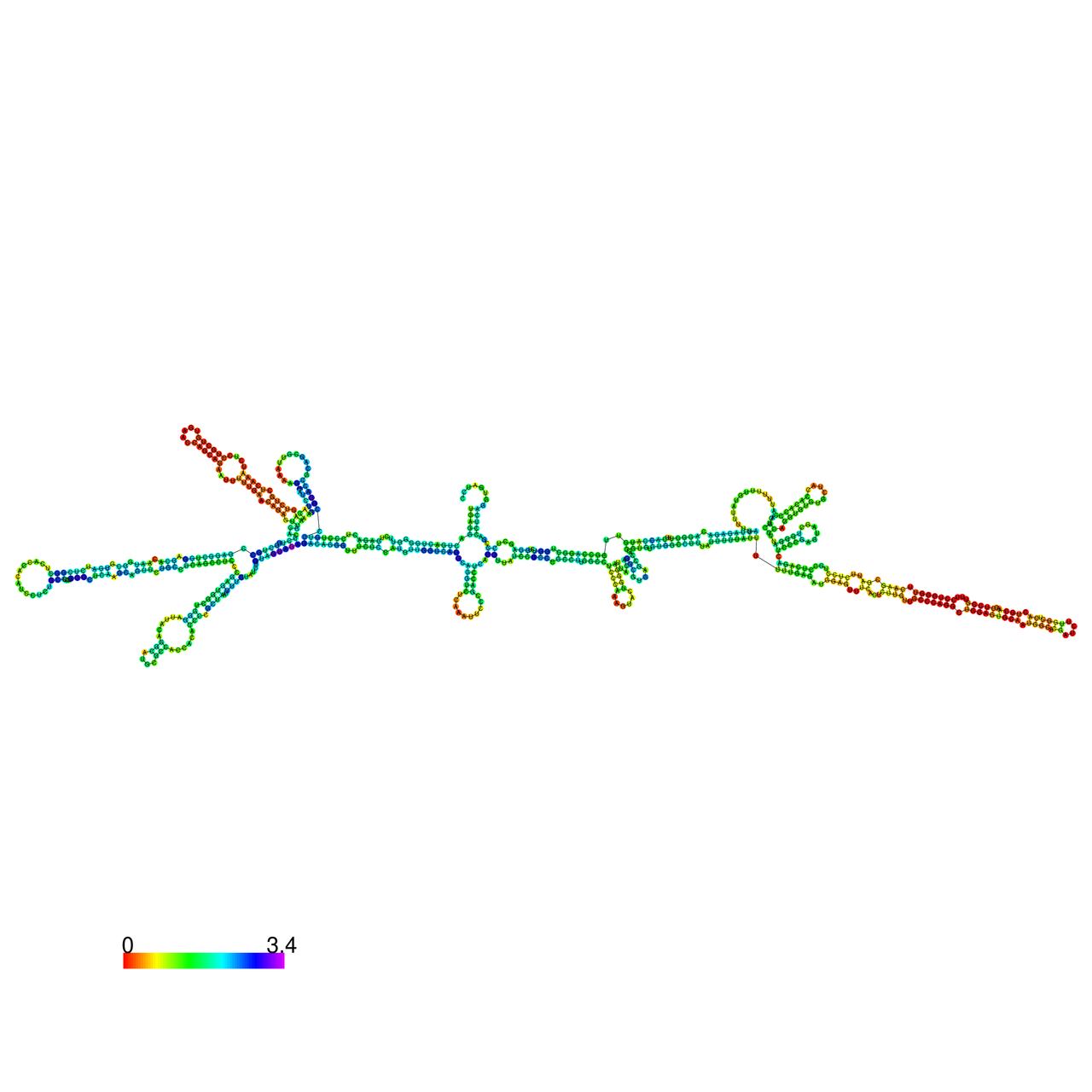
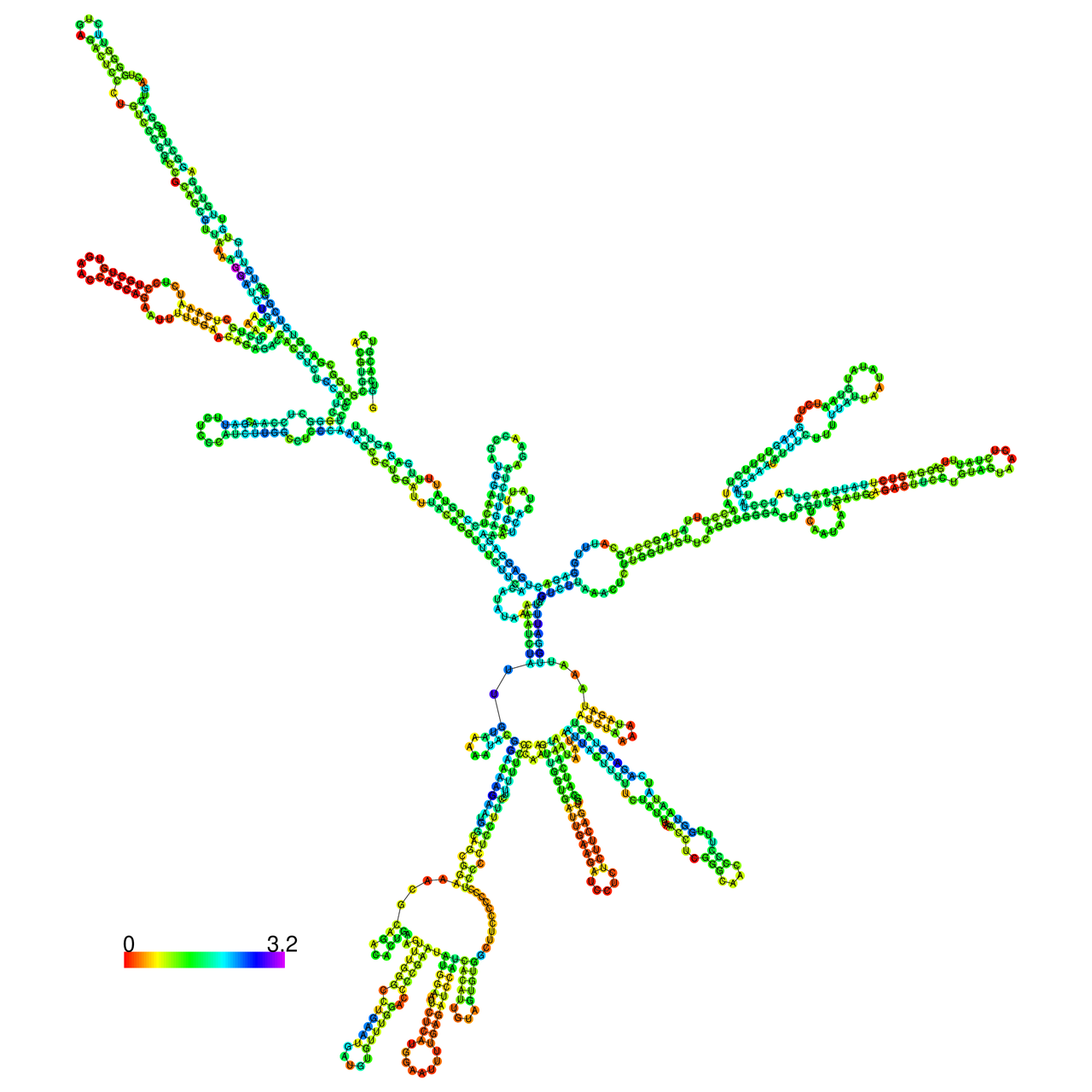
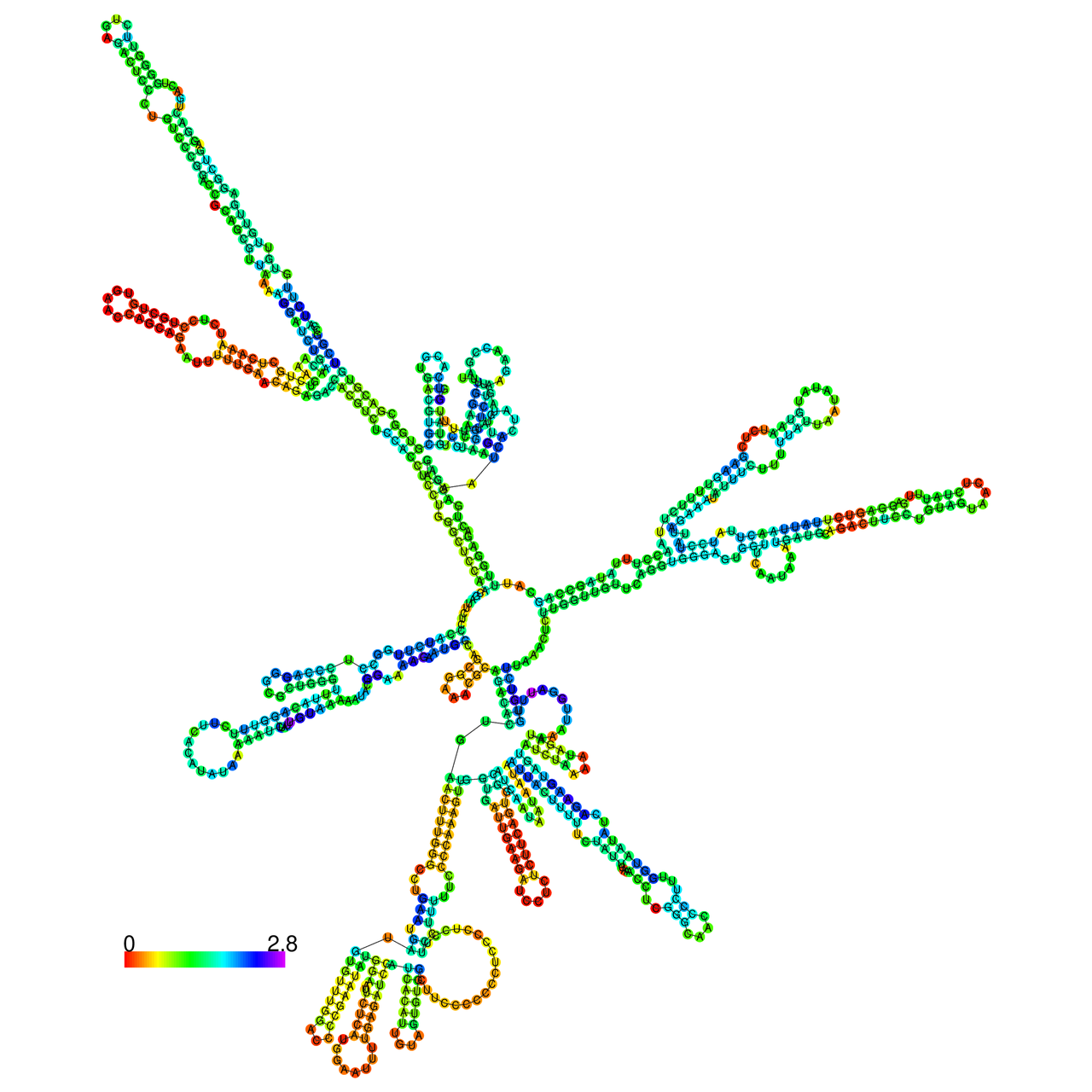
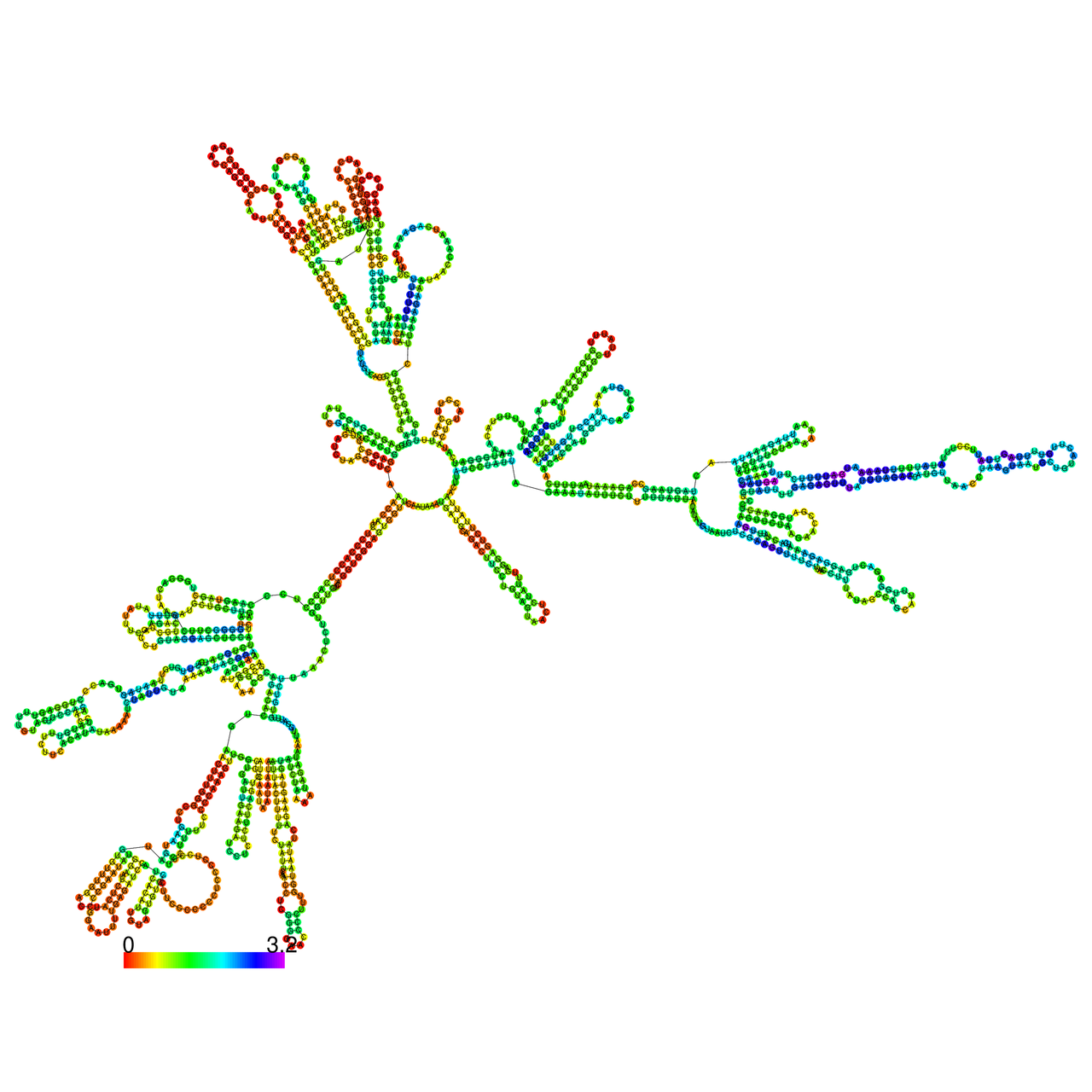
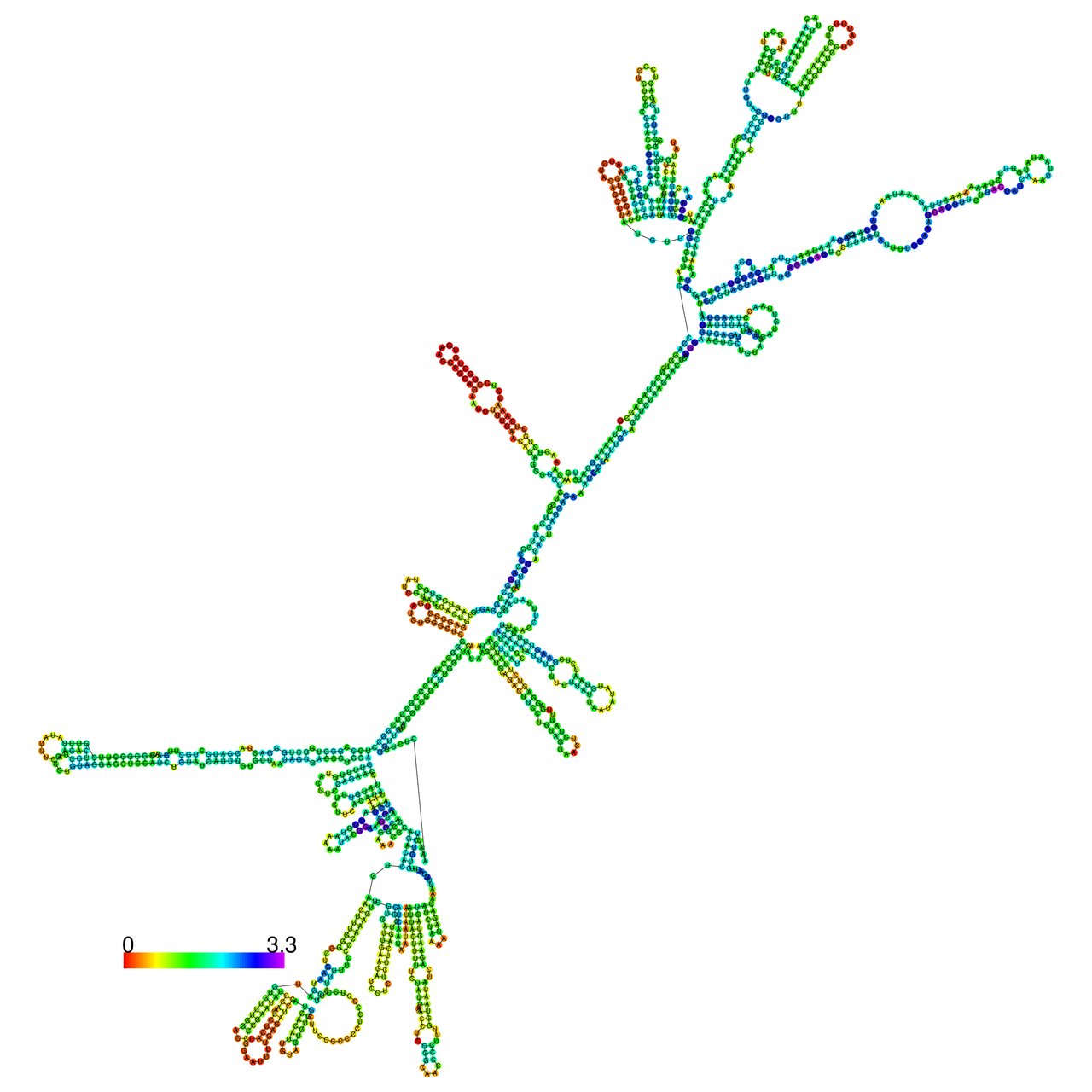
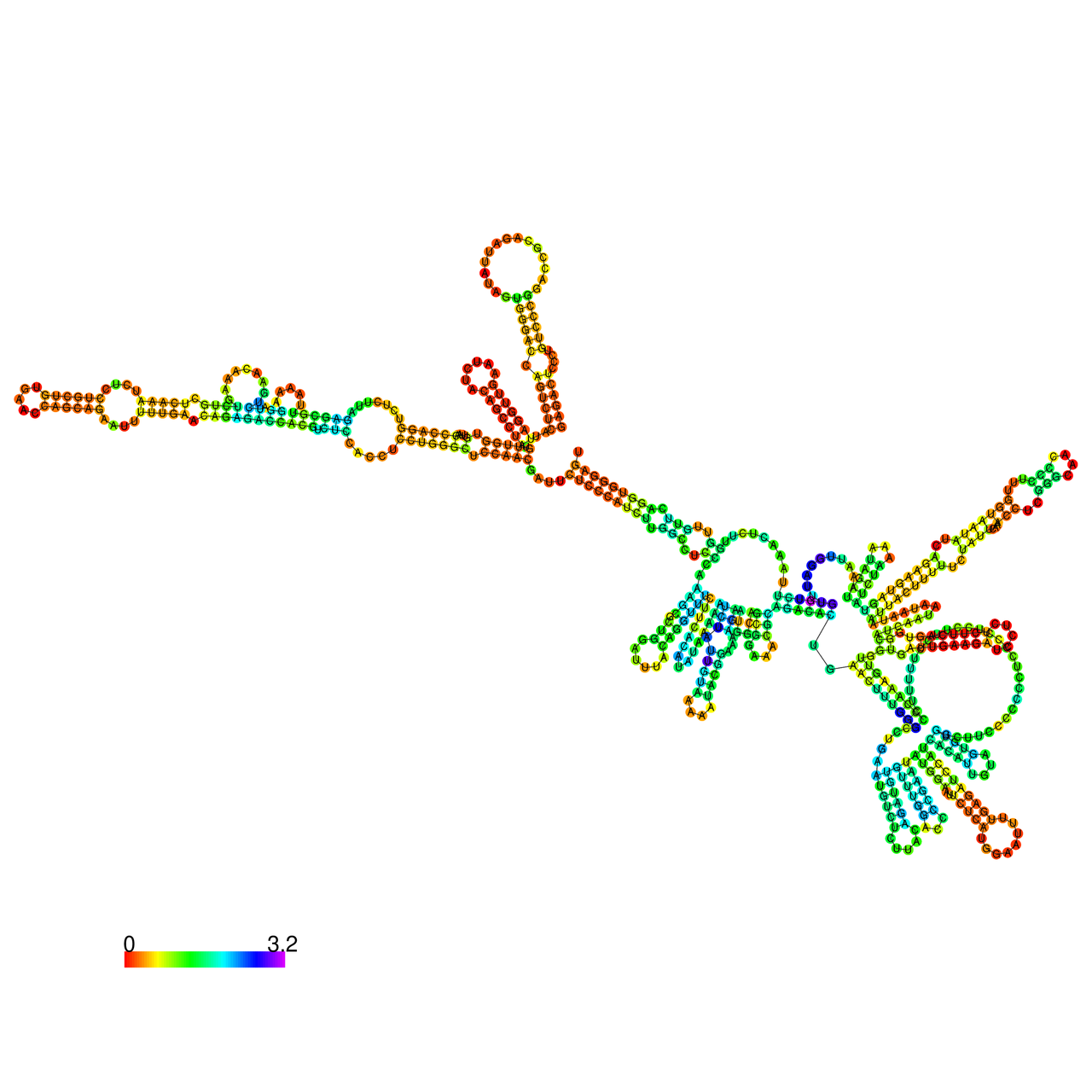
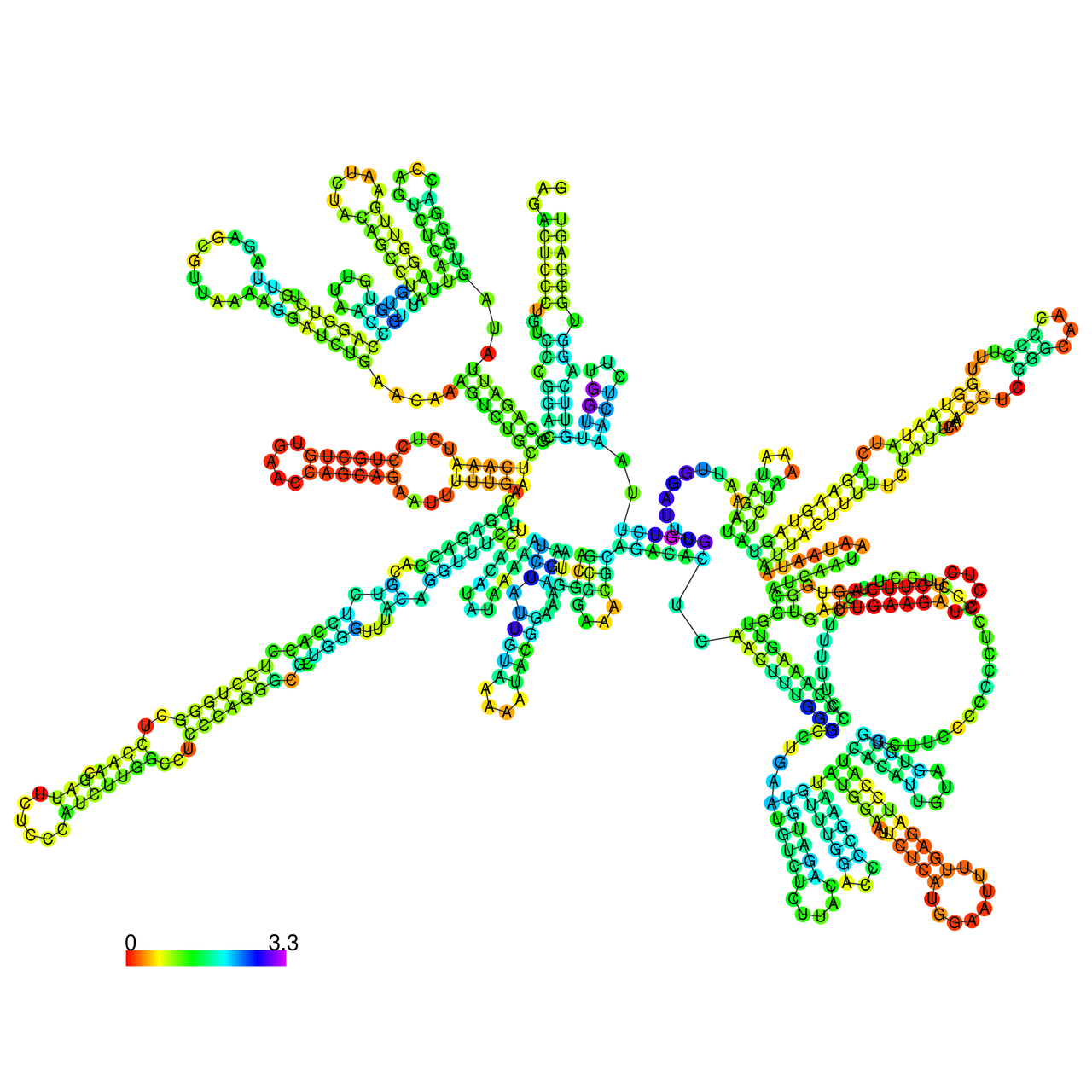
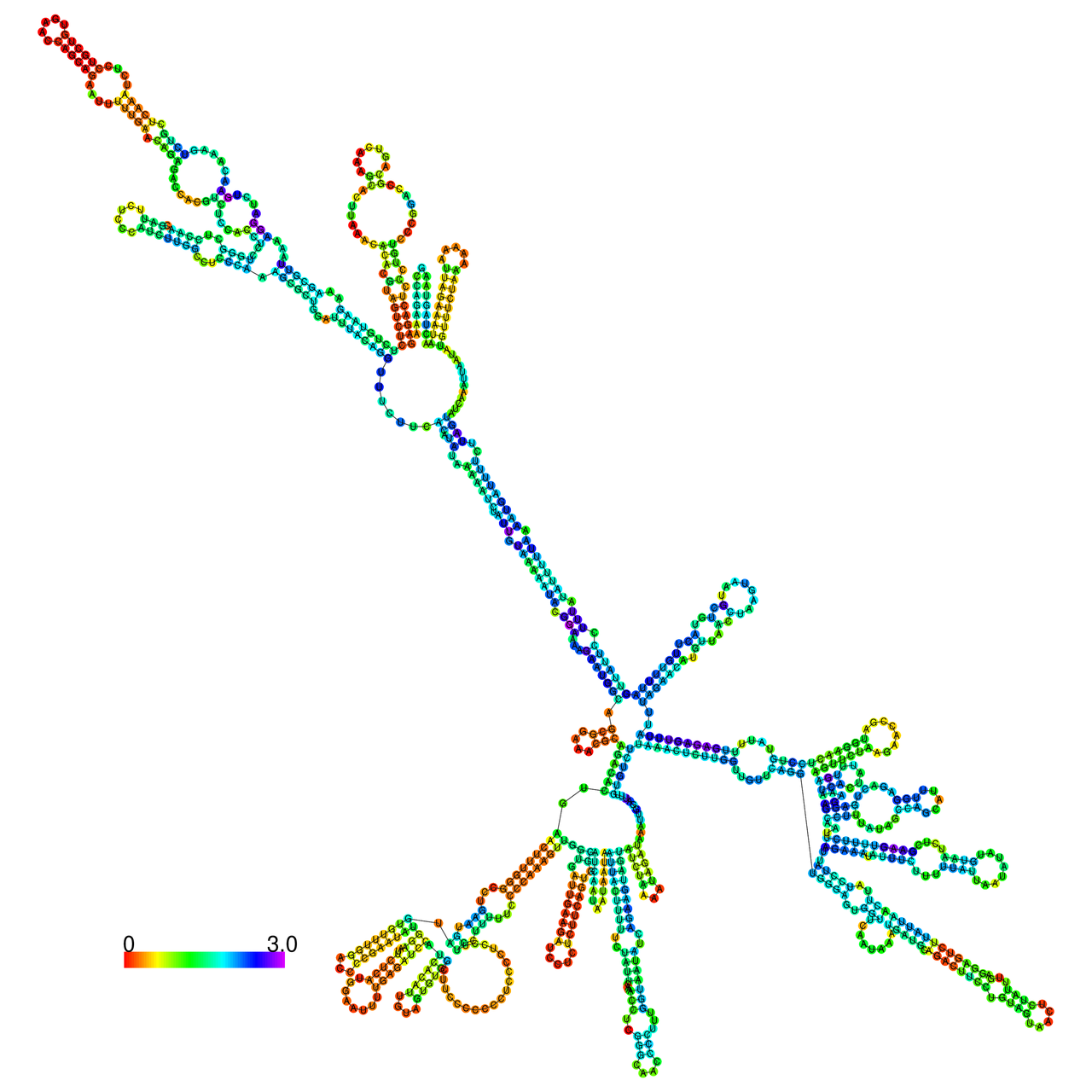
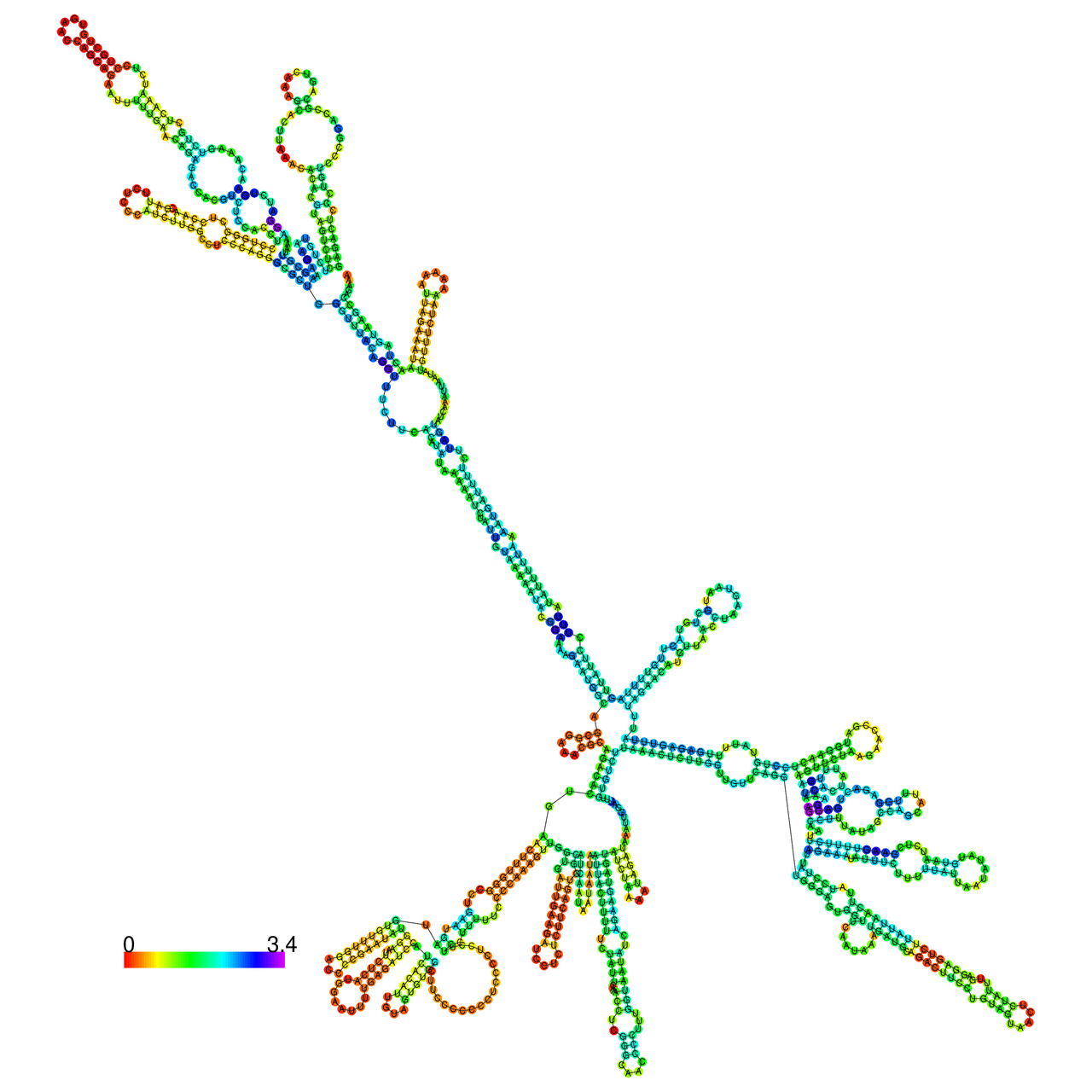
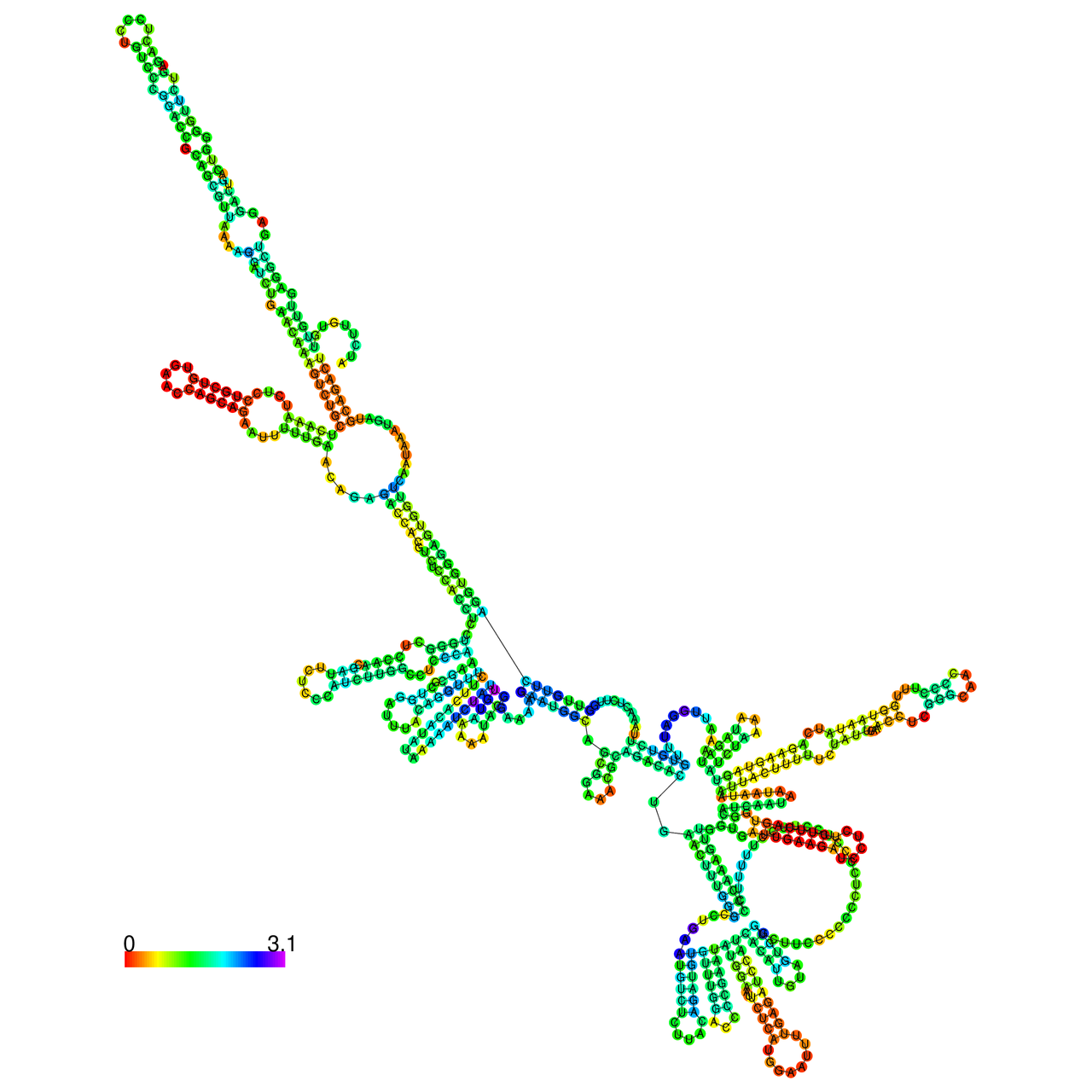
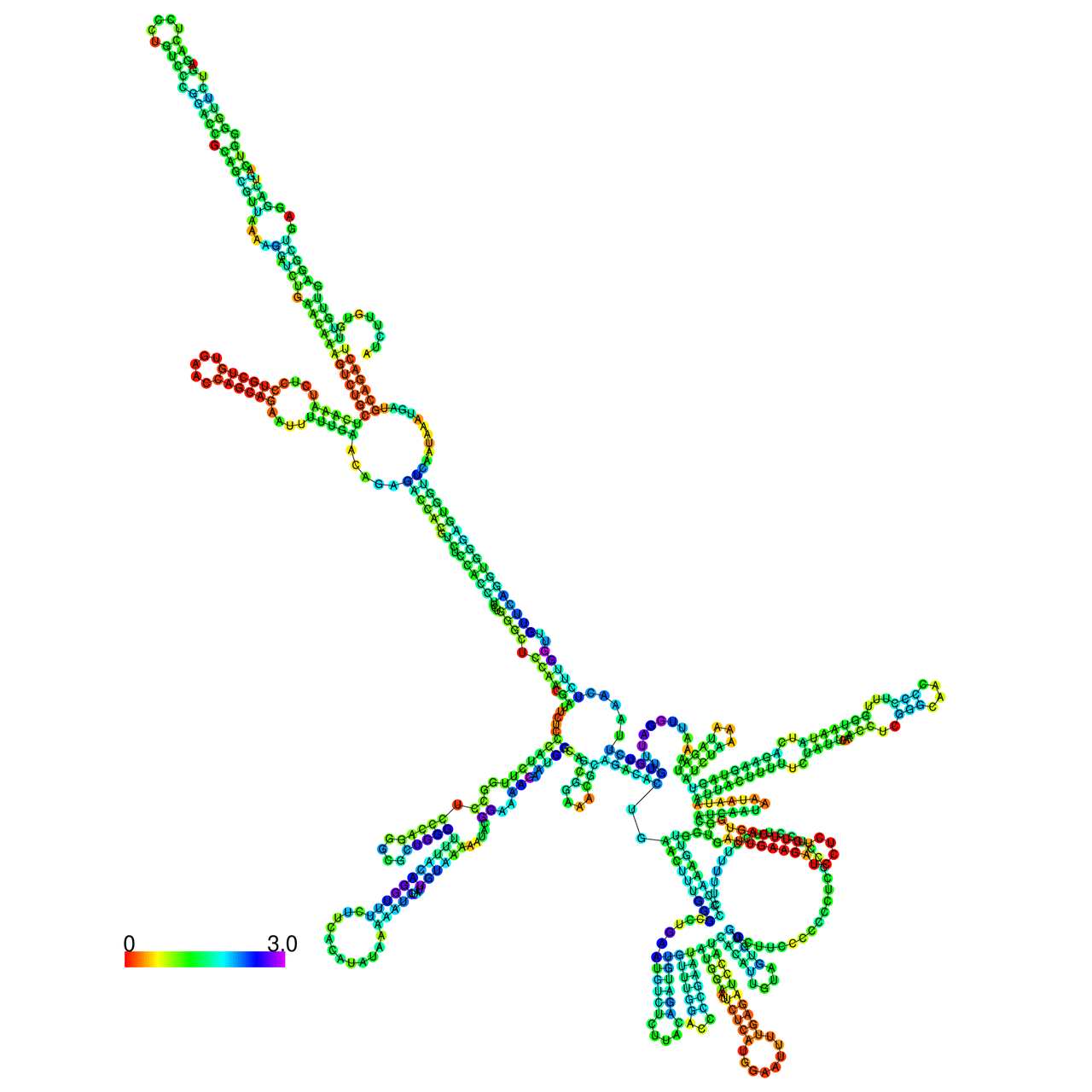
The effects of the RNA editing to the stability of the RNA structures for GIGYF2 |
 - RNA A-to-I editing in mRNA. - RNA A-to-I editing in mRNA.* Click on the image to enlarge it in a new window. |
 RNA A-to-I editing in RNA structures for Minimum free energy (MFE). RNA A-to-I editing in RNA structures for Minimum free energy (MFE). |
| CAeditom_ID | ENST | Editing_Position | MFE_withoneAI | MFE_withoutAI | MFE_withmultipleAI |
| CAediting_248954 | ENST00000409547.4 | chr2_232734470_+ | -1888.80 | -1887.50 | -1889.00 |
| CAediting_248955 | ENST00000409547.4 | chr2_232734471_+ | -1887.50 | -1887.50 | -1889.00 |
| CAediting_248956 | ENST00000409547.4 | chr2_232734479_+ | -1886.90 | -1887.50 | -1889.00 |
| CAediting_248870 | ENST00000425040.4 | chr2_232724576_+ | -172.90 | -172.90 | -185.90 |
| CAediting_248871 | ENST00000425040.4 | chr2_232724592_+ | -173.70 | -172.90 | -185.90 |
| CAediting_248872 | ENST00000425040.4 | chr2_232724601_+ | -172.20 | -172.90 | -185.90 |
| CAediting_248873 | ENST00000425040.4 | chr2_232724643_+ | -177.40 | -172.90 | -185.90 |
| CAediting_248874 | ENST00000425040.4 | chr2_232724649_+ | -173.10 | -172.90 | -185.90 |
| CAediting_248875 | ENST00000425040.4 | chr2_232724650_+ | -172.90 | -172.90 | -185.90 |
| CAediting_248876 | ENST00000425040.4 | chr2_232724651_+ | -172.90 | -172.90 | -185.90 |
| CAediting_248877 | ENST00000425040.4 | chr2_232724662_+ | -174.40 | -172.90 | -185.90 |
| CAediting_248878 | ENST00000425040.4 | chr2_232724667_+ | -172.20 | -172.90 | -185.90 |
| CAediting_248879 | ENST00000425040.4 | chr2_232724675_+ | -172.30 | -172.90 | -185.90 |
| CAediting_248880 | ENST00000425040.4 | chr2_232724676_+ | -176.50 | -172.90 | -185.90 |
| CAediting_248881 | ENST00000425040.4 | chr2_232724679_+ | -177.20 | -172.90 | -185.90 |
| CAediting_248882 | ENST00000425040.4 | chr2_232724690_+ | -175.20 | -172.90 | -185.90 |
| CAediting_248870 | ENST00000427233.4 | chr2_232724576_+ | -158.70 | -158.90 | -173.40 |
| CAediting_248871 | ENST00000427233.4 | chr2_232724592_+ | -160.20 | -158.90 | -173.40 |
| CAediting_248872 | ENST00000427233.4 | chr2_232724601_+ | -158.50 | -158.90 | -173.40 |
| CAediting_248873 | ENST00000427233.4 | chr2_232724643_+ | -163.40 | -158.90 | -173.40 |
| CAediting_248874 | ENST00000427233.4 | chr2_232724649_+ | -159.40 | -158.90 | -173.40 |
| CAediting_248875 | ENST00000427233.4 | chr2_232724650_+ | -159.00 | -158.90 | -173.40 |
| CAediting_248876 | ENST00000427233.4 | chr2_232724651_+ | -161.80 | -158.90 | -173.40 |
| CAediting_248877 | ENST00000427233.4 | chr2_232724662_+ | -160.00 | -158.90 | -173.40 |
| CAediting_248878 | ENST00000427233.4 | chr2_232724667_+ | -158.90 | -158.90 | -173.40 |
| CAediting_248879 | ENST00000427233.4 | chr2_232724675_+ | -159.70 | -158.90 | -173.40 |
| CAediting_248880 | ENST00000427233.4 | chr2_232724676_+ | -162.70 | -158.90 | -173.40 |
| CAediting_248881 | ENST00000427233.4 | chr2_232724679_+ | -162.40 | -158.90 | -173.40 |
| CAediting_248882 | ENST00000427233.4 | chr2_232724690_+ | -159.10 | -158.90 | -173.40 |
| CAediting_248954 | ENST00000428883.4 | chr2_232734470_+ | -169.40 | -168.70 | -173.20 |
| CAediting_248955 | ENST00000428883.4 | chr2_232734471_+ | -168.30 | -168.70 | -173.20 |
| CAediting_248956 | ENST00000428883.4 | chr2_232734479_+ | -168.70 | -168.70 | -173.20 |
| CAediting_248870 | ENST00000430720.4 | chr2_232724576_+ | -175.60 | -174.70 | -191.40 |
| CAediting_248871 | ENST00000430720.4 | chr2_232724592_+ | -175.50 | -174.70 | -191.40 |
| CAediting_248872 | ENST00000430720.4 | chr2_232724601_+ | -174.00 | -174.70 | -191.40 |
| CAediting_248873 | ENST00000430720.4 | chr2_232724643_+ | -179.20 | -174.70 | -191.40 |
| CAediting_248874 | ENST00000430720.4 | chr2_232724649_+ | -175.50 | -174.70 | -191.40 |
| CAediting_248875 | ENST00000430720.4 | chr2_232724650_+ | -175.60 | -174.70 | -191.40 |
| CAediting_248876 | ENST00000430720.4 | chr2_232724651_+ | -175.70 | -174.70 | -191.40 |
| CAediting_248877 | ENST00000430720.4 | chr2_232724662_+ | -178.10 | -174.70 | -191.40 |
| CAediting_248878 | ENST00000430720.4 | chr2_232724667_+ | -174.00 | -174.70 | -191.40 |
| CAediting_248879 | ENST00000430720.4 | chr2_232724675_+ | -174.00 | -174.70 | -191.40 |
| CAediting_248880 | ENST00000430720.4 | chr2_232724676_+ | -178.30 | -174.70 | -191.40 |
| CAediting_248881 | ENST00000430720.4 | chr2_232724679_+ | -177.40 | -174.70 | -191.40 |
| CAediting_248882 | ENST00000430720.4 | chr2_232724690_+ | -174.40 | -174.70 | -191.40 |
| CAediting_248870 | ENST00000464402.4 | chr2_232724576_+ | -152.00 | -152.00 | -165.00 |
| CAediting_248871 | ENST00000464402.4 | chr2_232724592_+ | -152.80 | -152.00 | -165.00 |
| CAediting_248872 | ENST00000464402.4 | chr2_232724601_+ | -151.30 | -152.00 | -165.00 |
| CAediting_248873 | ENST00000464402.4 | chr2_232724643_+ | -156.50 | -152.00 | -165.00 |
| CAediting_248874 | ENST00000464402.4 | chr2_232724649_+ | -152.20 | -152.00 | -165.00 |
| CAediting_248875 | ENST00000464402.4 | chr2_232724650_+ | -152.00 | -152.00 | -165.00 |
| CAediting_248876 | ENST00000464402.4 | chr2_232724651_+ | -153.60 | -152.00 | -165.00 |
| CAediting_248877 | ENST00000464402.4 | chr2_232724662_+ | -152.20 | -152.00 | -165.00 |
| CAediting_248878 | ENST00000464402.4 | chr2_232724667_+ | -151.30 | -152.00 | -165.00 |
| CAediting_248879 | ENST00000464402.4 | chr2_232724675_+ | -151.30 | -152.00 | -165.00 |
| CAediting_248880 | ENST00000464402.4 | chr2_232724676_+ | -155.60 | -152.00 | -165.00 |
| CAediting_248881 | ENST00000464402.4 | chr2_232724679_+ | -153.30 | -152.00 | -165.00 |
| CAediting_248882 | ENST00000464402.4 | chr2_232724690_+ | -152.60 | -152.00 | -165.00 |
| CAediting_248954 | ENST00000464805.4 | chr2_232734470_+ | -273.00 | -270.80 | -279.30 |
| CAediting_248955 | ENST00000464805.4 | chr2_232734471_+ | -273.90 | -270.80 | -279.30 |
| CAediting_248956 | ENST00000464805.4 | chr2_232734479_+ | -272.90 | -270.80 | -279.30 |
| CAediting_248823 | ENST00000474465.4 | chr2_232705535_+ | -215.60 | -215.60 | -216.30 |
| CAediting_248824 | ENST00000474465.4 | chr2_232705927_+ | -216.60 | -215.60 | -216.30 |
| CAediting_248954 | ENST00000476995.4 | chr2_232734470_+ | -203.10 | -200.50 | -205.30 |
| CAediting_248955 | ENST00000476995.4 | chr2_232734471_+ | -203.30 | -200.50 | -205.30 |
| CAediting_248956 | ENST00000476995.4 | chr2_232734479_+ | -201.70 | -200.50 | -205.30 |
| CAediting_248870 | ENST00000482666.4 | chr2_232724576_+ | -312.20 | -312.80 | -325.90 |
| CAediting_248871 | ENST00000482666.4 | chr2_232724592_+ | -312.80 | -312.80 | -325.90 |
| CAediting_248872 | ENST00000482666.4 | chr2_232724601_+ | -312.50 | -312.80 | -325.90 |
| CAediting_248873 | ENST00000482666.4 | chr2_232724643_+ | -317.30 | -312.80 | -325.90 |
| CAediting_248874 | ENST00000482666.4 | chr2_232724649_+ | -316.70 | -312.80 | -325.90 |
| CAediting_248875 | ENST00000482666.4 | chr2_232724650_+ | -313.20 | -312.80 | -325.90 |
| CAediting_248876 | ENST00000482666.4 | chr2_232724651_+ | -312.80 | -312.80 | -325.90 |
| CAediting_248877 | ENST00000482666.4 | chr2_232724662_+ | -312.40 | -312.80 | -325.90 |
| CAediting_248878 | ENST00000482666.4 | chr2_232724667_+ | -312.00 | -312.80 | -325.90 |
| CAediting_248879 | ENST00000482666.4 | chr2_232724675_+ | -312.50 | -312.80 | -325.90 |
| CAediting_248880 | ENST00000482666.4 | chr2_232724676_+ | -316.20 | -312.80 | -325.90 |
| CAediting_248881 | ENST00000482666.4 | chr2_232724679_+ | -313.70 | -312.80 | -325.90 |
| CAediting_248882 | ENST00000482666.4 | chr2_232724690_+ | -312.80 | -312.80 | -325.90 |
| CAediting_248954 | ENST00000483164.4 | chr2_232734470_+ | -156.20 | -152.70 | -156.60 |
| CAediting_248955 | ENST00000483164.4 | chr2_232734471_+ | -154.60 | -152.70 | -156.60 |
| CAediting_248956 | ENST00000483164.4 | chr2_232734479_+ | -152.70 | -152.70 | -156.60 |
| CAediting_248954 | ENST00000490229.4 | chr2_232734470_+ | -195.70 | -192.50 | -198.40 |
| CAediting_248955 | ENST00000490229.4 | chr2_232734471_+ | -195.20 | -192.50 | -198.40 |
| CAediting_248956 | ENST00000490229.4 | chr2_232734479_+ | -193.60 | -192.50 | -198.40 |
| CAediting_248954 | ENST00000490612.4 | chr2_232734470_+ | -153.60 | -151.40 | -156.40 |
| CAediting_248955 | ENST00000490612.4 | chr2_232734471_+ | -152.40 | -151.40 | -156.40 |
| CAediting_248956 | ENST00000490612.4 | chr2_232734479_+ | -152.80 | -151.40 | -156.40 |
| CAediting_249057 | ENST00000629305.1 | chr2_232860206_+ | -2498.20 | -2498.10 | -2500.00 |
| CAediting_249058 | ENST00000629305.1 | chr2_232860232_+ | -2498.30 | -2498.10 | -2500.00 |
Top |
Relation with ADAR for GIGYF2 |
 Correlation between ADAR gene expression and RNA editing frequency Correlation between ADAR gene expression and RNA editing frequency |
| Cancer | CAeditomID | position | ADAR1 (p-val) | ADAR1 (coeff.) | ADAR2 (p-val) | ADAR2 (coeff.) | ADAR3 (p-val) | ADAR3 (coeff.) |
| STAD | CAediting_248900 | chr2_232726209_+ | 0.0466214017343898 | 0.24217450092233 | . | . | . | . |
Top |
Relation with cancer stages for GIGYF2 |
 Correlation between Cancer stages and RNA editing frequency Correlation between Cancer stages and RNA editing frequency |
| Cancer | Stage_type | CAeditomID | position | P-val | Coeff. |
Top |
Relation with survival for GIGYF2 |
 Correlation between RNA editing and survival Correlation between RNA editing and survival |
| Cancer | CAeditomID | position | KMpvalue | Coxpvalue | CoxHR |
Top |
RelatedDrugs for GIGYF2 |
 Approved drugs targeting this gene. Approved drugs targeting this gene. (DrugBank Version 5.1.8 2021-01-03) |
| Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
RelatedDiseases for GIGYF2 |
 Diseases associated with this gene. Diseases associated with this gene. (DisGeNet 7.0) |
| Gene | Disease ID | Disease name | # pubmeds | Source |
| GIGYF2 | C4083045 | PARKINSON DISEASE 11, AUTOSOMAL DOMINANT, SUSCEPTIBILITY TO | 5 | CTD_human;GENOMICS_ENGLAND;UNIPROT |
| GIGYF2 | C1535926 | Neurodevelopmental Disorders | 1 | CTD_human |